“Low quantity, fragment heterogeneity, false results and analytical difficulties are some common limitations of using ctDNA as a biomarker.”
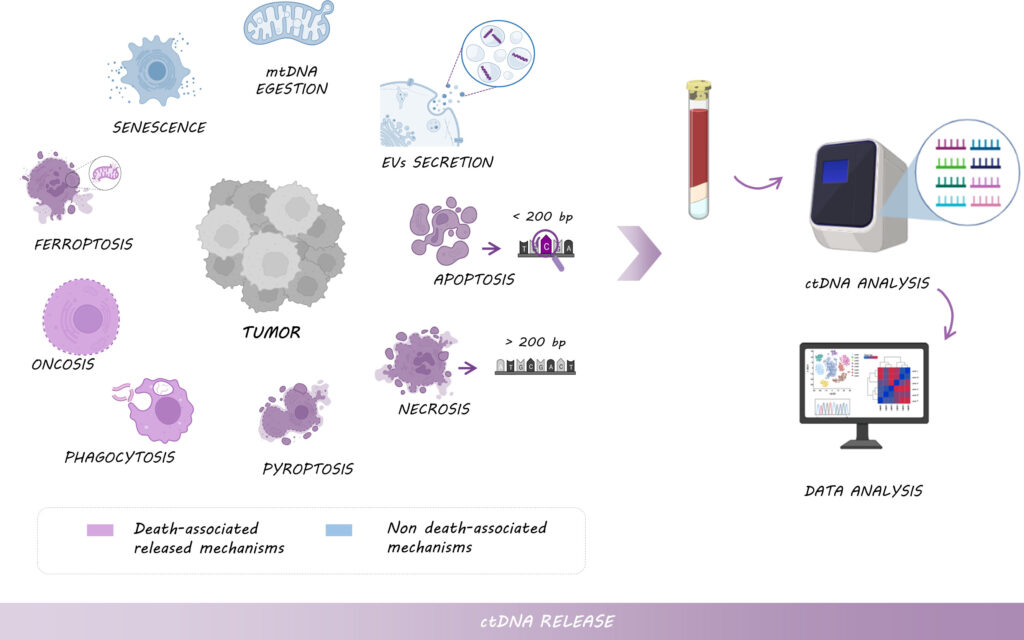
Circulating tumor DNA is free fragments of DNA released by tumor cells in the bloodstream or nearby tissues.
ctDNA acronym for cell-free tumor DNA is an amazing and futuristic genetic analysis technique. It’s minimally invasive, easy to perform and sensitive enough to test tumor DNA and thus is also an ideal biomarker too.
However, it has several serious shortcomings which limit its use as a clinical tool. Let’s discuss some of the limitations of the present biomarker.
Key Topics:
Limitations of ctDNA
Low quantity, fragment heterogeneity, low stability, differential sensitivity, limited copies, lack of universal set up and chances of false results are several common limitations of ctDNA.
Low quantity
The major limitation of the present technique is the quantity. ctDNAs are secreted in a low quantity by tumor cells. Studies suggest that the average quantity of ctDNA is between 10 to 100ng, which is comparatively very low.
The low strength of ctDNA causes two critical analytical errors or false results. First, false positive results and second false negative results. Two common factors for a false-negative result are low DNA and the location of the tumor.
Heterogeneity in fragments
The first thing to understand about the present marker is that the tumor cell destruction process produces random fragments. Meaning, we will get different numbers, sizes and types of fragments from different samples.
For example, from one sample we will get the TP53 and BRCA1 gene fragments while in another sample, we might not get them. The fragment size usually ranges from a hundred to a few thousand base pairs.
The stability
Due to the short half-life of up to 2 hrs, the ctDNA sample is less stable. This forces us to do rapid processing which leads to assay failures and inaccuracy in results. Once the ctDNA starts releasing, the cell’s natural clearance mechanism starts clearing it immediately.
Evidently, these data are for blood-derived ctDNA, studies demonstrate that ctDNA derived from other sources, for example, plasma has a comparatively higher half-life and can be stored for up to 48 hours using stabilizers.
Differential sensitivity
Blood is the major source of ctDNA, however, it can be isolated from other tissues as well. Studies suggest that the sensitivity of ctDNA testing also varies among sample types which limit the analytical efficiency of the test.
More research has been done on blood biopsy as it’s widely available, easy to isolate and majorly, minimally invasive. It’s proved more sensitive. However, at the same time, the yield of ctDNA from blood is also very low.
Lack of universal setup
No universal setup, SOP or working process is available for testing the ctDNA, however, for isolating ctDNA from blood, quite a few well-established SOPs are there. It is also important to understand that there are many challenges one has to face to prepare a standard protocol or SOP for ctDNA testing.
Methodological limitation
ctDNA could be the future of tumor testing. In addition, recent data also shows excellent scope from a clinical perspective, however, the overall sensitivity and accuracy of the ctDNA testing are very less as compared to other testing methods.
Limited mutant copies
A low copy number is a major challenge in this scenario. As cells shed off less tumor DNA, copy numbers are also less for testing. For even a highly sensitive digital droplet technology, at least 3 molecules of a mutant allele are required to indicate results.
Limited standards
As the technique is just emerging and yet growing, no standards, cut-off values or other testing parameters are available as a reference to evaluate the results.
Other factors
Technically, the process of ctDNA isolation is difficult.
It’s more prone to contamination as ctDNA is usually contaminated with normal cells’ DNA.
Blood-based liquid biopsy isn’t available for all types of cancer. For example, due to the blood-brain barrier, less ctDNA from brain tissues can be shed off in the blood which limits its use for testing brain cancer.
Read more:
Wrapping up
For now, ctDNA-based techniques aren’t well-penetrated clinically. However, several laboratories use such techniques routinely to test various tumors. Majorities of assays are based on the NGS platform which is time-consuming, costly and laborious.
Despite such limitations, scientists are doing research on establishing cell-free tumor DNA as a biomarker for various cancers due to its immense applications.