“RNA sequencing is a high throughput next-generation sequencing method used to analyse gene expression and transcriptomics studies.”
Ribonucleic acid is a type of nucleic acid majorly involved in the gene expression, gene regulation and coding/decoding of information.
mRNA- messenger RNA is a coding sequence of a gene implicated in the synthesis of protein, approximately, 4% of RNA pool consists of mRNA while rest are non-coding RNAs.
Though approximately 90% of RNA are non-coding, those are important for performing various functions.
For example,
- tRNA- transfers the amino acid to the ribosomal site, while the translation.
- rRNA- ribosomal RNA helps in translation.
- microRNA- gene expression and regulation of gene expression.
- siRNA- protects a cell from the exogenous RNAs.
An mRNA is a single-stranded nucleic acid transcribed from a gene and from which the protein is translated.
The mRNA or transcript contains only the coding sequences from a gene. Thus it only possesses the sequences which are required in the protein formation.
Henceforth, by analysing an mRNA, the expression pattern of the related gene can be determined.
Also, total RNA examination gives us an idea about which genes are linked to the protein formation and which are not.
Means, the amount of total RNA, mRNA and ncRNA can be determined by studying the total RNA.
Transcriptomics is the study of the entire mRNA- often called as transcriptomes or total RNA present in a cell or cell population.
By analysing the transcriptome of a cell, the entire pool of the gene expression can be examined. One of the important techniques used in transcriptomics studies is RNA sequencing.
The transcriptome of an organism is larger, complex and more uncertain, unlike the genome.
The transcriptome varies in different condition and a different environment. Lifestyle, diet, exercise, environment, conditions and other external factors play a significant role in transcriptome regulation.
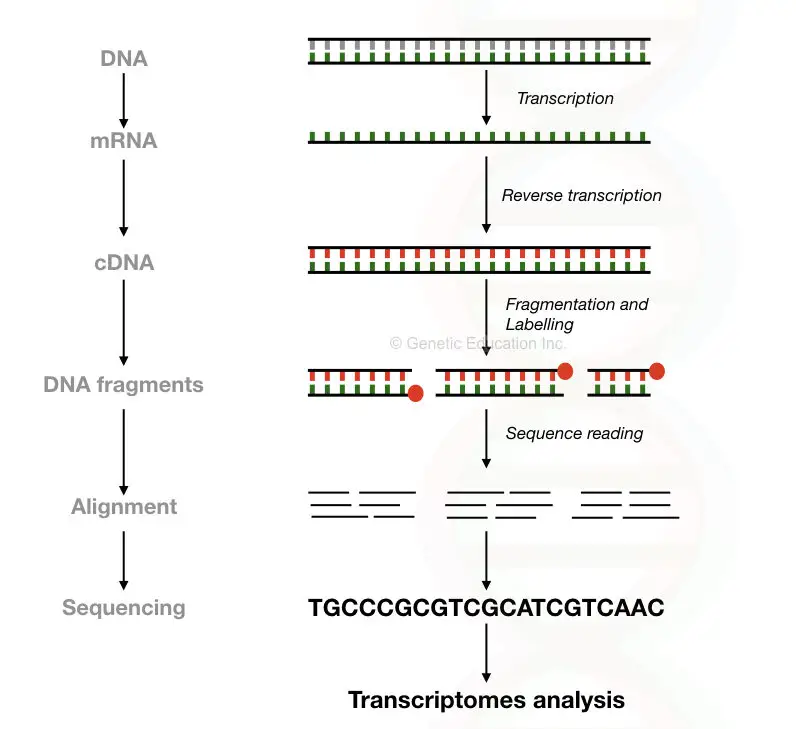
Here in the RNA sequencing, the cDNA synthesised from the mRNA is sequenced and quantified in a sequencer. In the present article, we will discuss the RNA sequencing only but before that read our previous article on transcriptomics: What Is Transcriptomics?
Key Topics:
Principle of RNA sequencing:
RNA sequencing is a next-generation, high throughput RNA sequencing and quantification method used for studying the transcriptomics and gene expression.
A cDNA is constructed from total mRNA through the process of reverse transcription and fragmented. Simultaneous, adaptor ligation and library preparation are practised before doing sequencing.
The sequencer reads and quantifies the cDNA complementary to the mRNA.
Steps in RNA-seq:
- RNA isolation
- cDNA synthesis
- Adaptor ligation
- Library preparation
- DNA fragmentation
- Sequencing
- Downstream applications
RNA isolation:
The first step in the RNA sequencing is the isolation of total RNA, mRNA or ncRNA for the experiment.
Isolating RNA is a tedious process in comparison with DNA isolation. The RNA can be breakdown easily. Also, the chance of contamination is high in RNA isolation.
Thus extra care must be taken while isolating RNA and expert hand in required to do so.
High yield pure RNA is must require for RNA sequencing.
The purity and quantity of the RNA are measured using the Nanodrop or qubit.
Note: for isolating mRNA from the total RNA pool, one additional step is required, using the oligo dT specific column in the purification or final step of elution, total mRNA can be isolated from the rest of RNAs.
Now our RNA sample is ready for the next step.
Reverse transcriptase PCR:
Another innovative set up for RNA sequencing is to do reverse transcriptase PCR in which the RNA is reverse transcribed into DNA.
