“The transposons are mobile genetic sequences that can move from one place to another place in a genome.”
“The transposons are the missing link in the path of evolution of multicellular organisms and therefore they are the crucial factor in the process of the overall evolution of life” biologist from the University of Georgia.
Key Topics:
“Evolution is a process by which different types of organisms suppose to originated from the primitive ones.”
The prime goal of any species on earth is to survive. For that, to fight with harsh conditions, various alleles are originated. The mutation is one of the forces involved in the evolution and creates alleles.
How the mutation occurs?
Well, there are so many reasons for that, some are intrinsic and some are extrinsic but transposons are one of them too. Since the discovery of the transposons in the 1950s, the mobile genetic elements are treated as a culprit for producing mutations.
The transposons are found in almost all organisms and species on the earth, either in the active form or inactive form.
The transposons can be divided into two broad categories:
- Class I transposons, a retrotransposon found in almost all eukaryotes.
- Class II transposons, DNA transposon found in prokaryotes.
RNA mediated transposition is performed by the retrotransposons. Through the reverse transcription, a newly formed DNA is inserted into the genome. on the other side, the DNA transposons can directly insert at the target site.
It is believed that the retrotransposons are derived from retroviruses or are retrovirus in the past.
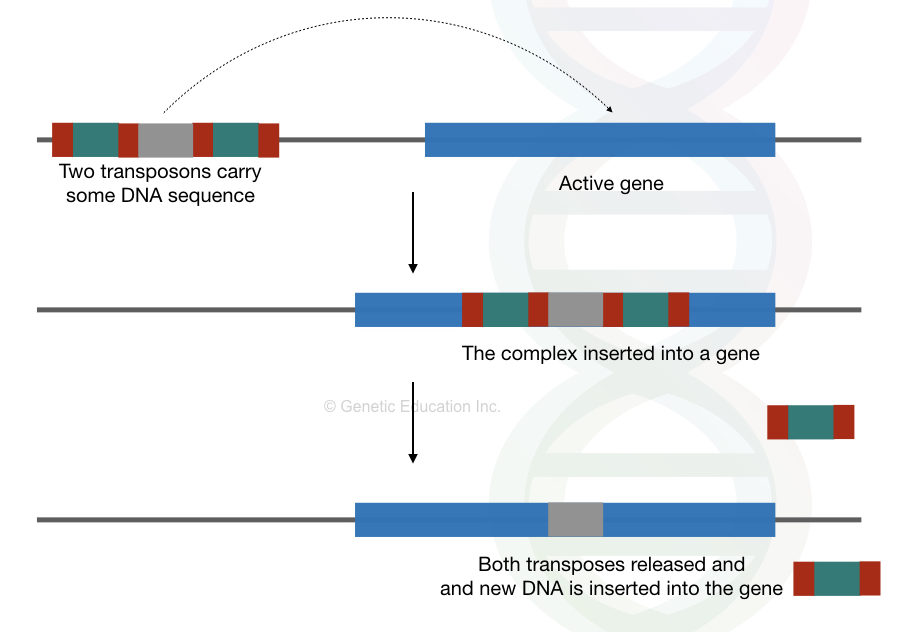
Transposons are known as selfish and parasitic peccant to the host genome. They jump from one location to another and creates new mutations and chromosomal breaks. See the image below first,
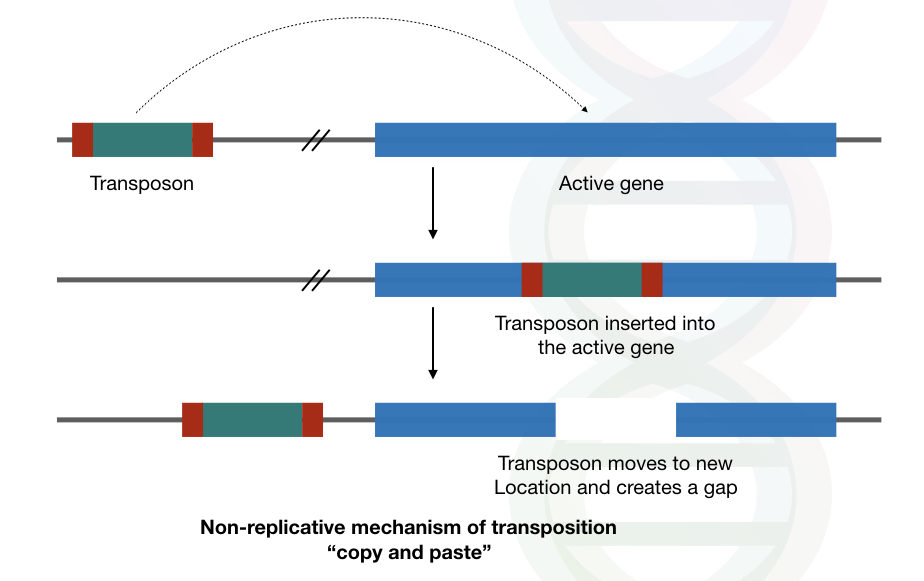
However, in modern science, the explanation is just a myth.
