“RNA extraction is a process to isolate various or a specific type of RNA molecules for gene expression studies.”
Download this article as an ebook: RNA Extraction pdf.
Snippet:
- The function of RNase is to cleave RNA. It is present everywhere on our hands, atmosphere, the bend on which we are working and in the environment.
- RNA is also a NA and less stable than DNA.
- RNA isolation and purification are performed under cold and aseptic conditions, it starts degrading immediately.
Like DNA, RNA (ribonucleic acid) is also a type of nucleic acid. So it should have a similar extraction process to the DNA, right.
Theoretically, Yes.
“Somehow” the process is similar
But practically, No.
It is substantially different and needs some different chemicals to extract RNA and not DNA. RNA though isn’t our genetic material but important for us. The final protein is manufactured from the mRNA only. It is the polynucleotide chain of only coding sequences which is also a reason to isolate mRNA from a total cellular RNA.
And that mRNA can encode a protein only if the tRNA and rRNA are there to help it. Meaning, the RNA is significantly important for our cells. Note that several pathogens have RNA as their genetic material, for example, influenza.
RNA is a polynucleotide chain made up of sugar, phosphate and nitrogenous bases, however, unlike the DNA it has Uracil instead of Thymine along with Cytosine, Adenine, and Guanine as nitrogenous bases.
Three specific forms of RNA are tRNA, rRNA and mRNA while other miscellaneous RNAs are miRNA, shRNA, piRNA and other smaller non-coding RNAs. Each type of RNA has a unique function to perform in a cell, you can read this article to learn more: RNA: Structure, Types and Function.
It is an intermediate ‘transcript’ that finally translates into an amino acid chain. So ideally, RNA extraction and studies will provide lucrative cellular function data, the “gene expression”.
Effective RNA extraction allows researchers to study gene expression precisely and accurately. RNA extraction is a prerequisite for pathogen identification. In this article, we will understand one of the most challenging genetic procedures, RNA extraction.
We will also look into the principle, steps, procedures, importance and a common RNA extraction protocol. My team has huge experience and expertise in such techniques, so this article will help you to understand every aspect of RNA extraction.
Stay tuned.
Key Topics:
RNA extraction- Basics
RNA extraction or isolation is a process to separate any type of RNA from a cell. It is useful for gene expression studies, pathogen identification and classification. As it’s a nucleic acid-based detection technique, it is accurate, reproducible and versatile.
Furthermore, the RNA-based NA detection techniques are robust, cost-effective and handy too, and henceforth in modern times, it is employed for infection detection and pathogen identification.
Recently popular method RT-PCR for COVID-19 detection is also an RNA-based DNA quantification method in which the viral RNA (SARS-CoV-2) is isolated, reverse transcribed into DNA and quantified.
In the past, for example, during H1N1 pandemics and epidemics, RNA-based nucleic acid detection techniques (PCR) were used for detection and management. However, to make these assays rapid and robust, ready-to-use RNA extraction kits are available.
Nonetheless, in the pre-genetic era, such RNA extraction-based RT-PCR detection techniques were not so popular. The RNA extraction technique is tedious and has higher contamination chances. Chomczynski and Sachhi (1987) postulated the first single-step RNA extraction method.
Their research article entitled, “Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction” explained the entire process using phenol and guanidinium thiocyanate.
Read-to-use RNA extraction kits are more advisable for pathogen detection and identification which relies on the principle of column-based separation. On the other hand, conventional techniques, however, challenging and laborious, are cost-effective and streamlined.
As a geneticist, everyone should have to learn conventional RNA extraction methods. So in this article, we will only discuss a conventional technique.
Principle of RNA extraction:
An RNA of a living cell (animal, plant, human, bacteria or virus) provides information on transcriptional activity, thus, important for gene expression studies. RNA lysis buffer lyses the cell wall/membrane while the use of organic solvents like phenol-chloroform separates RNA upon centrifugation.
