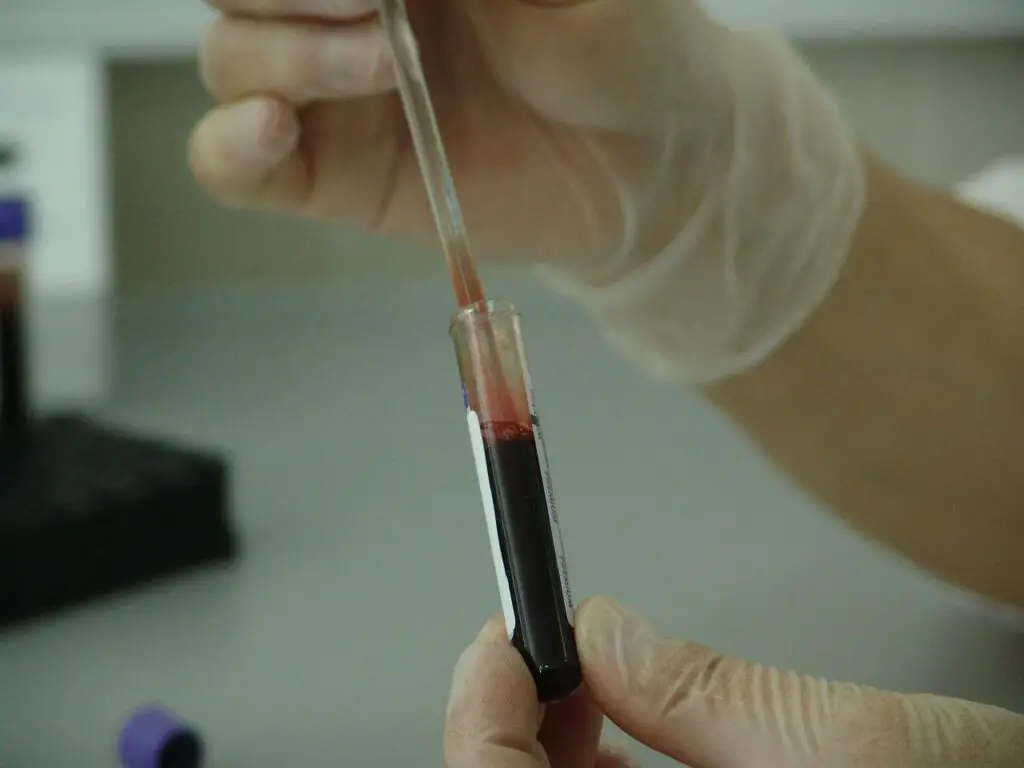
“RIN is an RNA Integrity Number showing the value of how intact the RNA is. RIN score >8.0 is ideally recommended for high throughput downstream processing.”
Unlike DNA, RNA is a highly sensitive nucleic acid and degrades rapidly by RNase. Other factors like heat, contaminated chemicals and inadequate buffering conditions also affect the intactness of RNA and lead to degradation.
So it needs special treatment without failure otherwise, we will lose our precious nucleic acid. The highly sensitive nature of the RNA makes the entire RNA extraction process more difficult. I, however, made it nearly easy for you in our previous article, you can read this article for more information: 21 Things to know for Effective RNA Extraction.
To understand the concept of RNA integrity, first, know why it is different from DNA and why needs special care. DNA is an intact double-stranded molecule but a total RNA is a cocktail of many different types of RNAs like tRNA, mRNA, various rRNAs and smaller non-coding RNA.
Moreover, it’s single-stranded and more fragile and the presence of RNase everywhere makes overall things difficult for RNA. With this, post-extraction processing, like adding RNA stabilizers and cryopreservation is also required to store the intactness of RNA.
So cumulatively things aren’t always in our favor while RNA extraction. On the other side, RNA has especially been used in gene expression, transcriptomics and RNA sequencing experiments, such assays are costly, time-consuming and labor-expensive.
Furthermore, the tissue-specific nature of RNA even makes things more difficult as tissue samples are usually less, precious and can’t be re-sampled. So overall, it’s a “one-shot wonder.”
We need to isolate a good quality RNA, which is indeed intact and to assess the integrity/intactness, the RNA integrity number is scored.
This article will mainly answer what the RNA integrity number is, why it is important, and how to calculate it. This guide is all about the RIN which certainly would help your RNA extraction endeavor.
