“A genome is an organism’s entire single set of DNA which have all the information needed for an organism to grow”.
Or
“A genome of us is made up of DNA having all the information for survival. The entire set of DNA present in a cell is called a genome.”
More precisely,
A haploid set of the DNA present in a cell is called a genome.
The genome of us is made up of DNA- a basic building block of life and manufactured of nitrogenous bases, phosphate and sugar.
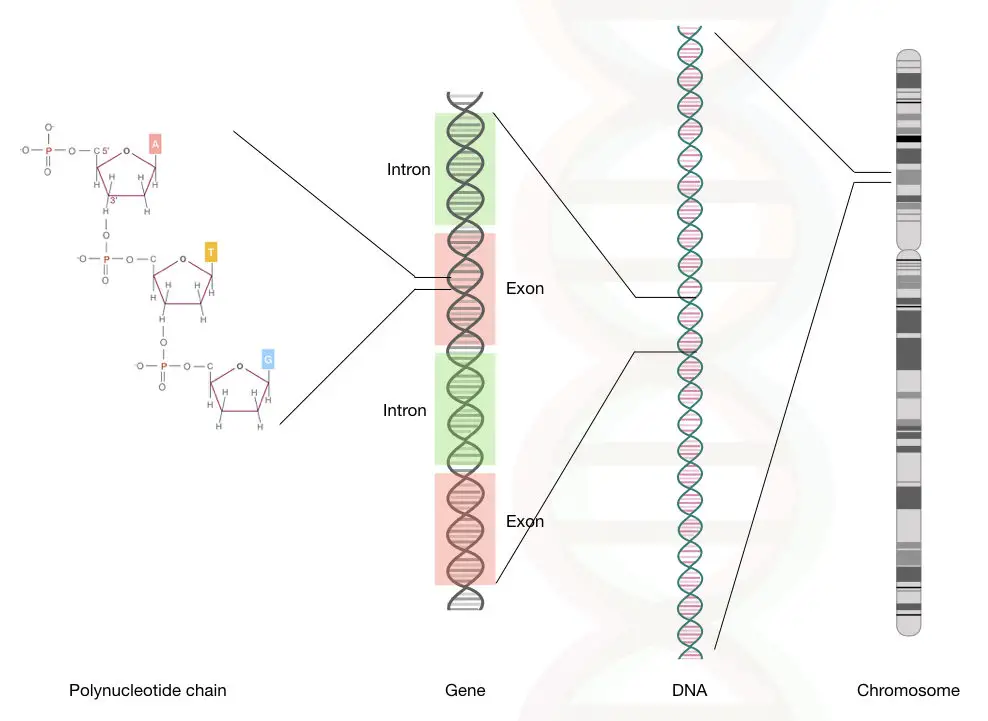
The DNA is located on the chromosomes- a complex network of DNA and proteins (mainly histones). 23 pairs of chromosomes are present in somatic cells with a pair of sex chromosomes.
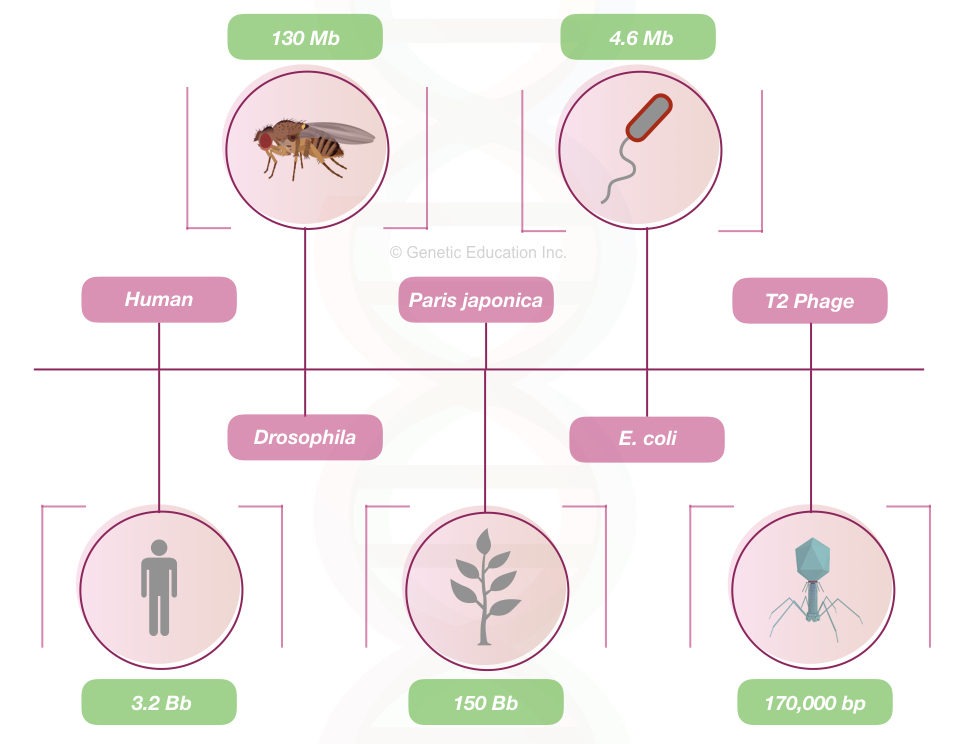
The human genome contains 3.2 billion base pairs. However, the size of the genome varies from species to species.
