“Extracting DNA from bacteria is a relatively easy technique due to their simple structure. Methods like CTAB, PCI and even ready-to-use kits give excellent results. However, to get good results some factors can be considered. Let’s take a look at the bacterial DNA isolation.”
DNA extraction from bacteria is a crucial scheme in microbiology, metagenomics, genetic engineering, infection studies and diagnosis. We can readily study the DNA of any bacteria which previously was a tedious task.
Conventional and traditional microbial culture, cultivation and microscopy can’t study DNA, take time and are prone to contamination and error. Conversely, utilizing DNA studies like PCR, DNA sequencing, 16s rRNA gene study and microarray bacteria can be studied effectively.
These techniques are more robust, fast, accurate and safe. To perform any of them we need pure and high-quality bacterial DNA. Bacterial DNA extraction is a relatively easy scheme but at the same time is complex and challenging.
Particularly, the diverse population of bacteria, varied genetic composition, and presence of cell wall, proteins and other components makes the present intervention even more difficult. No worries, We spent years working in genetic labs. Thus we have a substantial experience in DNA extraction.
In this article, we will learn how to extract DNA from bacteria. I will discuss various methods, protocols and common problems you will face. I will also provide one protocol for you to test.
Stay tuned.
Disclaimer: It’s priorly important to know that we are discussing the scheme for bacterial chromosomal or genomic DNA isolation, not for bacterial plasmid DNA isolation.
Key Topics:
Bacterial DNA Extraction
Bacteria are single-cell microorganisms living on earth even before us and dinosaurs. They are present, in nearly, every place on earth, even at some extreme conditions too and hence have a proven role in the food chain and earth’s overall ecosystem.
The single-celled bacterium is structurally very simple and contains its own genetic or chromosomal DNA, small circular plasmid DNA and other cellular organelles. Our target for isolation, here, is bacterial chromosomal DNA only.
It is believed that our own body has 10 times more bacteria than our own cells. They are present in our body, atmosphere, air, soil, water, and even on non-living things having different shapes. Where the majority of the bacterial population is harmless, a small group of bacteria is very dangerous.
Bacteria can be studied by traditional microbiology techniques, however, DNA studies give us more flexibility to explore the more uncovered areas in this field. The bacterial DNA isolation scheme is a complex process that highly depends on the type of bacteria.
Factors that affect DNA extraction from bacteria are the type of bacterial species, cell wall constituents of bacteria, the concentration of bacteria in the sample, age of bacterial culture and technical parameters such as technique selected, lysis method, purification process, etc.
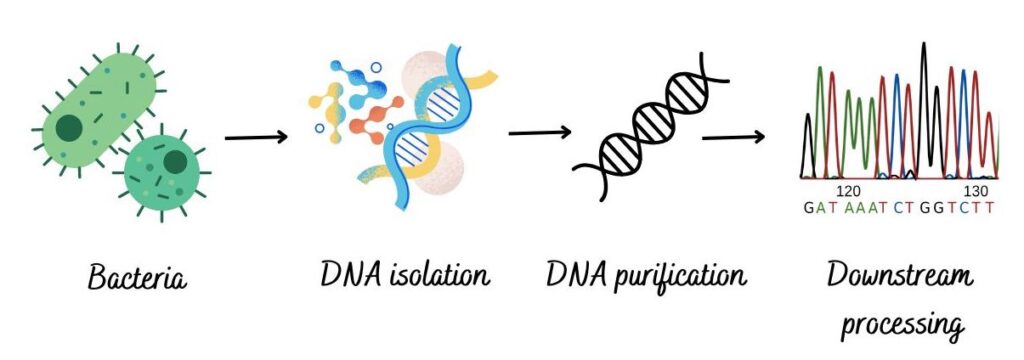
The general steps and process of bacterial DNA extraction are similar to the general DNA isolation process. The process is briefly explained here.
Culture or cell harvesting (if required)
Low-abundant bacteria having very low cell populations can be cultured prior to DNA isolation. In case, if it can’t be cultured, through harvesting a higher number of cells are obtained for isolation.
Note: To get optimum yield the bacteria should be harvested at a late log and early stationary phase.
