“A marker gene is a known DNA sequence located nearer to the target gene with detectable and inherited properties.”
Marker genes are applicable in the genetic engineering experiments, especially in the plant genetic research, often. Structurally, the marker gene is known as DNA sequence either gene, non-gene or DNA nearer to a gene used to locate the target gene.
The target gene is a DNA whose function we wish to study hence it is unknown for us. The marker gene works as a landmark to study the effect of some genes, stated by the National Human Genome Research Institute.
The antibiotic-resistant gene- a type of selectable Marker gene is one of the most common examples of the marker gene used in the genetic transformation experiments.
The techniques of genetic engineering are used to alter the genome of the target organism to achieve or change the desirable phenotype.
It is routine practice in the plant genetic engineering to manipulate their genome to achieve desirable characters or phenotypes which is important to us.
By using the marker gene sequence, the non-altered cell population and layered cell population can be distinguished.
In the present article, we will understand the concept of the marker gene and how it is useful in the genetic engineering experiments.
Related article: What is a Gene?- Definition, Structure And Function.
Key Topics:
What is a marker gene?
DNA sequencing located on chromosomes near to the target which is detectable is known as the marker gene.
There are two types of marker genes; selectable marker genes and Receptor genes.
Selectable marker genes:
The selectable marker gene is used in the plant genetic engineering and construction of genetically modified organisms. For example the antibiotic-resistant genes such as the streptomycin-resistant gene, Hygromycin resistant gene, Gentamicin resistant gene and ampicillin-resistant gene.
The function of the selectable marker is known to us, which means the use of these types of markers is to achieve a subsidiary secondary function along with the target gene.
While experimenting on the antibiotic resistance cells, when a marker gene of antibiotic resistance is inserted into the modified cell type, it may grow under the antibiotic rich media by the not-altered cells can’t.
Modified and unmodified cells are distinguished and used further.
Screenable markers:
The screenable markers are gene sequences used to screen cell population either altered or not. The altered cells having the screenable marker gives a signal that the target is inserted correctly.
For instance the green fluorescence protein-coding gene sequence.
A gene for a green fluorescence protein is isolated and instead into the plasmid along with the gene of interest and inserted into the target cells.
If the plasmid incorporated the genes correctly, the cells may glow green under the fluorescence microscope, which means cells are transformed.
Blue-white screening, GUS assay, opine synthesis and luciferase are several examples of the screenable markers.
Properties of marker gene:
- To select some gene as a marker gene, it should have several unique properties in it.
- It must be located near to the target gene
- It is also inherited along with the target gene
- It is amplifiable and can be detected easily
- The marker gene must be located on the same chromosome as the target gene
- It must be distinguishable
Use of selectable marker in genetic engineering:
In the present section, we will discuss how the selectable marker is used in the genetic engineering experiments to distinguish cell populations.
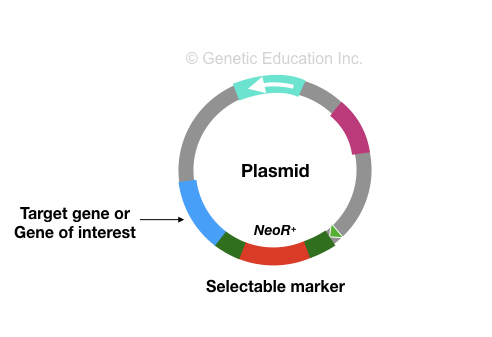
The target gene, plasmid DNA and technique selected for the experiment are three key elements of any genetic engineering experiment.
The target gene is a gene whose function we wish to study or willing to alter its function. Usually, this is a known gene.
The plasmid is an extrachromosomal DNA of bacteria that can transfer gene sequences we wish to study. It is a kind of vehicle that transfers genetic material.
Using an appropriate technique, a gene of interest can be inserted into the target cell population. But how can we distinguish the altered cells and non-altered cells?
Here the selectable marker is used!
The antibiotic that reduces the activity of the target cell is selected and used in the cell culture media. Which means when the normal cell population or the normal target cells are cultured in the antibiotic rich media, it can’t grow!
When an antibiotic-resistant gene sequence is inserted with the gene of interest into the plasmid.
The plasmid transfers both the gene and makes a protein. The protein that stops the activity of antibiotics also translates that naturally helps cells to grow.
Conclusively, if cells grow, it means our antibiotic gene is working properly and so our target gene is! That altered cell population is selected for further experiments.
Notably, the concept of the genetic marker and marker gene is different, don’t get confused!
The genetic markers are known DNA sequences, though are not marker genes. The genetic markers are usually non-coding DNA sequences like the RAPD, RFLP, AFLP or SNP.
It is just a known location for some unique sequences!
On the other side the marker gene can be a coding sequence, as we discussed; the antibiotic-resistant gene and green fluorescence protein-coding gene.
It is employed to confirm transformation experiments.
Notably, some marker genes are also used in the polymerase chain reaction like experiments to validate the experiment or assay.
It works as an internal control to know whether a gene sequence is correctly amplified or not!
Conclusion:
The “marker gene” is popularly used in the genetic engineering experiments, indeed the correct explanation lacks. While using it in the gene alteration experiments make sure to locate it exactly near to the target gene so that the activity and the expression of the target gene can be encountered correctly.