In the present article, we will cover all the information on the genome.
Topics Covered:
- What is a genome?
- Definition
- Structure
- Coding sequences of a genome
- Non-coding sequences of a genome
- Euchromatin and heterochromatin regions
- Function
- Genomics
- Conclusion
Key Topics:
What is a genome?
Total DNA in a cell is a genome.
Let’s understand it by taking an example. The genome is just like an entire cookbook having all the information on how to cook different recipes along with the list and information of different ingredients.
Each chapter in a cookbook dedicatedly contains information on one particular recipe and related ingredients. Similarly, the entire genome is just like a cookbook that contains all the information for coding and regulating gene expression.
Each gene from the entire genome contains information for encoding different proteins. Further, some non-coding sequences also possess information for gene regulation.
The eukaryotic organism is made up of billions of cells, a group of cells that makes tissue, tissues combinedly form organs for performing different functions, organs build the body.
All the information on how cells divide, how it forms, how it died and how it functions, are encoded in the DNA.
Different genes encode different proteins for performing different functions. For example, some genes regulate cell division while some genes control apoptosis- cell death.
Genes are functional of a piece of DNA because not all the DNA is capable of encoding proteins.
Read more on genes and DNA:
Table: Some organisms, their genome size and the number of chromosomes.
| Name of the species | Genome size in (Mb) | Number of chromosomes(n) |
| Human | ~3200 | 23 |
| Dog | 2500 | 39 |
| Mouse | 2600 | 20 |
| Rat | 2800 | 21 |
| Sheep | 3000 | 27 |
| cat | 3000 | 19 |
| baboon | 3100 | 21 |
| pig | 3000 | 19 |
| Cow | 3000 | 30 |
| Horse | 2700 | 32 |
By interacting with the histone proteins, categorized into- H1, H2A, H2B, H3 and H4, DNA coiled up into a structure called chromatid, four different chromatid forms a chromosome.
A single chromosome is a single part of an entire genome (entire 23 chromosomes) made up of arms, centromere and telomeres.
Important genes are located on two arms of a chromosome, denoted as “p arm” and “q arm” while the telomere has non-coding repeated sequences called satellite DNA.
The telomeric DNA sequences protect the gene coding segment of a chromosome from the end replication problem. The entire set of DNA- the genome is present within the membrane-bounded structure known as the nucleus.
However, some DNA is also present in a cytoplasm either chloroplast DNA/ mitochondrial DNA in eukaryotes or plasmid DNA in prokaryotes.
Those DNA molecules are circular or linear ins structure but scientists do not consider it has genomic DNA because of the presence of their own replication and transcription machinery.
Scientists had sequenced the entire genome of us in a projected named Human genome project- completed in 2003. In addition to this, the genome of different other organisms was also sequenced and compared with ours. The project was started by the U. S department of energy and the national institute of health.
The human genome contains 3.2 billion base pairs. It provides all the information to an organism required to function and survival. Further, it stores information and is inherited to new cells and offspring.
Approximately, 20,500 functional genes are identified and sequenced through the project.
From the HGP, scientists have constructed a database of the genetic map, Physical map, entire DNA sequence data, human sequencing variation map and other information of genes and their functions.
Fun fact:
If we print a total of 3.2 billion basepair letters on paper, 500 pages notebook can be filled with it.
