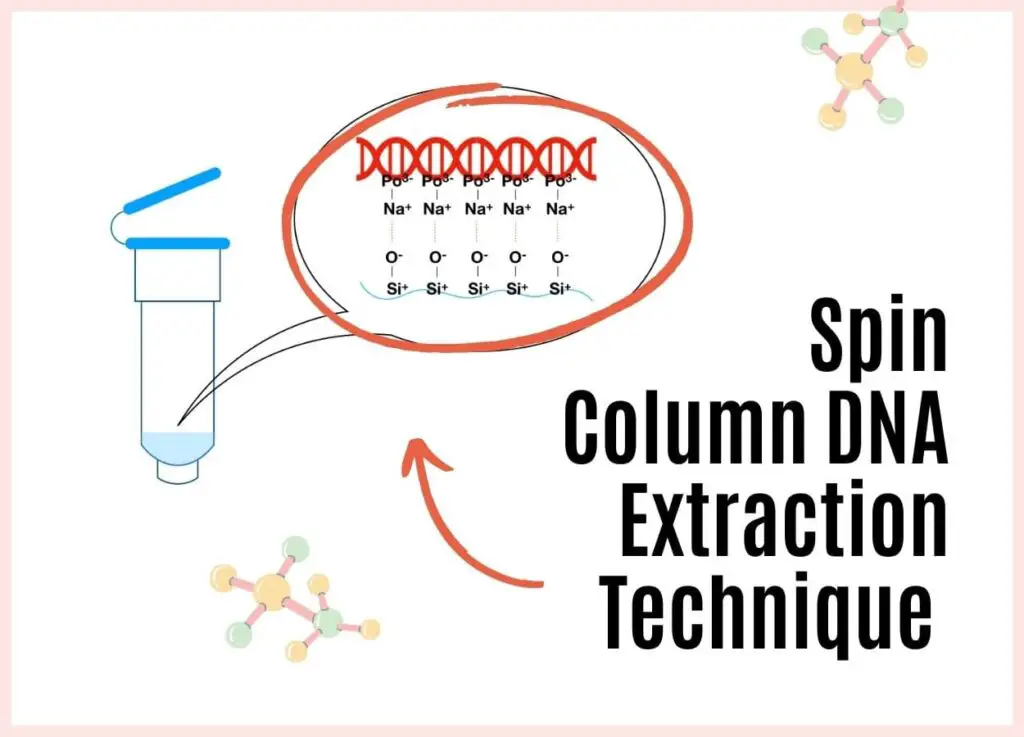
“Spin-column DNA extraction falls into the solid-phase DNA extraction category. It is quick, effective and popularly utilized in recent times. However, like all other technology, it has its own advantages and limitations. Let’s understand the technology and its background.”
DNA extraction or isolation has pivotal significance in genetic research. B’cos it’s an essential step that provides a starting material for the experiment. However, extracted DNA should have these two important qualities– first it should have excellent purity and second, it should have sufficient quantity.
Numerous ready-to-use kits and manual DNA extraction methods are available and scientists use them, but scientists face issues with the purity and quantity of extract. In addition, with such methods, speed and cost also remain considerable problems too.
So to solve many nucleic acid extraction problems, spin-column technology was introduced. In this article, we will understand the spin column DNA extraction technology including— working, principle, process, steps and requirements.
Most importantly, we will discuss the advantages and limitations of the present technology so that you can make your decision to go with it or not. Let’s dive into the article.
Key Topics:
Introduction to Spin-column Technology
Spin-column DNA extraction falls under solid-phase DNA extraction as it uses a solid phase, usually, silica to separate nucleic acid. The original concept was developed by McCormick and co-workers in 1989. Timely new concepts and optimizations led the technique to the top of all.
One of the major advantages of spin columns is the lack of dangerous and hazardous chemicals like phenol and chloroform. Instead, a few safe chemical solutions and two to three centrifugations, make the whole process easy and safe. The single uniform protocol is applicable on many biological samples.
The most talked part of this technology, “spin-column” contains solid silica gel where DNA binds. Only a few treatments afterward will exact our pure DNA without any precipitation required. The principle of the technique is explained here.