Simply put, the transposons can be considered as “agents” that are powerful enough to cause mutations or variations. Although, the mechanism of it is still unclear.
Now it is proved that some transposons are essential to survive for every organism. It had also involved in the genomic evolution of multicellular organisms.
Transposons are just a bunch of junk DNA without an apparent regulatory function in the genome, still, it can make proteins for themselves.
The transposons often referred to as transposable elements and are settled in the heterochromatic region of the eukaryotic genome as inactive sequences. The active transposons move across the genomes, between the genes, and produce new mutations.
In a more scientific language, “ their transposition into the protein-coding DNA sequences possibly diversify the transcriptional signals that result in loss of function or a faulty protein”.
The meaning of both sentences is the same.
In the present article, for understanding the role of the TEs in the evolution, we have to first understand how it disrupts the gene function?
Read further:
A gene…
A gene is a function DNA piece that makes protein or proteins. It is made up of introns and exons. Introns are non-coding intervening sequences while the exons make transcript and form proteins. By the exon splicing mechanism, the non-coding introns are removed, forming a pre-mRNA.
It transcribed into the mRNA and forms a protein by translating it.

The moving genetic elements disrupt the function of a gene in several ways.
If a mobile genetic element, a transposon inserted into one of the exons of the coding sequences, ultimately it mutates that exons. It can’t splice out and can’t form protein, consequently. This mechanism makes loss of function- proteins.
Scientists has now proved that the TEs are involved not only in the loss of function mutation but also creates new alteration (insertional mutation). Some of the coding region of transposons inserted directly in the intron and behaves like a fake exon,
That exon is counted for the protein formation and a totally new type of protein is formed from it.
In another way,
The “cut and paste” transposons create a gap or break in a DNA sequence, now, that gap is filled by some random nucleotides by the normal replication mechanism of an organism. Conclusively, it can block the function of a gene in these three different ways.
These explanations favor that transposons generate a new variation, a new allele, and viz a new protein
New phenotypes in nature are actually manufactured due to allelic variations and therefore allelic variation can be considered as a base for evolution.
Read this article on Gene vs Allele.
Let’s take an example,
It makes the point clear how transposons or transposable elements are involved in evolution.
How the antibiotic-resistant bacterial species are evolved?
The bacteria did not have an antibiotic gene before the discovery of it. The fact clearly indicates that the gene for antibiotic resistance has emerged. The antibiotic resistance gene is emerged due to the moving genetic elements present in their genome.
Once the allele happened, the transposons move it across different bacteria by introducing it in plasmids.
Even, the resistance gene can be transferred between different genera and species of bacteria. The transposition of E.coli R-plasmid is found in some of the genera such as Salmonella, Hemophilus, Shigella, Proteus and Pasturella.
The transposons result in the deleterious effects but, in fact, the actual function of it is to drive evolution, creating new alleles. Furthermore, transposon translocates DNA sequences within a genome, facilitates exon shuffling and repairs double-stranded breaks.