If we make a chain of all the bases it covers approximately ~3,000km.
Definition:
“Entire DNA present in a cell of an organism is called a genome which inherited information from one generation to another and regulates gene expression.”
Or in simple language, we can say,
“A genome is an information storage and distribution unit having all the information on how the organism will grow and develop.”
Structure of genome:
Structurally, the genome can be divided into two broader categories:
- Coding DNA sequences
- Non-coding DNA sequence
Coding DNA sequences:
The genome of a prokaryote is flat, and contains only a few important genes and a less non-coding sequence. Genes are structurally also not similar to the eukaryotic genes as well.
On the other hand, the eukaryotic genome is different.
Unlike the prokaryotes, the major portion of the eukaryotic genome is made up of non-coding “so-called” junk DNA.
Moreover, the genes are made up of exons and introns.
The unmethylated DNA sequences which make protein have introns and exons are called coding DNA sequences or a gene.
Less portion of the eukaryotic genome can encode proteins, especially in humans, only 2 to 3% of the genome is functionally protein-coding sequences.
Although the ratio varies from organism to organism. number of genes in a different organism is given in the table below,
| Species | Name | Number of genes |
| Homo sapiens | Human | 21,000 |
| E. coli | Bacteria | 4,200 |
| Oryza sativa | Rice | 38,000 |
| Daphnia Pulex | Water flea | 31,000 |
| Gallus gallus | Chicken | 17,000 |
The coding region is a function DNA piece called a gene that has a couple of important characteristics.
First, the repetitive DNA portion in a gene is less or nearly zero because the chance of replication error is very high in the repeated DNA sequences.
The second most important property of a gene is the unmethylated DNA sequence. DNA methylation prohibits transcription as enzymes can not function, once the methyl group is added.
The third most important characteristic of a gene is the packaging of genes on chromosomes.
Well, non-coding and coding DNA sequences are even arranged differently on chromosomes.
The coding region “so-called” gene-rich region is more loosely packed as compared to non-coding DNA. Thus it allows enzymes to function properly.
Furthermore, the genes or the coding- DNA are located on the “arms” of chromosomes.
Another important characteristic of a gene is its structure- as it contains introns and exons. The introns are removed from the final transcript and exons combinedly form a protein product.
Importantly, the length of the gene is also one of the crucial factors for protein formation. The larger the gene more chance of replication error results in a mutation.
Thus most of the genes are shorter and contain fewer exons.
Though 20,000 to 21,000 genes are identified to date, many more transcribed sequence with unknown function exists in our genome.
Non-coding DNA sequence:
Now, this is very important to understand.
Although this portion of the genome can not form any protein, it is very important for our cells to survive.
Approximately 97 to 98% of the human genome is made up of non-coding, repetitive junk DNA.
Here I am going to enlist some of the important types of non-coding DNA sequences present in our genome in this segment of the article:
Repeated DNA sequence:
Undoubtedly, the major portion of the human genome is made up of the repeated DNA sequence located on the centromeric and telomeric region of chromosomes often known as the microsatellite or minisatellite.
Fun fact:
Human chromosomes are protected by the telomeres which have a repeat sequence of TTAGGG.
In addition to this, the length of the telomere is directly proportional to the age of the person. Longer the telomere, the longer the age.
Related article: Role of Telomeres in aging.
Tandem repeats:
The tandem repeats are located one after another in the genome, also of two types. Short tandem repeats and long tandem repeats.
The short tandem repeats are made up of the 1 to 6 nucleotide long units while the long tandem repeats are made up of 10 to 50 nucleotide long units.
The short tandem repeats are known as STR while the long tandem repeats are known are the VNTR- variable number of tandem repeats.
Both types of non-coding DNA sequences is a very important marker used in genomic and genetic studies.
STR or VNTR is one of the unique properties of a person, no two people in the world have the same VNTR or STR profile.
Thus this type of repeated DNA sequence is used in the verification of individuals, DNA fingerprinting and paternity verification.
We have covered an entire article on STR hence we are not discussing it in detail here.
Related article: Short Tandem Repeats (STRs): A Secret of Every DNA Test.
One important point we have to discuss the repeated DNA sequences is that, even though it can not code for any protein, it can cause some inherited genetic disorders too.
For example, Huntington’s disease occurs due to the abnormal expansion of the CAG repeat.
