“RNA extraction is a process to isolate RNA from a cell for gene expression studies and pathogen identification.”
RNA extraction is a nucleic acid separation technique to collect various types of RNAs, however, it isn’t as straightforward as DNA separation. The highly sensitive nature of ribonucleic acid makes things hard for the lab personnel.
Unlike DNA, stabilizing, protecting and storing RNA are three crucial factors in the process that ultimately decide the fate of transcript and transcriptomics analysis. Again, unlike DNA, RNA needs special care, continuous monitoring and high-end safety.
High yield and ultrapure RNA is the key to get success during various RNA-dependent experiments such as qPCR, RT-PCR, transcriptomics, microarray and RNA sequencing analysis.
So only learning how to isolate RNA isn’t enough for you, one should have insight information and optimization range to get a decent RNA output. In the present article, I will share with you some of the tips and my personal optimization options for effective RNA isolation.
“RNA is more labile, weak and prone to degradation than DNA. The reactive C-OH (hydroxyl) group present in the RNA induces rapid formation, breakdown and reuse of RNA strands in a cell.”
Key Topics:
21 Things to know for Effective RNA Extraction
1. Sample collection remedy:
The sample collection process is an important step in RNA processing. It’s always challenging to collect a contamination-free and adequate quantity of samples. Leave the sample collection part to an expert.
Maintain a cold chain through the process and perform extraction as soon as possible. Snap-freeze the sample if required. Also, store the sample separately to avoid RNase contamination.
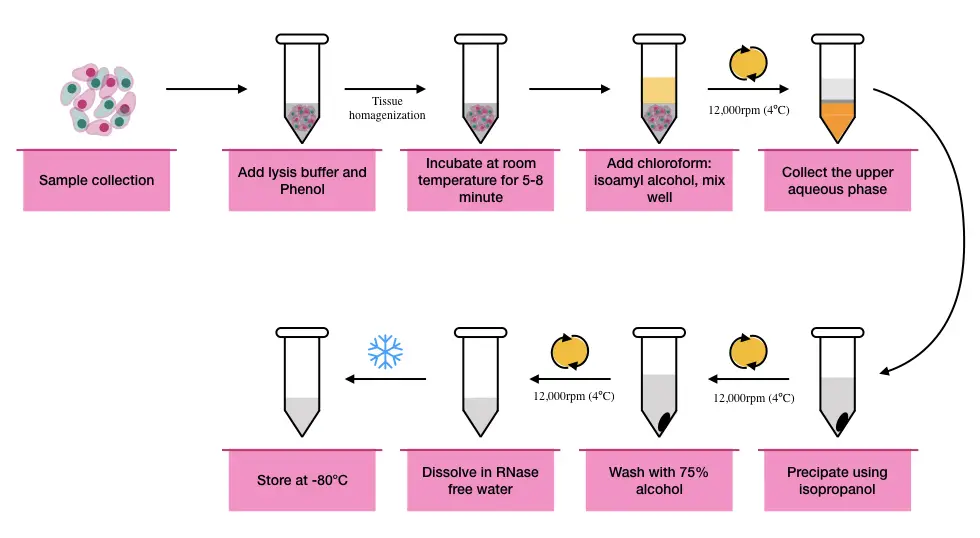
2. Type of RNA required:
The very first thing in your endeavor to extract RNA is to decide what type of RNA you need in your experiment, meaning, understanding the requirements of your experiment. mRNA, tRNA and rRNA are three major types of RNA that decide the fate of a gene.
While siRNA, miRNA and other non-coding, small and single-stranded RNAs have a role in gene expression, regulation and protection against microorganisms. Hence, for post-transcriptional modification studies, you need microRNA whereas for transcriptional activity studies you need only mRNA.
