dNTP stands for deoxyribose nucleotide triphosphate employed in PCR to expand the growing DNA strand. dATP, dTTP, dGTP and dTTP are four common dNTPs used in PCR.
The function of dNTPs in PCR is to expand the growing DNA strand with the help of Taq DNA polymerase. It binds with the complementary DNA strand by hydrogen bonds.
The PCR is an in vitro technique of DNA synthesis. The purpose of performing PCR is to generate multiple copies of DNA fragments of our interest to visualize it under gel electrophoresis.
The aim of PCR is to make millions of DNA copies for various downstream applications like DNA sequencing or DNA microarray.
The polymerase chain reaction is the unmatched tool used in molecular genetic research since its discovery. DNA template, primers, buffer, Taq DNA polymerase and dNTPs are the ingredients of PCR. To understand the mechanism of PCR we should have to learn the importance of every ingredient used in it,
In this article, We are discussing one of the key ingredients of PCR that is the dNTPs. We will also look into its structure and why it is so important.
Furthermore, we will also talk about the mechanism of interaction of dNTPs and Taq DNA polymerase and their working on the growing DNA strand.
Furthermore, we will talk about the mechanism of how dNTPs and DNA polymerase interacts and help to grow the DNA strand.
Read further on PCR related articles,
- Role of DMSO in PCR: DMSO a PCR enhancer
- Function of taq DNA polymerase in PCR
- PCR primer design guidelines
- Role of MgCl2 in PCR reaction
Key Topics:
What are dNTPs?
The dNTPs are the artificial nucleotides used in the PCR to synthesize new DNA strands much like DNA replication. dATP, dTTP, dGTP, dTTP are common nucleotides present in the PCR mastermix.
Structure of dNTPs:
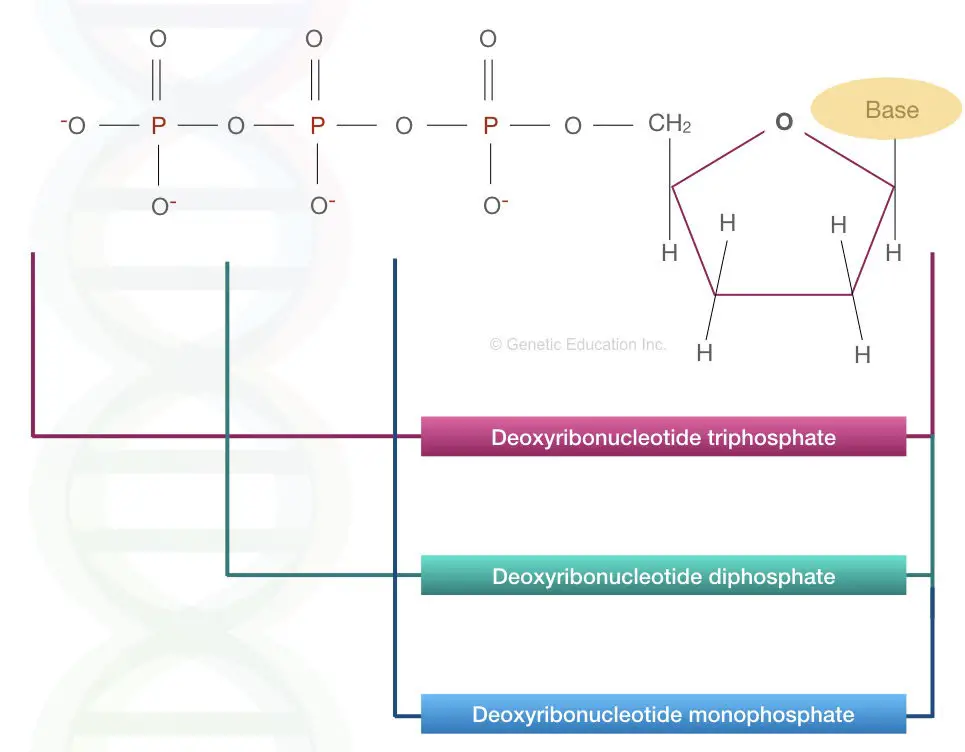
The dNTP stands for deoxyribose nucleotide triphosphate.
Firstly, we have to understand several basic terminologies such as base, nucleotides, nucleosides, ribonucleotides, deoxyribonucleotides, dideoxy ribonucleotides in order to understand dNTPs. These are some basic terms routinely used in genetics, still, people misunderstood it.
The Adenine and Guanine are Purine bases while Cytosine and Thymine are Pyrimidine bases. Nitrogenous bases can not form DNA directly. Phosphate backbone and pentose sugar are additionally required to create a DNA molecule.