Principle of spin-column DNA extraction
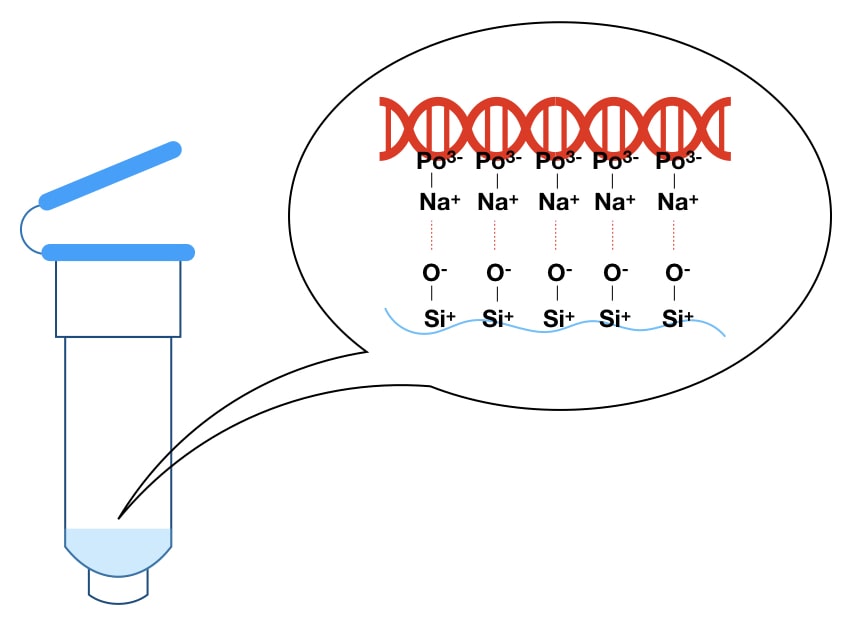
The biological sample containing proteins, lipids and other cellular constituents is digested using a lysis solution and Proteinase K; lysate is applied in the spin column where it binds to the positively charged solid silica surface. Washing buffer removes all other debris while the pure DNA elutes under the treatment of high pH and salt-containing elution buffer.
The entire process is elaboratively explained in the steps portion of this article.
Kit components
Spin-columns
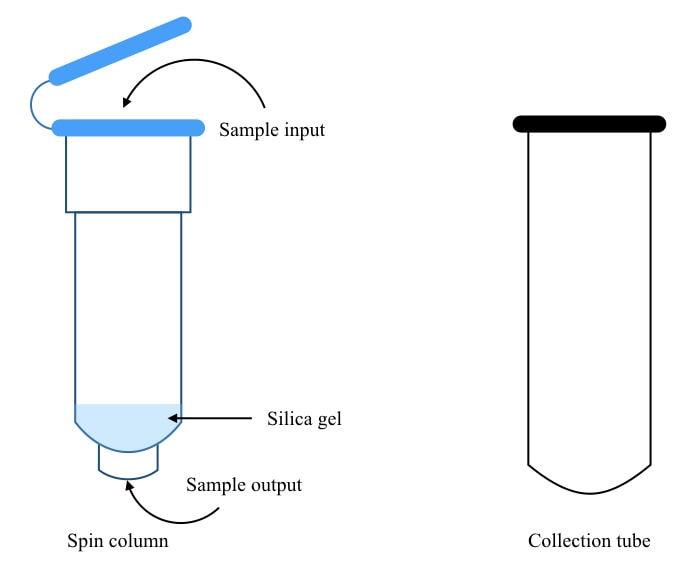
The important part of any ready-to-use spin-column-based DNA extraction kit is the spin column itself. The atypical design allows effective adsorption, purification and elution. The spin column is made up of high-quality plastic.
Technically the column tube is a bit smaller in size than the collection tube with a narrow opening at the bottom, so that it can fit into the collection tube. Just above the narrow opening, the tube is filled with silica gel- a solid matrix having positive charges that attract and bind DNA to the matrix.
The ideal column tube should be sterile and remain stable under heat, cold or chemical reaction. Other than silica column tubes, many other columns are often used in various types of nucleic acid extractions. Here are some.
Silica column
Since 1979, silica has been used in various extraction experiments. Under low salt and high alkaline conditions, silica selectively binds with the negatively charged DNA. The Silica column is filled with silica gel. It is commonly used for genomic, plant and bacterial DNA isolation.
Silica columns are widely used in kits, research and the diagnostic industry for DNA isolation and purification.