Transposable elements:
As the name suggests, the transposable elements often called transposons have the power to change location within a genome, although the DNA sequences are not mobile in nature.
“the transposable elements are the mobile genetic elements- non-coding DNA that moves from one location to another.”
We had covered an entire series of articles on transposons, therefore we are not discussing it in depth here. You can read all the articles here:
Category: Transposons
The transposons are either retrotransposons or DNA transposons, the retrotransposons are transcribed into the RNA and move to another location.
Based on the terminal repeats present on both sides of the retrotransposons, it is divided into long terminal repeats (LINEs) or short interspersed elements (SINEs). The majority of the transposons present in us are retrotransposon types.
Other types of transposons that are found in bacteria and other prokaryotes are DNA transposons.
Terminal inverted repeats are the main characteristic of the transposons, the DNA transposons encode enzyme transposase which recognizes the terminal sequences for the mechanism of transposition.
In addition to this, the transposons are also categorized based on the mechanism of transposition, either cut and paste transposons or copy and paste transposons.
We have covered a dedicated article on each type of transposon. Read in the category given above.
The transposons play an important role in creating new variations in nature and thus it is believed that, in past, transposons are one of the forces which drive evolution.
Besides this, once the transposons move to a new location, in between the gene, it dysregulates or inhibits the function of a gene or fuses two genes and creates a new genotype.
This is the reason, transposons like non-coding DNA sequences are a very important part of our genome. But!!
Transposons are inactive in eukaryotes before thousands of yours.
Methylated DNA sequences:
Yet another type of non-coding DNA sequence are the methylated DNA sequences in our genome.
A methyl group binds to the CpG-rich region of the genome and deactivates the gene function or makes the DNA sequence non-functional.
The tightly packed heterochromatin region of the genome is a methylated-rich region, mostly inactive.
Smaller RNAs:
Smaller double-stranded or single-stranded RNA molecules play an important role in the regulation of gene expression.
Called microRNAs, encoded by some of the genes but can not able to form a protein thus this type of DNA even though able to transcribe, is categorized in the non-coding type of DNA sequences.
Those microRNAs play a crucial role in the RNA interference mechanism in which it destroys some of the pathogenic viral RNAs other endogenous mRNAs and regulates gene expression.
Again, we have covered an entire series on different types of RNA and microRNAs. You can read more about it here: microRNA (miRNA) and Gene Regulation.
Besides this, promoters, enhancers, suppressors, insulators, locus control regions, core and proximal promoter sequences are also categorized into the non-coding DNA sequences as well.
Summary of the non-coding DNA:
| Type of non-coding DNA | Description |
| Satellite DNA | Microsatellite and minisatellites are located on the centromere and telomeric region of the chromosome. |
| Transposons | Mobile genetic elements are located throughout the entire genome and can move from one location to another. |
| Introns | Introns are present in a gene within the exons and are removed prior to mRNA formation. |
| Telomeres | The telomeric end of the chromosomes are made up of the six nucleotide repeat sequence that protects the chromosome from the end replication problem. |
| microRNAs | microRNAs are transcribed from the DNA but can not form protein and therefore it is categorized into non-coding DNA. |
| Regulator elements | Other regulatory elements help in replication and transcription. |
Euchromatin and heterochromatin regions:
Euchromatin regions are lightly packed DNA sequences mostly, gene-rich regions while the heterochromatin regions are tightly packed non-coding DNA sequences.