Using a special type of polymerase known as DNA reverse transcriptase, the cDNA is synthesised from the mRNA.
Learn more on RT PCR here: Reverse transcriptase PCR.
Second strand cDNA synthesis:
After the cDNA synthesised from the mRNA, the second strand DNA synthesis is must required. For that, PCR is performed using the normal Taq DNA polymerase in conventional PCR.
The Polymerase adds dNTPs to the growing DNA strand using the primer set.
Library preparation:
The first step in the library preparation or NGS library preparation starts with the fragmentations.
Using the special type of restriction endonucleases whole set of ds cDNA is fragmented for the NGS.
Note:
The library preparation varies from platform to platforms as some fragments mRNA before performing reverse transcriptase while some perform it later after the preparation of cDNA.
In the last step of the library preparation, the ends are ligated with the adapter sequences and amplified for repairing the ends. dA-tailing is optional in case of mRNA library.
Library purification:
The entire library is purified using the ready to use DNA purification kit.
In the last step, it is very crucial to purify the entire fragment library for that the concentration of the library is assessed using the quantitative PCR or bioanalyzer.
Note:
For a beginner I think, library preparation and DNA fragmentation is very hard to understand, so we think, we should write an entire article on Genomic DNA fragmentation, library preparation and its importance in NGS.
Anyway let’s move ahead,
In the next step, the sample of fragmented DNA is sent for sequencing.
DNA sequencing:
Now the sample is sequenced in the high throughput NGS machine which reads the sequence as well as quantifies the nucleic acid as well.
The chemistry behind sequencing the cDNA is depends on which platforms we are using, although, the widely used method is the use of fluorescence chemistry.
In the sequencer, individual fragments are “read” separately and sequenced. The final data are sent to the bioinformatics lab for post-sequencing analysis.
Different types of RNA and its function:
| Abbreviation | RNA types | Function |
| mRNA | Messenger RNA | Codes for protein |
| tRNA | Transfer RNA | Transfer amino acid to the site of translation. |
| rRNA | Ribosomal RNA | Catalyse the translation reaction |
| miRNA | microRNA | Gene regulation |
| siRNA | Smaller interfering RNA | Gene regulation and maintaining gene expression. |
| LncRNA | Long non-coding RNA | Transcriptional regulation and epigenetic regulations. |
| snRNA | Small nuclear RNA | Helps in mRNA splicing and related functions |
| snoRNA | Small nucleolar RNA | Helps in RNA nucleotide modification |
| piRNA | Piwi-intercalating RNA | Function in defence against transposon; transposon defence system. |
| scaRNA | Small Cajal body-specific RNA | Also helps in nucleotide modifications (a type of snoRNA). |
| shRNA | Small hairpin RNA | Synthetic RNA molecule helps in gene regulation and controlling gene expression. |
Transcriptome data analysis:
One of the tedious jobs, as we already discussed in the previous article is the transcriptome data analysis and interpretation of results. NGS data are huge and more complex.
Frankly speaking, teaching data analysis of transcriptomics is not possible, one should have to take hands-on practice to learn, still, I will try to teach you what is next in this process.
Different fragments are sequenced in the machine and data are collected. In the next step, based on the flanking region of the fragments, the so-called “contings” are arranged for getting splice variants.
In the next step, the transcriptome is compared with the reference sequence or can be assembled de novo.
Different RNA sequencing methods:
Whole transcriptome sequencing:
The whole transcriptome of a cell or tissue- all RNAs (mRNA, tRNA, rRNA, sncRNA, microRNA and siRNA) are sequenced and quantified. In other words, we can say, Both novel, as well as known transcripts, can be sequenced.
For doing it, the entire set of RNA is extracted from the tissue sample.
Target RNA sequencing:
A gene-specific, cluster of gene-specific, a pathway-specific or disease-related transcriptomes are sequenced in the target-specific RNA sequencing.
Target RNA sequencing is more affordable and accurate than the whole transcriptome sequencing. Moreover, less amount of RNA sample is required for doing the experiment.
Small RNA sequencing:
One of the most emerging technique in RNA-sequencing is to study the smaller non-coding RNA of a cell because those are associated with so many functions in a genome.
Smaller RNA molecules such as miRNA, siRNA and piRNA can be quantified and sequenced through the NGS based RNA sequencing method for various applications.
mRNA sequencing:
Sequence the entire mRNA transcript using poly(A) tail selection used for gene expression studies. Using mRNA sequencing known as well as novel transcript alteration can be detected.
Advantages of RNA sequencing:
One of the pivotal advantages of RNA sequencing is the no need for prior sequence information. As the entire process is not based on the probe-based chemistry, prior sequence information for designing the probe is not required here.
More accurate and sensitive gene expression study can be done.
Broader dynamic range.
Capture both known as well as novel alterations of the transcript, even if no sequence information is available.
Even though no reference sequence information is available, RNA sequencing can be applied for any species.
Related: Synthetic Genome- Methods, Applications And Challenges.
Conclusion:
Conclusively we can say, that the RNA seq method is more advanced and accurate as compared to the microarray. Also, de novo mutations can also be discovered using it.
Sources:
Kukurba KR, Montgomery SB. RNA Sequencing and Analysis. Cold Spring Harb Protoc. 2015;2015(11):951–969. Published 2015 Apr 13. doi:10.1101/pdb.top084970