Phenol: chloroform: isoamyl alcohol, by centrifugation, forms three layers viz the lower organic layer, middle DNA and protein-containing layer and upper aqueous RNA-containing layer. The upper aqueous phase is collected, precipitated, washed and dissolved in RNase-free water.
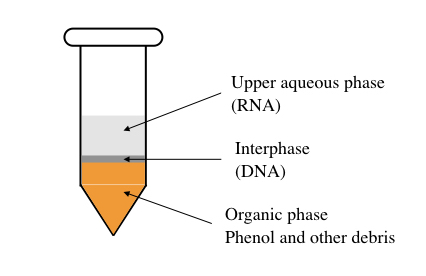
Phase reparation by phenol: chloroform: isoamyl alcohol.
“Organic solvent-based nucleic acid methods using phenol, chloroform and isoamyl alcohol are considered gold-standard and high yield methods.”
The problem with RNA extraction:
As aforementioned, there is a problem handling the RNA and restricts the reproducibility of any RNA extraction method. Some common problems are explained here,
- RNA is single-stranded and less stable than DNA.
- Heat and hydrolysis can degrade the sugar backbone of RNA easily.
- Less cellular quantity results in low yield.
- Less stability of RNA: The tissue should be immediately processed for extraction as the time increases the chances that RNase will degrade RNA increases. Using commercially available RNA stabilizers can help to extend RNA pre-processing time, though.
- And most importantly, the universal presence of the RNase enzyme degrades RNA more easily than DNA. RNase is predominantly present in the atmosphere, surrounding environment, on a working desk or even on our hands degrades the extracted RNA more easily.
These reasons make the RNA isolation technique more sensitive and specialized which needs extraordinary preparation and care, every time. The effective RNA extraction protocol relies on these 5 things,
- Inactivation of RNase
- Stabilizing RNA
- Isolating RNA from DNA and protein.
- Precipitating only RNA
- Removing the DNA
Steps in RNA extraction
- Sample collection
- Sample preparation
- Tissue homogenization
- Cell lysis
- RNA precipitation
- RNA washing
- RNA elution
- RNA quality check
- RNA quantification
- RNA integrity check
- Storing and handling
Process of RNA extraction:
Sample collection:
Cells, cell lines, cultures and tissues are common sample types for RNA extraction. The sample is isolated aseptically and stored at -80℃. It is advisable to collect samples by an expert. In order to make RNA stable, the tissues are stored at -80℃ before processing.
Note that additives are added to store the tissue sample for a longer period of time, as RNA starts losing its integrity under adverse environments.
Sample preparation:
Preparing a sample before RNA extraction is so crucial for higher yield and greater purity. Oftentimes, the tissue homogenization technique is utilized to prepare a homogenous mixture of cells.
Usually, for plant tissue or solid mammalian tissues, a mortar and pestle can be used (although professional tissue homogenizers are advisable). During tissue grinding, RNA lysis buffer is carefully added, slowly to the sample.
Upon getting fine homogenization, the sample is ready for extraction. Noteworthy, perform steps at 4℃ temperature and under the strict aseptic hood. Take necessary precautions to avoid RNase activities.
Sample quantity:
| Sample type | Concentration |
| Tissue sample | 50 to 100 mG |
| Monolayer cell culture | 1 × 10(5) –1 × 10(7) |
| Cell culture in suspension | 5–10 × 10(6) cells from animal, plant, or yeast origin or 1 × 10(7) cells of bacterial origin |
Optimization
Quickly perform RNA isolation after sample collection, RNA degrades rapidly, or store tissue sample immediately into liquid nitrogen at -80℃ for long-term use.
Cell lysis and chemical treatment:
Chemical processing followed by centrifugation separates RNA from other cell debris and chemical traces. Common chemicals are combinations of various organic solvents that separate RNA into the upper aqueous phase.
An additional step of DNase treatment can be applied to remove the DNA from the sample, which eventually yields a good quality and highly pure RNA.
Note:
Incubating a sample at room temperature for 5 to 8 minutes dissolves the nucleo-protein complex.