Bacterial lysis
The bacterial cell wall is lysed using chemical, mechanism or enzymatic techniques depending on the requirement of the experiment. Usually, any protease along with lysis buffer gives effective results. Lysis breaks the cell wall and releases DNA.
DNA isolation
The DNA isolate is collected in a separate tube and precipitated using alcohol.
DNA purification
Alcohol purification is the most abundant way for DNA purification, however, kits can also be used to purify bacterial DNA for sensitive experiments like DNA sequencing. For alcohol purification, ethanol wash is given 2 to 3 times.
DNA elution
DNA can be eluted in an elution buffer, TE buffer or dd/W.
Bacterial DNA extraction protocols:
In this section, we will discuss some of the most common schemes that scientists use for the present purpose.
Quickest DNA isolation method:
The very first method is simple, fast and highly reliable for college-level research and demonstrating the DNA extraction technique. It uses only a couple of chemicals, one incubation and a protease enzyme.
- Add 1.5 ml bacterial cell culture to the 1.5 ml tube and centrifuge to settle bacterial pellets.
- Now add a lysis buffer to the pellet, mix well and incubate at 80°C for 5 to 10 minutes.
- Cool at room temperature and add protease enzyme to the solution and incubate for a few minutes at 60 to 65°C.
- Centrifuge the sample and collect the supernatant in a new tube and remove pelleted cell debris.
- Add chilled isopropanol to the supernatant and precipitate it by inverting the tube. You can also add a pinch of salt to maximize the yield.
- Once the pellet appears, centrifuge it, settle it and remove the alcohol.
- Wash pellets with alcohol 2 times, air-dry them and re-suspend it in TE buffer or dd/w.
Note: Remember that the present method can only work effectively for gram-negative bacteria. Additional steps and chemicals are required to break the hard cell wall of gram-positive bacteria.
EtNa bacterial DNA extraction protocol:
Vintgataramin & Frost, 2018 published an optimized protocol for bacterial and yeast DNA isolation using Ethanol and NaOH. The present protocol is a combination of EtNa and spin-column purification by Qiagen kit.
- Take 100 µL of the sample in a sterile 1.5ml tube and add 455 µL EtNa solution to it.
- EtNa solution preparation: 24mM NaOH, 2.7mM EDTA and 74% ethanol.
- Heat the sample at 80°C for 10 minutes and centrifuge it to settle the pellets.
- Carefully transfer the solution to a spin column.
- Now follow the QIAamp mini spin column protocol.
The present protocol is very quick, and safe and doesn’t contain any hazardous chemicals, digestion enzymes or tedious mechanical lysis steps. It can effectively work for both gram-positive and gram-negative bacteria.
CTAB Bacterial DNA isolation protocol:
CTAB buffer is a very popular solution in various DNA isolation schemes. Numerous protocols and modifications using CTAB are available for various types of bacteria and other organisms, however, CTAB with PCI is the best combination for achieving excellent outcomes.
Three major constituents of CTAB buffer are CTAB, beta-mercaptoethanol and EDTA, having the function of cell wall lysis (work as a detergent), protein degradation and DNA stabilization, respectively. When phenol and chloroform are added to this scheme, the overall performance would increase. Take a look at the general protocol.
- Add bacterial culture in a 2 ml centrifuge tube. Centrifuge the sample and settle pellets in the bottom. Remove supernatant.
CTAB buffer for 100 ml: 100mM Tris HCl, 25mM EDTA, 1.5M NaCl and 2% CTAB. Make up the final volume with double distilled water.
- Now resuspend the pellets in around 400 µL TE buffer and 100 µL (5M) NaCl, mixed by vortexing.
- Add 50 µL CTAB buffer, mix the sample well and incubate in the dry bath for 60 minutes at 70°C. Mix the sample occasionally during the incubation.
- Add 500 µL chloroform, mix by vortexing and incubate on ice for 30 minutes.
- Centrifuge the sample at a low speed for 10 to 15 minutes at 4°C temperature.
- Collect the supernatant in another sterile tube and add 500 µL phenol: chloroform to the solution and mix well. Use vortexing for mixing.
- Now centrifuge the sample at high speed and collect the upper phase.