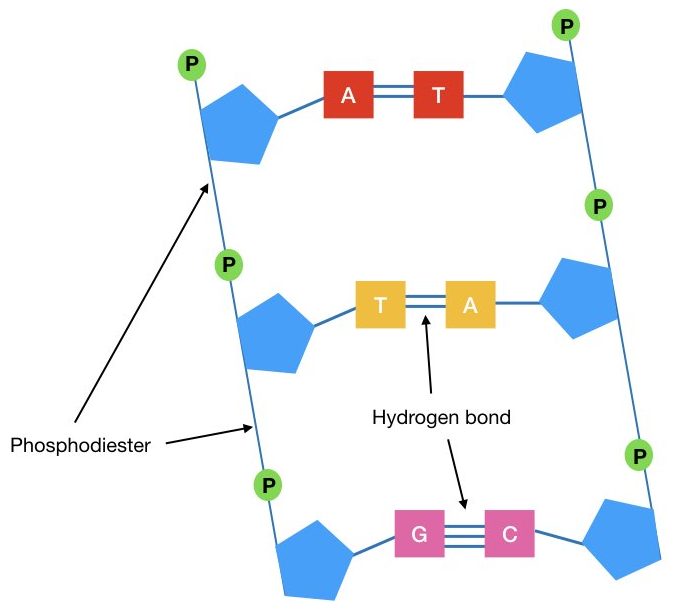
The nucleotide is made up of a pentose sugar, phosphate and nitrogenous base. One nucleotide binds with another adjacent nucleotide with a phosphodiester bond while one nucleotide binds with another nucleotide on the opposite DNA strand with hydrogen bonds.
Read Further on the Structure of DNA: DNA story: The structure and function of DNA
The phosphate present in nucleotide is triphosphate, (tri- three) when phosphate is not linked with the nucleotide (or absent), it is called a nucleoside (nucleotide without phosphate is termed as a nucleoside).
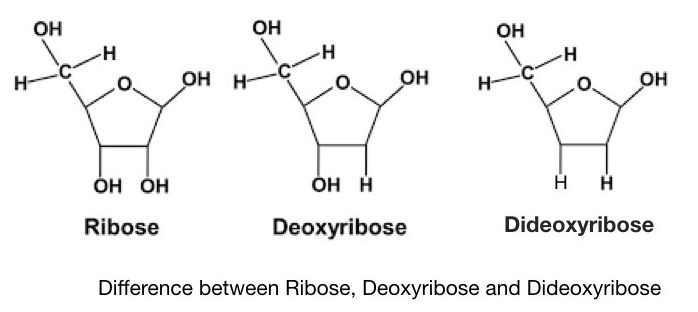
When a hydrogen atom replaces the hydroxyl group in the pentose sugar, the sugar is called deoxy sugar. The DNA is made up of deoxy pentose sugar while the RNA is made up of only ribose sugar.
The dNTPs are the chain of nucleotides made up of ribose, nitrogen base and phosphate.
Further, ddNTPs are different from dNTPs.
Dideoxynucleotide triphosphate does not have a free 3’ OH group, another 3′ OH group of sugar is replaced by hydrogen.
Hence it cannot cause the expansion of a growing DNA strand. So these types of ddATP, ddTTP, ddCTP and ddGTP are used in DNA sequencing. We will discuss the role of ddNTPs in sequencing elaborately in another article.
The ddNTPs are used in the chain termination method to stop the expansion of DNA synthesis.
The triphosphate is made up of three different molecules of phosphate which provides a backbone to the DNA.
The phosphate backbone of DNA has three different phosphate molecules called alpha, beta and gamma phosphate. When one phosphate or gamma phosphate is released, the structure is called deoxynucleotide diphosphate.
Similarly when two out the three phosphates are released the structure is called deoxynucleotide monophosphate.
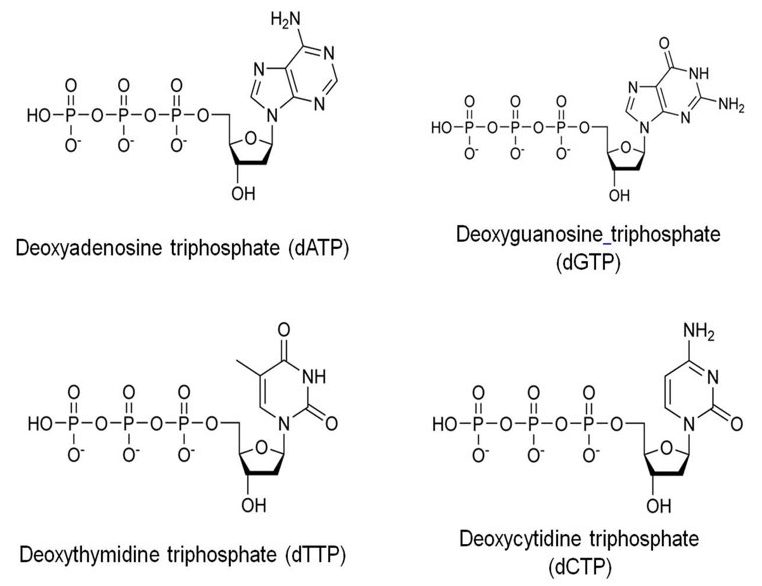
dATP and dGTP are purines while dCTP and dTTP are pyrimidine dNTPs used in PCR reaction. The function of dNTPs in PCR is the same as in vivo replication. If you are interested to read differences between nucleotides vs nucleosides and purines vs pyrimidines read these articles:
The function of dNTPs:
The dNTPs are the ingredients of PCR, RT-PCR, DNA sequencing or DNA microarray that helps to grow the DNA or amplification of DNA.