Why it is important! The reason why ‘types of RNA’ are important is that every RNA is different in size and has a unique function. The table below comprises different types of RNAs, their function and size.
| Type of RNA | Size | Function |
| Total RNA | – | Gene expression, gene regulation, PTM and protection |
| mRNA | >200nt | Forms protein from the DNA |
| tRNA | 70-90nt | Transfer amino acids to the growing polypeptide chain |
| rRNA | >200nt | Forms various ribosomal subunits like 5S, 18S or 28S rRNA. |
| microRNA | Up to 22nt | Post-transcriptional modifications. |
| Smaller non-coding RNAs | 20 to 200 nt | Various biological and cellular activities |
Once you have decided which type of experiment you want, thereby the type of RNA, you have to select an RNA isolation procedure or protocol on that basis. And here is another point on how to choose the RNA extraction protocol.
Related article: RNA: Structure, Types and Function.
3. Pick the RNA extraction protocol:
Every RNA is different in size, and therefore there are various RNA extraction protocols and kits available. Although the modular method for total RNA separation, like the Phenol-chloroform-based technique, is available; downstream processing is required to separate a specific type of RNA, however.
This adds an extra step in the process that eventually elevates risks for contamination and losing RNA integrity, so a specific type of RNA extraction kit or protocol is advised.
For instance, to isolate only the mRNA from the total RNA a specific type of oligo(dT) column is used in which the poly-A tail of mRNA (which is the characteristic) binds with the oligo (dT) and remains in the column, after washing.
In the next step, the oligo (dT) is washed off and pure mRNA is collected in another tube. The same process is followed in the magnetic bead-based separation technique in which instead of a column, oligo(dT) magnetic beads are used.
Depending upon the size of each RNA, various magnetic bead-based techniques or spin-columns are available in market that have the power to separate a specific type of RNA only. Note that magnetic bead-based separation is rapid but costly.
For high nuclease, fat and secondary metabolite-containing tissues like liver, brain/adipose and plant tissue, phenol-based RNA extraction is advised.
4. Quantity of starting tissue:
“Less is more.”
A 20 to 50gm tissue sample is good for homogenization of tissue for RNA extraction. However, higher and lesser amounts of starting material may give abrupt RNA output. A large quantity of tissue unnecessarily creates chaos in the experiment.
The sample may come out of the tube, fall on the working bench or contaminate other things. Scientifically, a more than sufficient amount of tissue prevents chemicals from lysis buffer to work properly.
The tube or spin-column when overloaded with a sample, can’t work effectively, eventually, it leads to reduced yield and quality of RNA. Starts with a small volume of tissue. Keep in mind that tissue type too.
5. Which tissue you are using:
Tissue to tissue the expression of transcriptomics differs and that is what we want to study. And in addition, the structure, composition and amount of RNA also vary. Such factors are also taken into consideration while performing RNA extraction.
Different tissues need a different treatment and so the protocol may vary. For example, for hard tissues like plant cells, lab personnel need to do extra effort in tissue lysis.
CTAB, SDS like detergents and other chemicals are used to do the job.
While processing the tissues like the liver with a high nucleic acid content, aggressive use of DNase is advised.
6. Creating RNase-free working atmosphere:
RNA extraction is a sensitive molecular biology procedure that needs special care. The predominant prevalence of the enzyme RNase makes the entire procedure more difficult. The RNase enzyme digests the RNA molecule.
Make the entire surrounding of your work free of RNase. Several suggestions regarding that are here,
- Wash off all the utilities with RNase-free water and autoclave them.
- Clear the working bench with RNase-free water.
- Use clear and sterilized gloves and a PPE kit during the experiment.
- Maintain sterility during the entire procedure.
I am planning to write a whole article on this topic on how to avoid RNase contamination during RNA extraction.
7. Using a specialized chem-combo:
Inactivating only the RNase isn’t enough here. We need extra chemicals or reagents to remove other things like DNA or protein. Such treatments are very crucial for some tissues.
For example, for plant tissue, proteins while for liver tissues DNA removal is so important. Enzymes like DNase and proteinase K are employed to destroy DNA and protein, respectively from a tissue. So to make the RNA isolation procedure more effective, such optimizations are advised.
For hard plant tissues, CTAB-like chemicals are used to remove the polysaccharides from the samples.
8. Be careful about accidental RNase contamination:
“You did it and you will never know it.”