- Now precipitate the DNA using alcohol and salt and wash it with the alcohol. Read this article for a step-by-step guide (Role of alcohol in DNA extraction).
Disclaimer: the protocol and chemical composition may vary among protocols, some minor adjustments and optimizations are advisable.
PCI Bacterial DNA extraction protocol:
PCI is yet another excellent protocol for getting high yield and high pure bacterial DNA. Phenol, chloroform and isoamyl alcohol are three major ingredients of the present protocol. The final nucleic acid exact is precipitated using the alcohol method.
The present solution along with a standard lysis buffer is capable enough to break the cell wall, digest proteins, isolate DNA and remove impurities. As I have mentioned the PCI method several times in various articles, so will not discuss it here. You can read the article and take a look at the protocol here: Phenol: chloroform and isoamyl alcohol DNA extraction method.
Bacterial DNA isolation using bead beating:
Bead beating is an excellent scheme for metagenomic DNA isolation containing bacteria. Various-sized beads, depending upon the type of cell wall bacteria have, are used for homogenization. Beads break the cell wall and release nucleic acid.
Further steps like purification and elution can be done using the conventional alcohol method or spin-column kit. The present method gives excellent results in comparison to any chemical-chemical isolation method.
If you want to learn more about what beads are and how the technique works, we have covered an article on it. You can read it here: DNA extraction using bead beating.
Ready-to-use kits:
Isolating bacterial DNA is a highly sensitive technique. Ready-to-use kits are also available for the present objective. For instance, Qiagen spin-column solid phase bacterial DNA isolation kits are valuable ‘add-ons’ to obtain excellent results.
Such kits having greater precision, accuracy, speed and performance becomes optimum for getting DNA from crucial testing samples having low-volume bacteria.
Lysis by heating protocol:
The simpler protocol for bacterial DNA isolation is heating. It disrupts the cell wall and readily releases DNA into the sample. It completes in a few steps. Add any good lysis buffer to the sample and heat it for ~30 minutes at 65 to 70°C temperature.
You can directly use the suspension for PCR. Else, follow the alcohol precipitation or spin-column purification to obtain good yield and purity. The present technique is widely used for tuberculosis DNA isolation and PCR.
Factor affecting bacterial DNA extraction
High yield and purity is our prime requirement for downstream applications. There are several factors that directly affect both the parameters in any bacterial DNA isolation scheme. Here I am listing some.
Bacterial species: Cell structure and composition vary from bacterial species to species. Gram-negative bacterial species have smoother cell walls which give more flexibility during DNA isolation, while the hard cell wall of gram-positive bacteria makes the entire process more difficult.
In addition, some species also contain secondary metabolites and byproducts having a direct impact on DNA isolation.
Cell wall structure: Gram-positive bacteria usually comprise thick and hard peptidoglycan cell walls. Such a rigid structure creates challenges in DNA isolation. Additional setup and lysis strategies are required.
Sample size: The sample size is yet another considerable factor here. Usually, bacteria can be isolated from their host and hence may have a low cell population. An insufficient sample results in low yield while an overloaded sample may negatively impact the overall extraction outcomes.
Extraction protocol: The purity and quantity of the DNA isolate highly depend on which protocol, lysis solution, purification scheme and chemicals are used. For instance, one protocol can work for gram-negative bacteria but the same can’t work for gram-positive bacteria.
The use of CTAB and PCI in the module provides high yield while using spin-column increases the purity. So the overall success depends on the extraction protocol selected.
Besides, other factors are bacterial culture conditions, growth phase, age of culture, cell density, type of host sample, buffer composition, presence of contaminants and overall handling.
Wrapping up:
In conclusion, DNA isolation from bacteria is an important process in genetic, microbiology and medical research as well as in infection studies. There are plenty of optimizations and protocols available but the choice highly depends on the type of bacteria present in the sample.
In this article, I have covered some popular methods, but not all. I know I haven’t covered the enzymatic method for lysis. An additional protease lysis step can be added to improve the overall quality of the extraction but isn’t necessary.
I hope you like this article. Please share it and bookmark it. Download our ebook on DNA isolation to PCR or get our highly impactful bacterial DNA isolation protocol.