“Transcription Start Site abbreviated as TSS is the first nucleotide from which the transcription is initiated. Notedly, TSS is different than the promoter region. The present article explains the transcription start site.”
Transcription is a part of our cells’ central dogma. It forms an mRNA transcript which is a readable form of the genetic code, to synthesize a protein. Technically, it’s a complex process and so yet not completely understood.
I’m planning a series for transcription, so I’m reading randomly about it on the internet and came to know about the Transcription start site. At first, I understand that it’s a site from where transcription initiates.
But as I surf more, I came to know that TSS is crucial and there’s less data available, even for me it was hard to find more relevant sources. The concept of TSS would help us to learn more about how genes form transcripts more precisely.
So in this article, I will explain what the transcription start site is, why it is important and how to find it. I will also explain why one has to learn about TSS. Present content would add more value to your gene expression learning.
Related article: “Transcription And Translation” A Brief Overview.
Stay tuned.
Key Topics:
What is a Transcription Start Site?
TSS is a location from where the process of transcription begins. More precisely, the first nucleotide which starts transcribing into mRNA is a transcription start site.
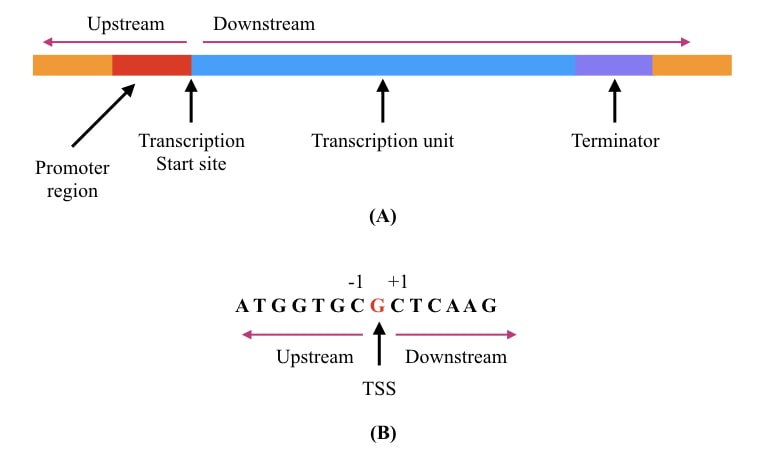
The region before TSS is known as the upstream region while the region after TSS is known as the downstream. Henceforth, the coding region that is up for transcription is located in the downstream region while the regulatory elements are located upstream.
Technically, the nucleotides located upstream are denoted with negative numbers while the nucleotides located downstream are denoted as positive numbers. So the first nucleotide that starts transcription, is labeled as +1.
Note that some genes may also contain more than one transcription site meaning, for such genes, transcription occurs at many places. On the transcript side, in the mRNA, TSS becomes a site for mRNA capping. Meaning, once the mRNA forms, the 5’ capping occurs at the site which is previously TSS.
What is the importance of a TSS?
Less available data is the major limitation of the present topic. However, after reading tons of literature, I came to know that the TSS is very crucial in order to study and understand gene structure and regulation of gene expression.
In addition, by locating the TSS in a gene, scientists can identify the promoter and regulatory element region, understand the structure of the 5’ end and determine the RNA polymerase binding site and capacity.
So TSS determination is crucial for knowing a gene, structurally and functionally.
How to determine TSS?
Three experimental methods for determining the TSS for a gene are Primer extension, nuclease mapping and RACE (Rapid Amplification of cDNA ends), among which RACE is the most accurate and popular.
RACE (Rapid Amplification of cDNA ends):
RACE is a bit complicated and highly sensitive technique that needs an expert hand for evaluation. Here is a broad overview of the technique.
First, the mRNA is isolated and allowed for cDNA synthesis using the standard cDNA synthesis protocol.
The cDNA is purified using the RNase enzyme which cleaves off RNA and provides pure cDNA.
Poly C tail is added to the 3’ end of the cDNA using the TdT- terminal deoxynucleotidyl transferase enzyme.
Two types of primers one specific for the 3’ poly C tail and another specific for a gene region are allowed for amplification. Another set of nested primers is also used for amplification of the final product.
The product is then sequenced using the RACE-seq protocol. Ideally, the first nucleotide after the G, from the 5’ end is our TSS. Olivarius et al., 2009 mapped the TSS of 17 genes from a single cell line using RACE-seq.
This approach is, however, tedious. We are not discussing the other two approaches as are not used often.
Read more: What is a Palindromic DNA Sequence?
Wrapping up:
A transcription start site can also be predicted computationally using bioinformatic tools. I will try it later and update this article. Right now I have this much information regarding this topic.
I will read more, discuss it with other fellow researchers and add more value to this topic. Although the present content is enough to understand the topic.
FAQs:
Sources:
Olivarius S, Plessy C, Carninci P. High-throughput verification of transcriptional starting sites by Deep-RACE. Biotechniques. 2009 Feb;46(2):130-2. doi: 10.2144/000113066. PMID: 19317658.