Precipitation:
The RNA precipitation process is much similar to the DNA precipitation process. Here using any alcohol the RNA is precipitated into a visible form. However, to precipitate only RNA, a specialized agent Lithium chloride is used.
RNA washing:
Precipitated RNA pellets are washed before dissolving and downstream processing. Washing removes other contaminants and debris like DNA, protein and traces of phenol and other chemicals. Again the process of nucleic acid washing remains the same as DNA washing. Washing gives a pure, whitish RNA precipitate.
Dissolving RNA:
RNA is dissolved into RNase-free water (Which is a type of nuclease-free water), TE buffer and specialized RNA dissolving solution. Commercially available RNA dissolving agents also contain RNA stabilizing agents which is more advisable.
Take a look at some recommendations for dissolving RNA.
- Dissolve RNA in RNase-free water or RNase-free pre-autoclave TE buffer.
- Do not dry the RNA pellets by heat.
- Store dissolved RNA at -80℃.
- Use RNase inactivation solution to store the RNA.
- Do not store it with other tissue or DNA samples.
RNA quality checks:
The conventional agarose gel electrophoresis technique is used to determine the presence of RNA and investigate the quality of RNA. The total extracted RNA is run on a 0.8 to 1% agarose gel.
RNA quantification:
Spectroscopic and fluorescence analysis are two common methods to quantify RNA. The 260/280 spectroscopic ratio is taken into consideration for the RNA quality check which is between 1.9 to 2.1.
The fluorescence technique relies on the use of fluorochrome that binds with the single-stranded RNA and quantifies it by detection.
RNA integrity check:
RNA integrity analysis is so crucial in the RNA extraction process. It shows us how much RNA is remained intact and is usable. It is a numerical value known as RIN. We have covered a comprehensive article on RIN- RNA Integrity Number. You can read it to know more. The whole principle, process and RIN results are explained in this article.
A common RNA extraction protocol:
A common RNA extraction protocol is somehow similar to DNA extraction protocol and completes in steps like lysis, precipitation, washing and elution, however, the use of some special chemicals helps to extract total RNA and not DNA.
Note that RNA extraction is a sensitive procedure and requires extra care so every instrument, glass and plasticware used in the experiments is pre-sterilized and treated with RNase- free water.
Follow common laboratory safety guidelines, and RNA extraction handling routines for extraction. This protocol is available online, we have made slight changes while using it.
Instruments and equipment:
- Tissue homogenizer
- Cooling centrifugation system
- Vortex
- Mortar and pestle
- Freeze and deep-freeze (-20℃ and -80℃)
- Incubator
- Laminar hood or extraction chamber
Chemicals and reagents:
- RNA lysis solution
- Saturated-phenol
- Sodium Acetate
- Phenol
- Chloroform
- Isoamyl alcohol
- Ethanol
- RNase-free water
- Lithium Chloride
- Glycogen (If required)
Preparation of RNA lysis solution
- 4M Guanidine Thiocyanate
- 25mM Sodium Citrate, pH 7
- 0.5% N-lauroylsarcosine
- 0.1M beta-mercaptoethanol