The Tol2 transposons of Fish not only jump but also alters the regulatory region of a gene and creates varieties of phenotype. The Tol2 Transposon is a DNA transposon, follows the “cut and paste” mechanism for transposition.
In the presence of the Tol2, the fish develops different types of pigmentation patterns, but once the transposon is removed, the fish remains albino. Here the Tol2 transposons are work as a pseudogene.
However, if the TOl2 remains in the regulatory region, it causes a variety of alleles for different types of pigmentation patterns (Koga et al., 2006). The Tol2 jumps within the regulatory regions and fabricates new variation in a Fish.
Now, the present example is the best explanation for how transposons are included in the process of evolution.
A regulatory inspector…
“The effect of transposons on a nearby gene is epigenetically regulated” B. McClintock. She designated them as elements, “Ac/Ds elements” remember!!
Why named it as “elements”?
She postulated that some of the mobile genetic sequences influence the epigenetic profile of the nearby gene and it could be attuned by environmental changes. Which means,
The gene expression is regulated by the transposons.
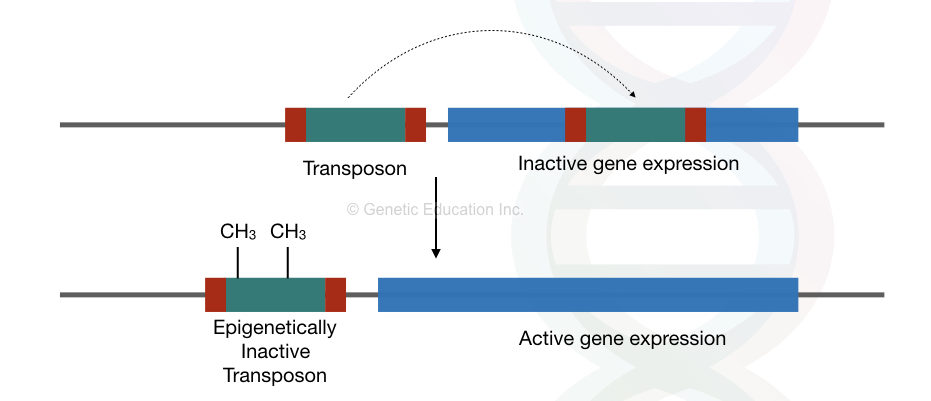
The moving elements move from one location to another location within a genome. If it jumps between an active gene, that gene is down-regulated and protein cannot be formed, thereby, interfering in protein formation it regulates gene expression.
The transposons are now epigenetically silent, situated in the heterochromatin region of chromosomes. Interestingly, it can on/off the gene expression of a nearby gene by playing with its epigenetic profile.
Even, the epigenetic profile of the transposons is also altering the gene expression. See the figure above,
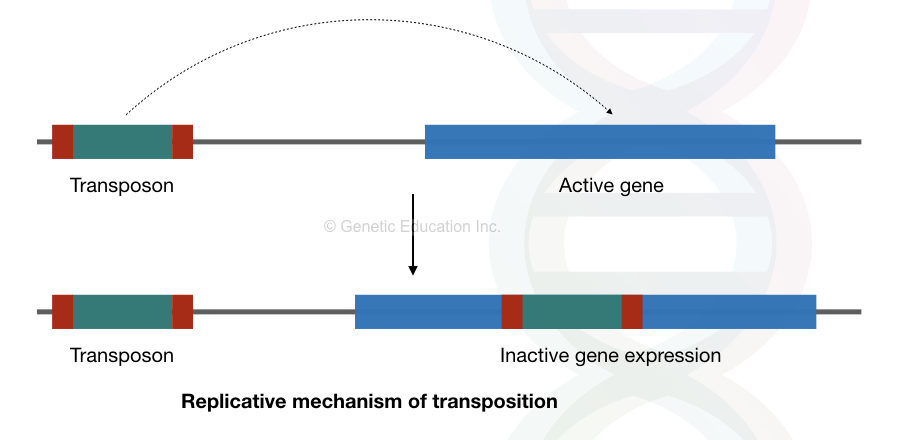
Let’s take another example,
The stress-responsive elements are the best example of the plausible role of transposons in the evolution of new characters.