Stay tuned.
Key Topics:
What is RIN (RNA Integrity Number)?
It’s an industry standard and state-of-the-art RNA integrity, intactness and quality assessment tool for gene expression, sequencing and microarray analysis.
The RNA Integrity Number is a numerical value that shows RNA integrity/intactness on a scale of 10 to 1 which is excellent to worse, respectively. 10 shows excellent RNA integrity while 1 shows worse. Technically, it works on microfluid chip-based separation that allows us to visualize the bands as well as measures the integrity value, computationally and by using a mathematical model.
In past years, scientists measure the RNA integrity by looking into the 260/280 ratio, electrophoretic analysis and banding pattern of 28S and 18S rRNA fragments which would be 2:1 in the ratio. The purity of RNA, although can’t tell us more about the RNA integrity. On the other side, the 28S:18S rRNA ratio is a kind of physical examination by electrophoretic analysis and is certainly more inconsistent.
The technique was based on the conventional gel electrophoresis principle that uses the EtBr stained RNA. When RNA migrates into a gel, by looking into the banding pattern, the integrity of RNA can be determined. As aforementioned, two major marker bands are 28S and 18S rRNA.
Unfortunately, the technique requires a huge amount of RNA as a starting material, manual assessment and expert personnel. Scientists have to look into the gel to study the integrity, thus, results vary from lab to lab.
Also, this technique wasn’t robust, reproducible, accurate and consistent so over a period, needed replacement. RIN evolution by capillary gel electrophoresis solves almost all the shortcomings enlisted here. Let us start wit the principle.
Principle of RIN assay:
The methodology of RIN is actually developed by Agilent technology whose mathematical modeling is based on the Bayesian adaptive learning method. In principle, a smaller volume RNA sample runs in smaller-sized capillaries and separates in capillary electrophoresis.
Note that the CE has the potential to distinguish smaller-sized nucleic acid fragments, sharply and accurately. The electropherogram construct shows RNA fragment bands as peaks by fluorescence emitted against the length or time of migration on the X and Y axis, respectively.
Noteworthy, the intensity of fluorescence emitted is directly proportional to the peak size of a particular RNA species in the graph. Thus more the fluorescence emitted, the higher the intensity of the band and the larger the RNA fragment. Take a look at the image below.
To know more about the CE, you can read this article: Capillary electrophoresis of DNA- Explained
Understanding the electropherogram of RNA:
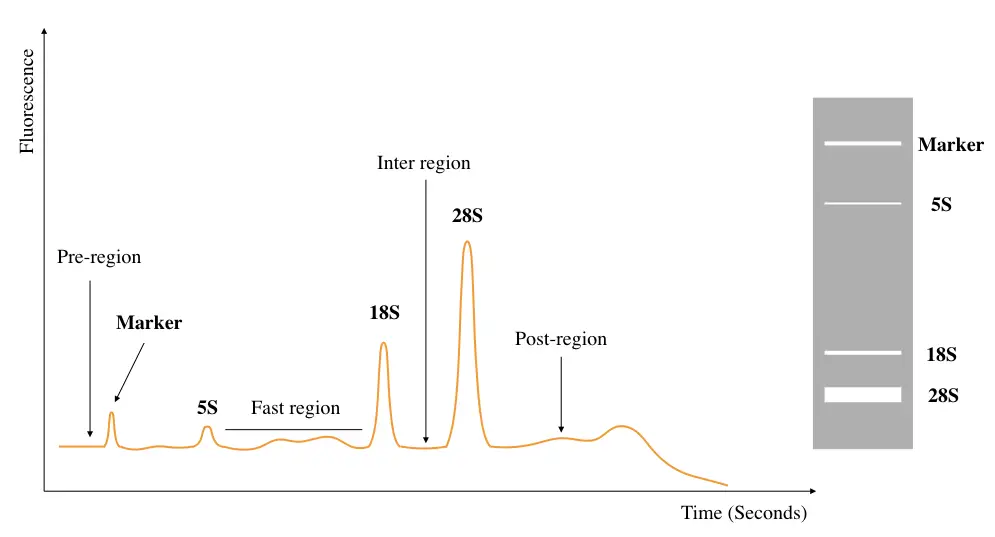
The figure above depicts a typical electropherogram for RNA integrity analysis, it’s a kind of hypothetical representation! Not a real image. To evaluate the RIN score, parameters taken into consideration are bands of 28S, 18S and 5S rRNA, marker and Fast region. All parameters are there in the graph.