Anion exchange column
The anion exchange column has a unique IEXM- Anion exchange membrane which attracts and binds with the negatively charged particles, like the DNA. The remaining extraction process will remain the same like other column-based techniques. This column is used for plasmid DNA extraction.
Commercially available IEXM has a higher binding capacity and selectivity. Research also shows that it has almost 100% plasmid DNA recovery (Yang et al., 2008).
Glass fiber spin column
The present column uses glass fibers– silica glass, borosilicate or flint glass powder instead of silica gel or anion resins. It has slight superiority, as it has high sample binding capacity and negligible sample loss. The rest of the principle remains the same and such columns are commonly used for genomic and plasmid DNA extraction and RNA extraction and purification.
The less penetration of the glass fiber spin column is because of its higher cost.
Cellulose column
Yet another interesting alternative to the silica column is the cellulose column which contains cellulose paper instead of silica or glass particles. Papers as a column are low cost, low throughput and highly effective. And hence is employed for plant genomic DNA and plant total RNA extraction. It is also used for the purification of PCR products.
Unfortunately, having low binding affinity limits its use for plasmid DNA isolation. Noteworthy, cellulose paper pipette filter tips and commercial papers are now also available for effective DNA isolation (Shi et al., 2018).
Diatomaceous earth
Broom et al first described this material as a solid phase matrix for column technology. It consists of around 90% silica, alumina 2% and hematite up to 2%. The principle and process remain the same as the silica column.
Diatomaceous earth is more simple and handier. It has less pipetting errors and recovers single and double-stranded DNA and plasmid DNA, effectively. However, it is costlier than other options so not commonly used in labs.
Collection tubes
Collection tubes are small high-grade plastic tubes in which the columns are placed and simultaneously, liquid waste is collected.
Proteinase K solution
Proteinase K solution or protease is available in commercial kits and is an important component of any ready-to-use kit. It has the function to digest protein parts of a cell. Either enzyme present in the kit is free from any DNase or RNase.
Technical note
Generally, the proteinase K is provided in lyophilized form and remains stable for 12 months at room temperature, however, the solution form should be stored at 4 to 8°C. So it is recommended to store proteinase K (any form) at 8°C temperature for optimum activity.
Lysis buffer
Along with proteinase K, the lysis procedure completes using the lysis solution which lacks phenol and chloroform, like the conventional method and effectively performs cell lysis. It’s important to know that different manufacturers have different recipes for lysis buffers and don’t reveal their formulas.
However, I certainly know that the basic chemical foundation for lysis buffers remains the same. If you wish to read more about the lysis buffer, I strongly recommend reading this article: How to prepare lysis buffer for DNA extraction?
Binding buffer
A binding buffer helps DNA or RNA to bind with the silica membrane in the column. Some manufacturers provide lysis and binding buffers in the same composition. It reduces the cost, extra steps and complexity.
We will discuss the process of binding in the upcoming section.
Washing buffer
Washing buffer usually contains ethanol or isopropanol. Washing buffer selectively removes only cell debris, proteins and other cellular molecules excluding DNA.
Elution buffer
Elution buffer is a high pH solution that detaches DNA from the silica matrix, dissolves it and elutes it. The ideal elution buffer contains TE buffer solution (pH 9.0). To know more about the Tris-EDTA buffer, read this article: Tris-EDTA buffer.
Technical note
Steps
The ready-to-use spin column DNA extraction kit performs the action in only 4 separate steps and 4 spins. The upcoming section explains each step, extensively.
Lysis
Spin column lysis occurs much like the traditional/manual process of DNA extraction. The solution of proteinase K and lysis buffer (both provided by the manufacturer) are added to the sample. And mixed well. The techniques of mixing are vortexing, inverting the tube, homogenization, tissue grinding using physical methods, etc.
As we explained, various chemicals of lysis/binding buffer and proteinase K lyse proteins, lipids and other cellular debris. Once the lysis completes, the lysate is transferred to the spin column tube.
Lysis is performed in a separate 2 ml Eppendorf tube– not provided in the kit. It takes 15 to 20 minutes.
Technical note
Adsorption
Now the lysate mixture, in the second step, is transferred to the column which is pre-placed on the collection tube. Technically what happens during adsorption is explained here.