Function of RNA extraction chemicals:
| Chemical | Concentration | Function |
| RNA lysis buffer | Given above | Cell lysis |
| water-saturated phenol | – | Cell lysis |
| Sodium Acetate | 2M | |
| Chloroform | – | Phase separation |
| Isoamyl alcohol | Phase separation | |
| Ethanol | 70 to 74% | RNA washing |
| RNase free water | – | Remove RNase activity |
| Isopropanol | 70% | precipitation |
| Lithium Chloride | 7.5M | Precipitation |
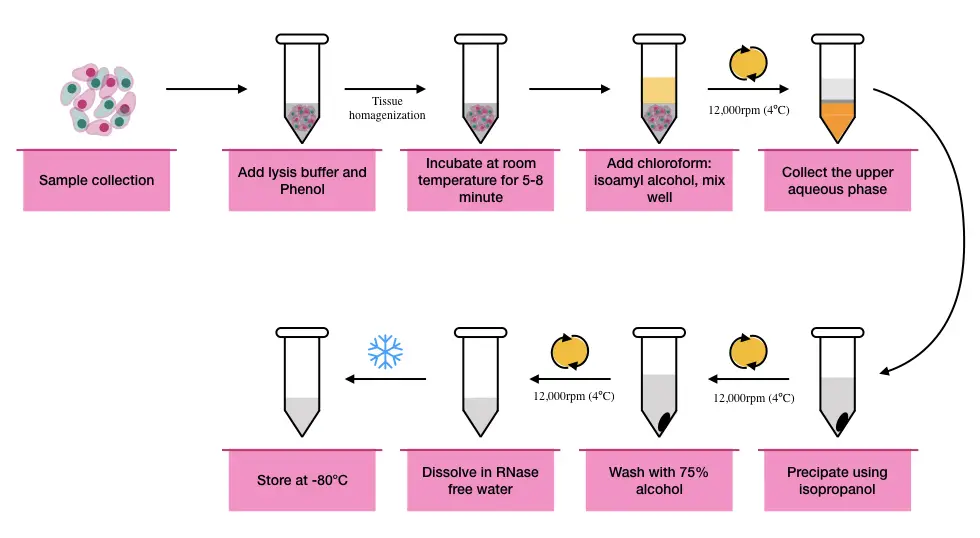
Protocol:
- Take a tissue sample of 5 to 100mg and grind using a mortar and pestle. (if you have a harder tissue use liquid nitrogen to break tissue). You can use a tissue homogenizer too.
- Add 1ml of RNA lysis buffer to homogenize (lyse) tissues and allow incubation for 5 min at room temperature to break the nucleoprotein complex.
(Recent studies suggest that guanidium thiocyanate, a chaotropic salt, is also used in the RNA lysis buffer which effectively isolates the RNA, prevents it from RNase and maintains integrity).
- Tissue homogenization should be ideally performed in cool or at 4℃ temperature to maintain the stability of RNA and inactivating RNase.
- After completion of homogenization, take the sample into a sterile, RNase-free tube and add 1ml phenol, 0.2 ml chloroform: isoamyl alcohol (49:1) and 0.1ml of sodium acetate to the per 1ml homogenized sample.
- Mix well by inverting the tube several times.
- Immediately, centrifuge the sample at high speed for 15 minutes at 4℃.
- After centrifugation, transfer the top aqueous phase to the new RNase free-sterile tube and add precipitant either isopropanol or Lithium chloride.
- Add an equal volume of (70 to 75%) isopropanol or 2.M LiCl solution to the aqueous phase, incubate at -20℃ for 1hr and precipitate by inverting the tube several times.
Pro-tip
For effective precipitation, use only LiCl and incubate the sample at -20℃ overnight.
- Once the precipitate appears, centrifuge the tube at 12,000 rpm and remove the aqueous phase.
Note:
The use of Lithium chloride (LiCl) only precipitates the RNA present in the sample and not DNA or protein. It is advisable to use LiCl.
- Add 75% ethanol to wash the pellets and centrifuge the tube.
- Air-dry the pellets and dissolve them in RNase-free water.
- Store the RNA in aliquots at -80℃ to avoid repeated freezing and thawing.
Optimization:
You can also add commercially available DNase solution (use concentration as per the manufacturer’s advice) during the lysis to destroy the DNA.
Note that this one is the simplest and most basic protocol for RNA extraction, tissue to tissue and organism to organism the protocol and chemical composition may vary. We will discuss various protocols in the upcoming article.
Safety precautions are advisable, read manuals and safety guides before performing the experiment. Chemicals like phenol and chloroform are hazardous.
Related article: Effectual Trizol RNA Extraction Protocol using Tri Reagent.
Importance and applications of RNA extraction:
The pre-requirement of gene expression studies is RNA extraction. As said above, mRNA transcribes into the chain of amino acids, by studying a tissue-specific mRNA will give us an idea about the amount of protein formed.