Accidental RNA contamination is so common among beginners. Because sometimes they contaminate things by a minor mistake and they even don’t know how it occurred.
Each step of RNA isolation would be performed accurately. Repetitive use of gloves, tips, tubes or dipping tips to different reagents contaminates RNA easily.
Also touching unsterilized areas, tips or pipette also ruin the entire experiment. Such mistakes are unnoticed but are so important, especially during RNA isolation.
RNase is present everywhere, touching contaminated or untreated surfaces will mistakenly contaminate the sample with RNase. And you know what will happen at the end.
Furthermore, repetitive use of extraction reagents in open environments, aerosols created by blank pipetting and previously shared working place or laminar hood for microbiology experiments also fail RNA extraction.
Note:
Document everything related to the isolation experiment to minimize the repetitive chances of RNA extraction failure.
9. Be equipped:
“Be well-equipped to win the battle.”
Skin, hair, skin oil and perspiration are common sources of RNase therefore one should have to equip themself before the actual extraction process. Use pre-sterilized utilities which must be treated with nuclease-free water prior.
Clear the working place, decontaminate it, and if possible do fumigation one day before. Wear a sterilized mask, gloves, mouth & head cap, and PPE kit.
Important note:
Use certified RNase-free utilities.
10. Store safe, with integrity & stability:
The single-stranded RNA is unstable and prone to degradation. As its enemy- RNase present everywhere, it remains always at a risk of losing integrity. So RNA integrity should be evaluated first. This article will help you to learn more: What is RNA Integrity Number (RIN)? and Why It Is Important?
Usually, it is stored at -20°C for short-term and -80°C for long-term use. Ethanol bath, the use of dry ice and liquid nitrogen are several options to effectively store the RNA for long-term use. RNA stabilizers are also available commercially in the form of a buffer which protects the integrity of RNA.
11. The use of liquid nitrogen:
Liquid nitrogen is immensely important for the RNA extraction process to store and process the sample. Adding liquid nitrogen snap freezes the sample and stores it at -80°C temperature. Such practice stores the cellular activities as it is and the integrity of the RNA.
Besides, it prevents tissue contamination and alterations in gene expression in vitro. RNA should be extracted as soon as possible but by adding liquid nitrogen, researchers can process the sample later, even after a few days.
Interestingly, liquid nitrogen will homogenize hard tissues like plant leaves or bark finely. Using mortar and pestle, grind tissues well by adding liquid nitrogen slowly. Wait until the LN evaporates completely to add a lysis buffer otherwise it becomes ice. In short, liquid nitrogen is used to store and lyse the tissues.
Caution:
Care must be taken while handling the liquid nitrogen as it can freeze & damage tissues, freeze eyes or other fluids and do permanent damage.
Some tips:
- Add a detergent to the lysis buffer for tissues with high lipids.
- Use guanidine or urea to deactivate cellular RNase.
- Perform all the procedures of RNA extraction under a cold environment to minimize RNase activity.
- Maintain the required pH during the entire process.
- RNase are ubiquitous in nature even by chemical treatment and autoclave, it remains partially active. Use certified RNase-free utilities and chemicals.
- Plasticware is pre-washed with 0.1M NaOH, 1mM EDTA and DEPC or RNase-free water.
- Heat-proof glasswares are first washed with detergents, autoclaved and baked at 180°C for an hour.
Pro-tips
19. Use pri-chilled pipettes, tips and glasswares to suppress the activity of RNase.
- DNA can be preserved in ethanol or any alcohol in the form of precipitate but RNA can’t as it degrades in alcohol.
- Maintain the cold chain throughout the process, starting from sample collection, to processing and storing.
Wrapping up:
RNA extraction needs mastery in molecular genetics but by following some routines, tips and optimizations, you can achieve expertise in it. However, as you do trial and error, you will learn new things and correct recipes for isolation.
Lastly, from my side, you have three most important pieces of advice for all beginners: store and lyse the tissues well, stabilize the RNA and maintain an RNase-free atmosphere. Strictly follow these three points and trust me you always get a good amount of RNA.
I hope this article will help you in your genetic learning.