Mechanism of action:
The process of PCR is divided into three temperature-dependent steps:
- Denaturation
- Annealing
- Extension.
In the denaturation step, the double-stranded DNA is denatured into single-stranded DNA; In the annealing step, the primer binds at the exact position of where its complementary sequence is present and,
In the extension step, the Taq DNA polymerase adds the dNTPs at the growing DNA strand.
Once the strand is opened and the primer binds with single-stranded DNA, the Taq DNA polymerase starts its catalytic activity.
The Taq DNA polymerase uses the single-stranded DNA as a substrate for the enzymatic activity and settled on the primer-DNA junction.
The one end of the Taq DNA polymerase binds near the 3’ OH group of the oligonucleotide primer, this complex is called as P- DNA complex.
In the next step, the dNTP addition started.
The dNTP binds with the P-DNA complex with weak affinity if the exact complementary nucleotide is present. Soon after the Taq DNA polymerase holds it and the Hydrogen bond interaction between a complementary base and dNTP occurs.
Here, in the beginning, not the entire nucleotide but the base (nitrogenous base) present on the dNTP decides whether to bind or not.
Firstly, if it finds the complementary base on the template ssDNA (A for T and G for C), it will form the hydrogen bonds between them. Three hydrogen bonds between C and G and two hydrogen bonds between A and T are manufactured.
Once the hydrogen bond forms, the Taq DNA polymerase conforms that the binding of dNTP with the growing DNA strand by forming a phosphodiester bond. Now after the formation of the phosphodiester bond, the Taq DNA polymerase moves one step ahead for adding new dNTP.
The phosphodiester bond is formed between 3’ OH of primer and 5’ P of the dNTP.
After the hydrogen bond formation, the Taq DNA polymerase catalyzes the reaction by removing the gamma and beta-phosphate from the triphosphate of dNTP. Two pyrophosphates (PPi) are released after the completion of the reaction.
The exact kinetics of how the dNTPs and Taq polymerase interact during PCR reaction is still not known.
Read further on an agarose gel electrophoresis,
The concentration of dNTPs in PCR:
In general, for PCR reaction with 30 to 35 cycles of a run, 200μM of each dNTPs are sufficient.
200μM each which means a total of 800μM of 4 dNTPs mixes is utilized in a single PCR reaction. We have to prepare our working concentration from the stock solution of 100mM.
However, in recent days ready to use mastermix is very popular and effective. The ready-to-use mastermix contains all the important ingredients such as dNTPs mix, gel loading dye and Taq DNA polymerase.
As we are science students we have to learn, how to prepare the working solution from the stock. Suppose we have to prepare 2mM working from the 100mM of stock.
now,
Remember V1C1= V2C2?
Here, the given concentration C1 is 100mM (which is the stock concentration of dNTPs)
V1 is the volume given is unknown (?)
C2 is concentration required = 2mM
V2 is volume required= 1000μL
Now V1C1= V2C2
V1= V2C2/ C1
V1= 1000 * 2/ 100
V1= 20μL
Here, 20μL of each dNTPs is needed for the preparation of the working solution so the final volume for our mix is 80μL of (dATP, dCTP, dGTP and dTTP) in 920μL of D/W.
This will make the final concentration of 2mM of each dNTP in 1000μL of working solution. Calculate for 200μM by yourself.
My ultimate guide for using dNTPs in PCR
Always prepare two stock solution tubes and store all the tubes at -20°C.
Calculate how many reactions you perform in a week accordingly prepare the working solution from the stock and store it at 4°C. Because repeated freezing and thawing will decrease the activity of dNTPs.
Always wear glows and maintain sterile conditions while preparing the working solution, because contamination of foreign DNA will inhibit PCR reaction.
The 200μM concentration is enough for PCR reaction, however, for long-range PCR 2mM to 3 mM concentration of each dNTP can be used.
As the concentration of dNTPs increases the rate of non-specific binding will increase. Similarly, the scarcity of dNTPs leads to incomplete PCR products. So always use an appropriate amount of dNTPs.
Conclusion:
Conclusively, The function of dNTPs in PCR reaction is as important as Taq DNA polymerase.
With the help of Taq, dNTPs bind with the growing DNA strand and expand it. If you don’t want to mess with all this stuff, you can use ready to use PCR kit which is more preferable. Yet, you have to learn the manual method of reagent preparation.
Comment below and let me know if any point is missing from the explanation. Do share your views here.
Subscribe to our weekly newsletter for the latest blogs, articles and updates, and never miss the latest product or an exclusive offer.




Thank you for your efforts
I luv u
Lol. Thank You