The negative charge of the DNA reacts with the positive charge of the silica gel matrix and makes a stronger bond in a competitive reaction. While the rest of the cellular components remain in the solution. This eventually occurred due to controlled salt and pH conditions.
Load only 200µL solutions on the column. Avoid overfilling/ flowing which contaminates the final elute. Allow settling and reacting DNA for 5 to 10 minutes.
Washing
In the third step, washing is performed. The wash buffer usually contains either ethanol or isopropanol and removes debris other than DNA. Keep in mind that our DNA is still adsorbed on the silica matrix. From the pores of silica, and under the higher centrifugal force or vacuum, the lysate is passed from the column and collected in the collection tube.
Two-step washing is performed to gain better-quality DNA. However, frequent washing causes DNA loss and reduces yield.
Elution
In the final/fourth step, our extract-DNA is eluted out from the column. High pH and salt elution buffers are added to the column which dissolves DNA by removing the bonding between the phosphate of DNA (Negative charge) and silica (positive charge).
The common composition of the elution buffer is a TE solution of pH 9.0. Oftentimes, dd/w is also used to elute DNA.
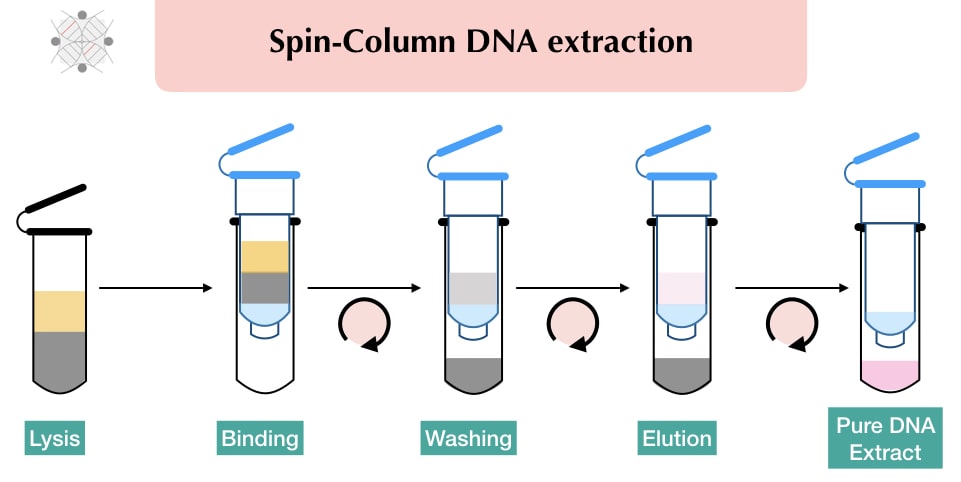
Process of Spin column DNA extraction (Protocol)
Every spin column kit manufacturer provides their own working protocol, nonetheless, is almost similar, super handy, easy to perform and quick. I am here just to provide the general outline of the protocol for better understanding.
I recommend you follow the steps provided in the kit. Also, this protocol is mainly adopted from the original Qiagen DNA extraction protocol.
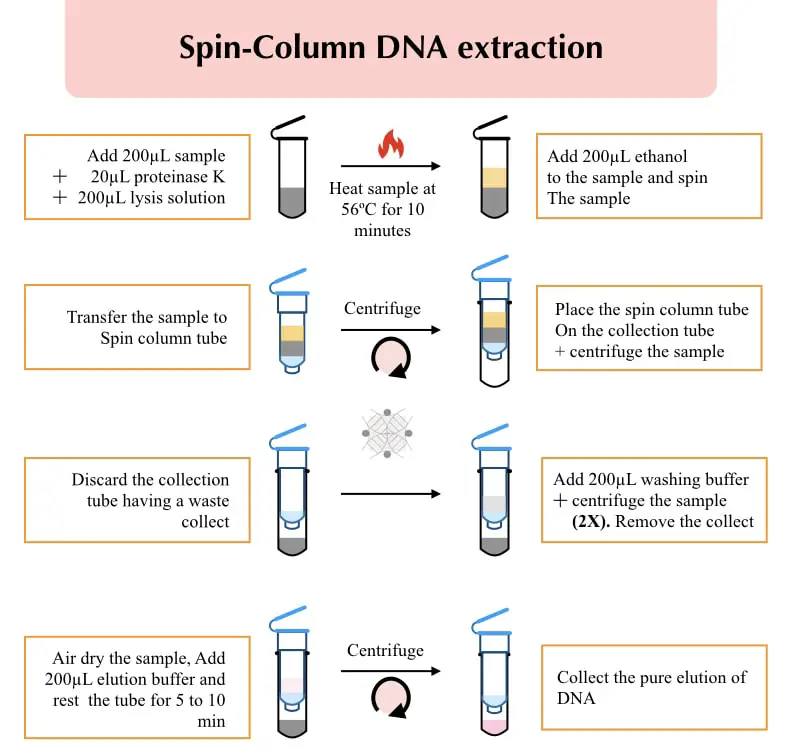
- Add 20µL Proteinase K at the bottom of a 2 ml microcentrifuge tube.