During the cytological analysis, when we perform GTG banding, different bands of light and dark pinkish-blue regions are observed. The darker portions or bands are the heterochromatin, tightly packed regions. while the light-colored regions are euchromatin regions. See the image below,
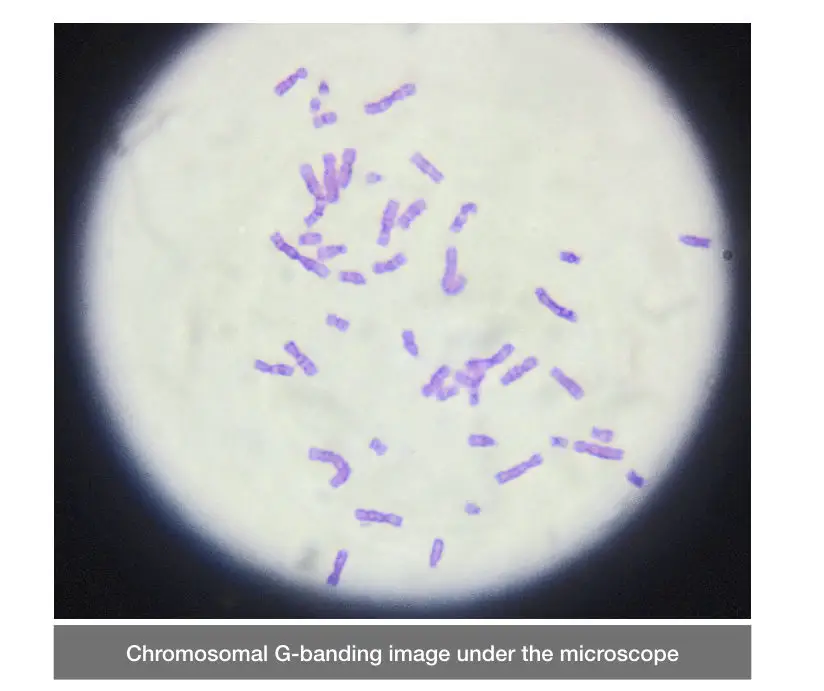
Related article: Euchromatin vs Heterochromatin: Differences and Similarities.
Functions of a genome:
The genome is just like the hard disk of our computer.
A computer hard disk stores all the data and crucial information for the computer on how it works and how to perform different functions for a computer.
Similarly, the genome stores all the information for an organism- for its growth, metabolism, development, reproduction, etc.
The computer runs on the binary language of 0 and 1, the genome stores information on A, T, G, and C nitrogenous bases and inherited in the same manner.
Genes within the genome encode different proteins for different cells to function properly while the non-coding sequences such as microRNAs help to regulate the gene expression.
Based on that, the genome can be divided into three categories:
- Coding gene sequences
- Regulatory elements
- Maintenance elements
Coding elements or coding genes are unmethylated loosely packed DNA sequences that encode a protein.
Regulatory elements regulate gene expression and in which amount the protein forms. Enhancers, promoters, suppressors and insulators are some of the regulatory elements found in the genome.
Maintenance elements are the sequences that help is DNA repair and maintain it. Origin of replication, telomeres and centromeres of a chromosome are some of the examples of maintenance elements.
How the genome function?
Our cells developed different pathways for doing different processes related to DNA.
Through replication, the DNA becomes doubled and inherited to the daughter cells. An exact copy of the entire genome is stored in another cell nucleus.
Through the process of transcription, mRNA is formed from the DNA. The mRNA has all the coding information, the entire mRNA set of the genome is called transcriptomes.
By analyzing the transcriptomes or entire set of mRNA, the amount of total gene expression can be determined.
Through the process of translation, a chain of amino acids is formed from the mRNA in the cytoplasm.
The entire process is known as “central dogma” which is regulated by the regulatory elements.
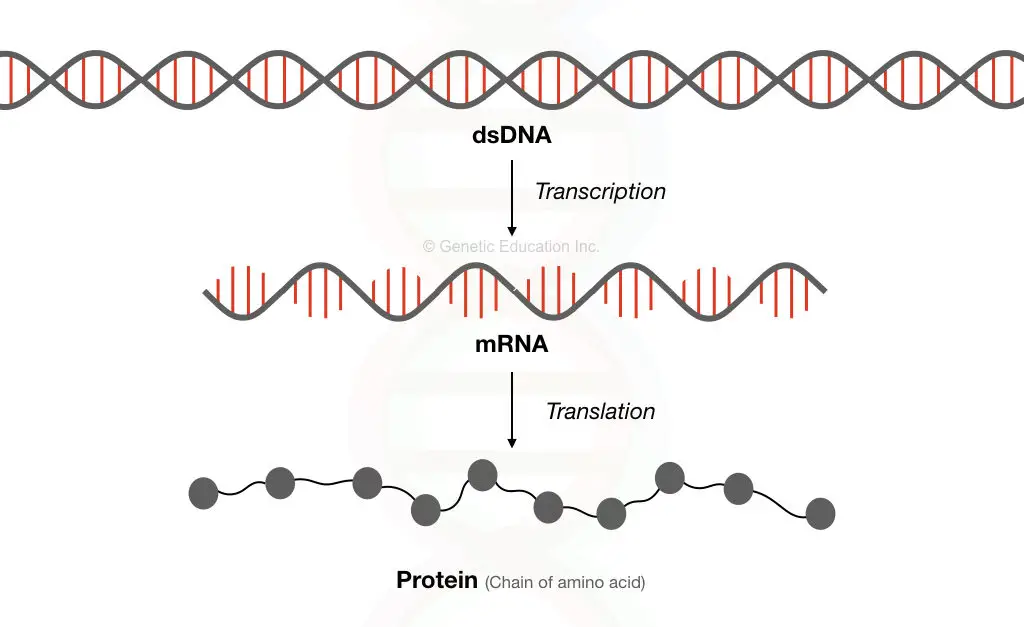
Sometimes, DNA polymerase inserts the wrong nucleotide or some external factor damages our DNA, it happens!