Also, it is important to note that this electropherogram or RIN method by 28S:18S rRNA analysis is only applicable to mammalian RNA. To determine the RIN score of prokaryotic RNA and other organisms, different parameters are set.
Now coming to the point, 28S rRNA emits more fluorescence, travels less distance in a gel and shows a prominent large (doubled than 18S rRNA band) peak in the graph. As it has a large size, it takes more time to migrate to CE.
Studies show that the 28S rRNA is more prone to degradation than other rRNA species. Thus, its peak size is an important marker here, which should be doubled in comparison with the 18S rRNA peak, visually.
Next, the 18S rRNA which is smaller than the 28S remains ahead, emits less fluorescent and shows a smaller peak than the 28S rRNA. A small peak of 5S rRNA species is also found in the graph as this RNA species is also present in mammals.
A marker is nothing but, technically, a tiny peak that shows the amount of degradation, meaning, the smaller the size of a marker, the less the RNA degradation is! The ‘fast region’ is a distance between the 5S and 18S rRNA whose distance is ideally fixed in the computational analysis. change in this distance indicates RNA degradation.
So this is a small overview of the RNA electropherogram. Now see in which conditions we can demonstrate RNA degradation and reduced RNA integrity.
- 28S rRNA peak too small, smaller than 18S or less prominent peak.
- Large 18S rRNA peak or very small like the 5S.
- Absence of 5S rRNA peak.
- Too large marker region.
- Absence of 28S or 18S rRNA region.
- Absence of both or any of the two rRNA species.
- To larger or small fast region.
Pre-region, post-region, inter-region and precursor region are other parts of the electropherogram of the 2100 bioanalyzer instrument.
Very important note:
Not only the 28S or 18S but all other RNA species run in the capillary gel are taken into account for the RNA score calculation. This makes the RIN score more accurate, trustworthy, reliable and usable than only the two rRNA species ratio.
Agilent described that anomalies or errors may occur during the reporting which are of two types; more critical and non-critical. For example, the peak after the post region is non-critical while the peak in the fast region, between the marker and 5s region and any peak between the 18S and 28S rRNA, are considered critical errors.
Related article: 5 Interpretation Manifests RNA Degradation.
How RIN is calculated?
As aforementioned, all the above factors enlisted; the peak of 28S, 18S and 5S rRNA, marker region, an aberration in the fast region, mathematical model and other computational analyses are taken into account by the software to calculate the RIN value. Ideally, it is assessed between 1 to 10.
No mathematical calculation, equation or manual ways are available to calculate the RIN score.
Why RIN is important?
Gene expression, RNA sequencing and microarray studies need RNA as starting material with an excellent integrity score. Such experiments are costly and labor-expensive and thus accurate pre-sample preparation is required. If the sample fails, it eventually fails the reaction and researchers have to bear the cost.
On the other side, RNA is more degradative by RNase. RIN value shows how intact the extracted RNA is. RIN value stamps the sample to use further.
What information does RIN provide?
On a scale of 1 to 10, RIN gives us an idea of how intact our RNA is. RNA has many enemies and is itself more delicate to operate. RNase enzyme, present everywhere from the extraction bench to pipette and on our hands to in environment which destroys RNA.
So starting from extraction to elution, it remains under the threat of RNase. With extraction, RNA protection is important too. Various techniques and chemicals are used to protect RNA intactness and integrity but still, it may get deteriorate and eventually fail our downstream assay.
RIN score shows whether the RNA is useful for further process or not. Henceforth, it, tentatively, save our experiment, cost, time and manpower.
What is the acceptable RIN score?
The higher the RIN score, the exceptionally intact the RNA is! Meaning, that RIN 10 shows highly intact RNA while RIN 1 shows greatly degraded RNA. The RNA integrity number between 8 to 10 is considered as highly intact RNA, RIN between 1 to 5 is considered as low intact, poor quality or highly degraded while RIN score between 6 to 8 is considered as moderately intact or low-degraded RNA. So the acceptable value is ideally, between 7 to 10.
Note:
The Agilent Bioanalyzer 2100 analyzes the entire electropherogram to develop the RIN score.