- Add 200µL sample– blood, serum, buffy coat or preparation.
- Add 200µL lysis buffer provided by the manufacturer. Mix them well using vortex or gentle inverting many times.
- Put the sample at 56°C for 10 minutes for effective lysis. Now add 200µL alcohol, mix well by vortexing and centrifuge to settle down.
- Transfer the mixture carefully to the spin column. Keep in mind that the lysate should not leak or wet the rim of the tube.
- Allow the sample to settle for 5 minutes and centrifuge it at 10,000 rpm for 2 minutes.
- Remove the collection tube and place the spin column on a new collection tube.
- Add 200µL washing buffer and centrifuge at 10,000 rpm for 2 minutes.
- Remove the collection tube, place the spin column on a new collection tube and add 200µL elution buffer.
- Place the sample for at least 15 to 20 minutes and centrifuge at 10,000 for 2 minutes.
- Transfer the elute into a new sterile tube (our pure DNA) and put it at 4 to 8°C temperature.
Now coming to the most important part of our discussion which eventually helps to understand the importance of the present techniques. The pros and cons of the spin column DNA extraction technology.
Advantages
The present technology has changed the entire scenario for not only DNA but RNA extraction and purification; so is widely penetrated. All the important advantages are explained in this section.
Safer
It doesn’t require hazardous and dangerous chemicals like phenol and chloroform which is the most lucrative advantage. Previous techniques highly rely on organic chemicals and solutions. Among which some have serious health hazards.
Phenol is volatile and corrosive and can burn the skin while chloroform also has very serious adverse effects on human health. Thanks to the spin column, we no longer need such chemicals.
Zero pre-preparation
The present technique is so super handsome as it nearly doesn’t need any extensive pre-preparation. In previous manual methods like the other chemical-based methods, we have to prepare each chemical and solution which is indeed a time-consuming process.
For example, to prepare a saturated phenol solution, lab personnel needs almost a day while preparing solutions like TE buffer, and lysis buffer take additional time. All the essential chemicals in the spin column kit come as read-to-use. Hence no extensive prerequisite is required.
And so comes another important advantage.
Super fast
The present technique as we mentioned, lacks extensive prerequisites and fewer steps thus is super fast. The extraction completes within an hour, exactly saying- it takes 28 to 35 min. So it saves so much time for experiments. Here is a chart of how much time each extraction technique took.
| Technique | with pre-preparation | without pre-preparation |
| Phenol, chloroform and isoamyl alcohol DNA extraction | 26 to 28 hours | 2 to 2.5 hours |
| CTAB DNA extraction | 6 to 7 hours | 1.5 to 2 hours |
| Proteinase K DNA extraction | 1.5 hours | 1 to 1.5 hours |
| Spin column DNA extraction | 1 hour | 28 to 34 minutes |
High purity
If someone asks why is the spin column we use? The answer should be because of the high purity. The kit available by Qiagen is so precise that every kit every time gives great purity which is so important in genetic research.
Abdel-Latif & Osman used three genomic DNA extraction methods– Qiagen spin column DNeasy plant DNA extraction kit, conventional CTAB method and Qiagen mericon DNA extraction kit. The highest purity obtained was between 1.80 to 1.95 using the Qiagen spin column DNeasy plant DNA extraction.
However, the purity obtained by any spin column kit depends on the quality of chemicals used, the preparation of chemicals, the quality and pores size of silica. Human hands have less to do with such kits.
In our in-house lab experiments, as though we have years of experience and even excellent working SOP, we get purity at 260/280 ratio between 1.70 to 1.85, which is indeed excellent. Notedly, the purity varies from sample type to sample type.