DNA repair pathway helps to seal those damaged gaps or repair the wrong mismatched nucleotides.
Non-homologous end-joining and direct DNA repair are the two DNA repair pathways by which our DNA is maintained in proper order.
Interestingly,
“Change in the nucleotide sequence or alteration in the DNA structure is called a mutation.”
Cis and trans acting elements of the genome:
Cis-Acting elements are the type of non-coding DNA sequences that regulates the level of gene expression. Remember, the cis-acting elements are actually the non-coding sequences.
On the other side, the trans-acting elements are the proteins, enzymes and transcriptional factors, act on the cis-elements that actually encoded by some other genes that help to control gene expression.
Thus the trans-acting elements can work on different cis-acting elements of a different or single gene.
Promoters, enhancers, silencers, repressions and insulators are the type of cis-acting elements.
Promoter plays an important role in gene regulation, we will cover a dedicated article on promoters in some other segments.
Genomics:
Now coming to another important point related to the genome- genomics.
A gene can be easily be studied but studying the entire genome is a tedious, laborious and time-consuming task.
The study of gene or DNA sequence is called genetics while the study of the entire genome and its function is called genomics.
Read our amazing article related to it: “Genome Vs Gene”, An Unusual Comparison.
The genome of eukaryotes or higher eukaryotes is more complicated and larger than the prokaryotes thus studying the entire genome of an organism is quite difficult.
However, some techniques such as DNA sequencing and DNA Microarray have the power to analyze the entire genome and are therefore used in genome-wide association studies.
Whole-genome sequencing:
Unlike the sanger sequencing whole genome sequencing can analyze the whole genome- whole-genome shotgun sequencing, high throughput DNA sequencing and next-generation DNA sequencing method are three of the best method used to do so.
Briefly for doing that, first, the entire genome is cleaved or fragmented into smaller fragments using the restriction digestion and created the library of fragments.
In the next step, each fragment is ligated with the known adaptors and sent for DNA sequencing.
Each fragment is sequenced multiple times and “contings” are generated.
Finally, the fragments are analyzed and arranged back on each chromosome and the entire sequence is generated.
Note: this is the broader overview, the method, principle and chemistry of sequencing varies based on the selected method.
What do we get from the whole genome sequencing?
A gene sequencing will tell us about sequence variations in a gene while in a genome sequencing variation such as SNPs, copy number variation, deletion, duplications and other alteration is determined.
New mutations or alterations are also identified using it which is beneficial in the genome-wide association studies.
Related article: 3 Of The Best Genome Sequencing Methods.
DNA microarray:
DNA microarray is a state-of-the-art technology works based on the mechanism of hybridization.
In microarray, especially whole chromosome microarray, all the mutations, alterations or copy number variation are screened in a single assay- on a chip.
However, it is dedicatedly working only for the known mutation, the new alteration can not be determined.
The method is used majorly for comparing alterations between two genomes or more than two genomes thus new variation can be established using it.
Further, gene expression can also be determined by it.
Related article: Genome-On-A-Chip: DNA Microarray.
Conclusion:
Although we have sequenced the entire genome, we have less information about it. Our genome is a more complicated thing than we think. To date, we have just identified only several thousand genes but scientists are constantly studying our genome to solve the mysteries of life.
Nonetheless, the human genome project helped us a lot in understanding the genomic properties of humans and other organisms’ genomes. Read the article on HGP: The Human Genome Project: Aims, Objectives, Techniques and Outcomes.