RIN score for gene expression, RNA sequencing, and microarray:
Not all molecular genetic assay needs 10 RIN RNA, and we can’t get it every time, in fact, most of the time. Some assays are manageable with a low RIN score while some require an exceptionally high RIN score. Here is the list of assay and RIN scores required.
- Gene array: RIN scores 6 to 8
- RT-qPCR: RIN scores 5 to 6
- qPCR: RIN score >7
- RNA sequencing: RIN score between 8 to 10.
- Microarray: RIN score between 7 to 10.
RIN score information provided by the machine:
Table: Standard information provided by Bioanalyser 2100 during RNA integrity assay.
| Parameter | Sample results |
| RIN score | 8.1 |
| RNA area | 1400.00 |
| RNA concentration (ng/μL) | 1370 |
| 28S/18S rRNA ratio | 1.4 |
| 18S rRNA | |
| Start size (nt) | 1529 |
| End size (nt) | 1960 |
| Area (%) | 24.7 |
| Total area (%) | 14.5 |
| 28S rRNA | |
| Start size | 2997 |
| End size | 4101 |
| Area (%) | 35.5 |
| Total area (%) | 20.5 |
Factors affecting RIN:
A good or bad RIN score depends on many factors starting from selecting extraction methods to how to manage the extract. Several common factors are discussed here.
- RNA extraction protocol:
What quality and quantity of RNA we will get solely relies on the RNA extraction protocol we use. Some protocols can’t inactivate RNase while others can, thus the protocol selection has been an integral part that influences the RIN score.
I personally, recommend using the Trizol protocol, you can read the entire article on this here: Effectual Trizol RNA extraction protocol.
- Tissue type:
Tissue type is yet another factor in this. Some tissues give exceptionally good RNA while some can’t. For example, it is very difficult to isolate RNA from highly nuclease-rich, parafilm-embedded, brain or plant tissues. While isolating RNA from a cell suspension is comparatively easy. Such hard-to-exact tissues negatively impact the RIN score.
- RNA elution process:
The elution process is also as important as other factors. After eluting the RNA, it is important to add the RNA stabilizer and cryopreserving it to maintain its integrity and intactness. If not stored well, RNase may attack which eventually affects the RIN score.
- The concentration of RNA extract
As per the report of the Agilent, the concentration of RNA has crucial implications on RIN Score. Usually, RNA concentration >50 ng/µL gives a uniform RIN score, however, a concentration < 25 ng/µL isn’t advisable to score RIN.
- An instrument for validation:
Microarray assessment is a standard RIN value evaluation tool, that helps scientists to decide the threshold value, however, it largely depends on the microarray system. So it may change the RIN score.
- Sample shipping:
Sometimes, nonstandard sample shipping changes the RIN. RIN should be measured pre- and post-shipping of RNA samples.
Note
RIN needs only a 1 µl sample to measure RNA integrity.
In this article, I am mentioning special thanks to Leussa A. (Ph.D.). Her article on LinkedIn is amazing and comprises all the necessary information on RIN. I got inspiration from her to write this article. Her article is here: What value of RIN is considered suitable/ sufficient for RNA seq analysis.
Wrapping up:
RIN score isn’t, indeed, a uniform standard to evaluate RNA integrity, however, certainly is accurate in results. But as a scientist, we need to develop our own threshold RIN value for our needs and purpose.
For example, some research suggests that a RIN which can’t form for microarray or sequencing works excellently for RT-PCR. Still, before deciding what to do with our RNA extract depending upon the data from literature and manufacturer, we should have to manually validate our own.
It is possible, perhaps, you can get excellent results in qPCR at RIN 6 or 6.5 but others can’t, in that case, you can save your sample. Nonetheless, over time, you get an idea of what RIN works for you and what not!
Best of luck.
Sources:
- Gene quantification ebook by Agilent Technology.
- Lightfoot S, Salowsky R, Buhlmann C. RNA integrity number: towards standardization of RNA quality assessment for better reproducibility and reliability of gene expression experiments. Breast Cancer Res. 2005;7(Suppl 2):P7.05. doi: 10.1186/bcr1197. Epub 2005 Jun 17. PMCID: PMC4233618.
- RNA Integrity Number (RIN)- Standardization of RNA quality control by Agilent technology.