Efficient
Ready-to-use kits have a problem with yield as only a limited volume of samples can be processed. But it gives amazing yield for sensitive and low-volume samples like plasmid DNA, serum or buffy coat. Even in comparison with the manual methods, for samples like amniotic fluid or CVS, the spin column technique gives excellent yield every time.
As we said, the overall yield is still very poor in comparison with manual methods. The present technology is powerful enough to yield 3 to 6 µG DNA from only a 200 µL sample.
Less starting material
Spin column technology is so optimized that it can even work excellently with lower-volume samples. Even it can handle 50-100µL samples and can still give maximum yield and purity which no other technique can do.
Newbie-friendly
The present technique is also newbie friendly. Less centrifugation and pipetting steps make the technique so user-friendly, that a person with only some basic knowledge of DNA extraction can perform it.
Reproducible results
Each kit has high reproducibility as every time you will get nearly as good as previous results (quality and yield). Keep in mind that the final results depend on how effectively you have performed each step, each time.
Limitations
No technology is completely advantageous, and so the spin column is! Some crucial shortcomings, the spin column technique has. Here are a few.
Loss of material
Our experts face one crucial downfall– a loss of material or DNA. With each wash, a substantial amount of DNA is lost. So as we increase the number of washing steps, we get less yield.
Furthermore, during the processing and due to poor binding, some DNA may also be lost.
Impurities
Some impurities still remain in the sample which creates problems in downstream testing like amplification and sequencing.
Low yield
The overall yield of DNA is very low, compared to manual methods. Any spin column kit can yield a maximum of 5 to 6 µG DNA while techniques like PCI can produce up to 10-15 µG of DNA or even more for blood.
Fewer optimization options
As the technique comes almost as ready-to-use, there are very few options for optimizations. For the majority of samples like plasmid, blood or culture, it works uniformly but for some hard samples like the plant, we need to perform optimization at every level.
Every plant species has a different polyphenolic constituent and hence needs a separate extraction protocol. Here a single kit can’t give uniform results. So manual methods and optimization strategies have their own importance which eventually does not come up with spin columns.
If you would like to sharpen your DNA extraction skills for nearly every sample type. I strongly recommend checking out your handbook: From DNA extraction to PCR.
Costly
Column-based extraction kits are affordable but not super cheap. On the other hand, manual methods, though are tedious but cheap, I mean very cheap. A single kit can perform 25 to 50 samples.
But when you prepare each chemical manually and optimize your protocol well, you can use those chemicals for so many samples. Manual methods are super cheap while column-based methods are costly, comparatively.
Furthermore, the overall cost of the experiment becomes even costlier when additional options like a vacuum machine or automated extraction machine are used.
Wrapping up:
Spin-column DNA extraction technology is by far the most penetrated technology in the industry, people are using it and is widely accepted, not only in research but in diagnosis as well. Undoubtedly, it is a kind of semi-automated, speedy and nearly perfect technique to separate and purify any DNA.
It uses a silica matrix as a solid phase on which DNA binds and separates. No additional chemical or solution preparation is required here. The technique is safer to use and hence is popular. Qiagen is the biggest manufacturer of various spin column kits, though there are many companies available in the market.
I strongly recommend using a spin column extraction kit for sensitive samples like— bone marrow, solid tumor tissue, semen, amniotic fluid, chorionic villi, cell-free DNA or tumor DNA.
For routines like blood or plant DNA extraction, manual methods are good options. Spin columns save time and give working yield and purity.
Sources:
Yang, Yu et al. “High-performance DNA purification using a novel ion exchange matrix.” Journal of biomolecular techniques: JBT vol. 19,3 (2008): 205-10.
Shi R, Lewis RS, Panthee DR (2018) Filter paper-based spin column method for cost-efficient DNA or RNA purification. PLOS ONE 13(12): e0203011.
Abdel-Latif, A., Osman, G. Comparison of three genomic DNA extraction methods to obtain high DNA quality from maize. Plant Methods 13, 1 (2017).
McCormick, R M. “A solid-phase extraction procedure for DNA purification.” Analytical biochemistry vol. 181,1 (1989): 66-74.
Boom, R et al. “Rapid and simple method for purification of nucleic acids.” Journal of clinical microbiology vol. 28,3 (1990): 495-503.