It is used in qPCR assays like reverse transcription PCR, and real-time PCR, which can quantify the amount of a specific type of mRNA that translates a specific protein.
Genome-wide association studies like microarrays often need a total mRNA for studying the total transcriptomes of the genome or wide varieties of protein-governing genes and their expressions.
Sequencing techniques like whole-transcriptome sequencing or RNAseq also need a pure mRNA and total RNA, respectively. Total transcriptomic sequence the entire mRNA governing DNA. It provides insight into alterations in gene expression.
RNA extraction, therefore, is important for expression studies in plants, animals, humans or bacteria.
A total RNA or mRNA extraction also has significant importance in cancer genetics to demonstrate the role of a particular gene (oncogenes or proto-oncogenes), indels or SNPs.
Other than these classic applications, advancements during the recent pandemics suggest that mRNA extraction can often be used for the precise identification and characterization of pathogens whose genetic material is RNA, not DNA.
For instance, for SARS-CoV-2 or Influenza H1N1, isolated RNA is reverse transcribed into DNA, and a specific gene can be investigated in a Real-time PCR. The quantification of the template gives us an idea about the amount of infection and the presence of a specific type or strain.
RNA extraction, a pre-step in RT-PCR can be used for pathogen identification, classification, characterization, nomenclature and detection needs pure pathogen RNA.
Recent advancements in RNA extraction technology:
RNA extraction is nowadays popular. It has applications in expression studies and identification. Point-to-care diagnostic industries use ready-to-use isolation kits. In principle, such kits are based on spin-column technology, unlike the nucleic acid precipitation that directly elutes RNA.
RNA will remain in the silica at a high pH buffer and elute only when all the contaminants and cell debris are removed. The final elution is collected in another tube. Spin-column extracted RNA is highly pure.
Green and Sambrook (2019) published a protocol using DNase in RNA extraction. Genomic DNA is a common contaminant in RNA and causes downstream result failure or false-positive results. The use of DNase removes genomic DNA traces from the RNA sample.
Ñique et al., (2021) explained the cost-effective RNA extraction method using the proteinase K which is crucial to degrade proteins during the extraction of RNA. The use of proteinase K in the RNA extraction removes nucleoproteins, nucleases, other enzymes and proteins which affect the purity of RNA.
However, the role of proteinase K in RNA extraction was first described by McGookin in 1985. Other than this, there are numerous chemicals, solutions and protocols available to extract RNA, stabilize and store RNA.
Wrapping up:
Unlike DNA extraction, the RNA extraction technique is tedious, laborious, time-consuming and contamination prone and hence one should have substantial experience and expertise.
For a conventional method, phenol and guanidinium thiocyanate are two important chemicals, although the function of other chemicals is crucial too. For pathogen RNA, ready-to-use kits are advised as it adds ease and speed to experimentation.
The goal of this article is to give you a brief RNA extraction process. We are planning a whole series on it. In upcoming articles, we will explain various protocols, optimization tips, the role of chemicals and other related information.
I hope this article helps you.
Resources:
- McGookin R. RNA extraction by the proteinase k method. Methods Mol Biol. 1985;2:109-12. doi: 10.1385/0-89603-064-4:109. PMID: 21374178.
- Green MR, Sambrook J. Removing DNA Contamination from RNA Samples by Treatment with RNase-Free DNase I. Cold Spring Harb Protoc. 2019 Oct 1;2019(10). doi: 10.1101/pdb.prot101725. PMID: 31575796.
- Ñique AM, Coronado-Marquina F, Mendez Rico JA, García Mendoza MP, Rojas-Serrano N, Simas PVM, Cabezas Sanchez C, Drexler JF. A faster and less costly alternative for RNA extraction of SARS-CoV-2 using proteinase k treatment followed by thermal shock. PLoS One. 2021 Mar 24;16(3):e0248885. doi: 10.1371/journal.pone.0248885. PMID: 33760876; PMCID: PMC7990203.
- Chomczynski P, Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem. 1987 Apr;162(1):156-9. doi: 10.1006/abio.1987.9999. PMID: 2440339.