For example, in A. thaliana, the Copia like ONSET retrotransposons are resistant to heat. It inserts into nearby genes due to heat-responsiveness.
The function of the nearby gene might disrupt but it creates a new allele that generates genetic diversity in the A. thaliana for the heat resistance.
Artificial genetic diversity and resistance species in plants can be generated by using the transposons.
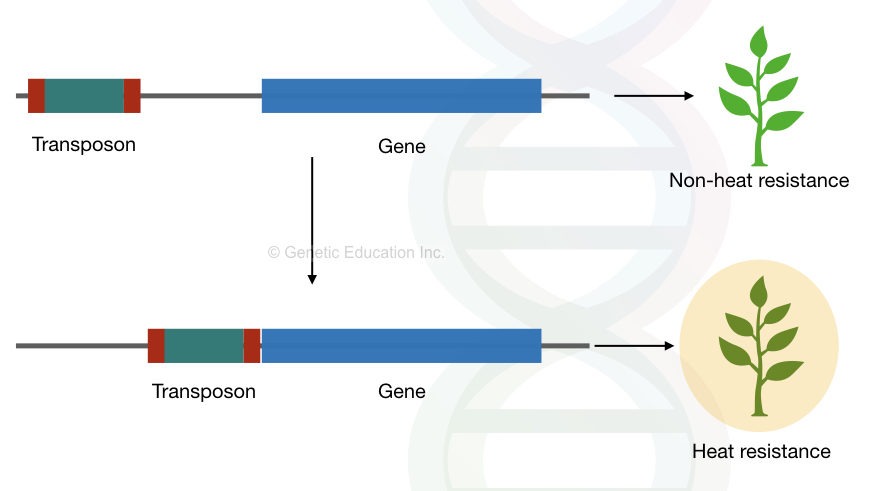
A DNA carrier…
Besides working as a regulatory element, the transposons even transfer DNA sequences from one place to another.
By creating a staggered break, the transposon inserted into a new location.
The break is then filled by the replication with the help of the DNA polymerase (target site duplication).
The duplicative sequences of the target site are generated on both sides of the transposon and moves along with it.
Read related articles:
A TSD inserted at a new location and ultimately a new variation is originated and ultimately a new variation and phenotype are produced.
So, the transposons itself perform five different functions:
- It can carry any of the DNA sequences
- It can create new mutations hence new variations
- It can regulate gene expressions
- Alter gene function through the insertion
- Can do chromosomal rearrangements
The genetic force that drives the evolution requires almost all these changes in the genome to drive new variation. So it is clearly evident that the transposons and transposition process responsible for the evolution of a multicellular organism.
The genome…
The major portion of the genome is made up of the repeated DNA sequences called junk DNA which can not code protein. A transposon is repeated DNA sequences positioned in the centromeric region of the genome and a major constituent of a heterochromatic region of the genome.
The heterochromatic region of the genome gives the structure to it and hence helps to arrange DNA on the chromosomes. And this is the reason why non-replicative transposons create nick or cut on the chromosome.
Related article: What is a genome?
Conclusion:
One important question arises in mind that,
Suppose, if the transposons do not have any significant role, why they are conserved and stored in the genome of an organism?
Although the majority of the transposon sequences are epigenetically silenced through methylation and other silencing mechanisms, their presence is indeed more important in the genome.
Sources:
- Feschotte C, Pritham EJ. DNA transposons and the evolution of eukaryotic genomes. Annu Rev Genet. 2007;41:331-368.
- Ayarpadikannan S, Kim HS. The impact of transposable elements in genome evolution and genetic instability and their implications in various diseases. Genomics Inform. 2014;12(3):98-104.
Subscribe to our weekly newsletter for the latest blogs, articles and updates, and never miss the latest product or an exclusive offer.



