“Nucleic acid extraction is a technique to separate or isolate nucleic acid from a cell. A pure NA is an utmost requirement for downstream molecular analysis such as PCR amplification, sequencing or microarray.”
Two types of nucleic acid are DNA and RNA. DNA is predominating the entire living kingdom while RNA is present in a few types of viruses. Though RNA in varied forms is present in mammals, only DNA is present as “genetic material”.
The usual location of nucleic acid is a cell nucleus but some in a circular or linear form are also found in the cell cytoplasm. For instance, plasmid DNA in bacteria, mitochondrial DNA in humans and plastid/chloroplast DNA in plants. Such nucleic acid is popularly known as organelle DNA.
Nucleic acid is commonly abbreviated as NA which allows researchers to study DNA sequence, gene or gene expressions. DNA grants genetic studies while RNA grants epigenetic and gene expression studies.
However, for any purpose, we as a researcher need pure NA and we know that extraction of nucleic acid is a tedious and difficult task, indeed. So, I’m writing this article for beginners as well as for researchers who need advanced knowledge, optimization tips and protocols.
This article is a kind of a course, you can say or a collection of all the articles I already have written on various techniques to separate NA, however, it’s not just an overview. The article actually comprises an in-depth explanation and review on the entire Nucleic acid Extraction topic.
I hope this will boost your knowledge of such techniques.
Stay tuned.
Key Topics:
Basics of nucleic acid extraction:
Our blog Genetic Education Inc. is all about genetics and thus somewhere in some articles, you may find thorough information on the nucleic acid. But in the context of ‘how to extract nucleic acid,’ we will have to re-state information here.
In 1986, Miescher F isolated nucleic acid for the first time in history and named it “nuclei” but was actually the NA. Soon after, many new discoveries in NA extraction were conducted, optimized protocols were generated and new techniques and technologies for nucleic acid extraction were developed.
Theoretically, the process to isolate NA sounds quite easy. Just by breaking the cell wall or cell membrane and nuclear membrane, we get our nucleic acid. Nonetheless, things aren’t ‘in a straight line’ in molecular genetics.
In the practical approach, the entire process comprises many separation steps, chemical preparation, centrifugation, incubation, and freeze and thawing. The principle of isolation is as follows.
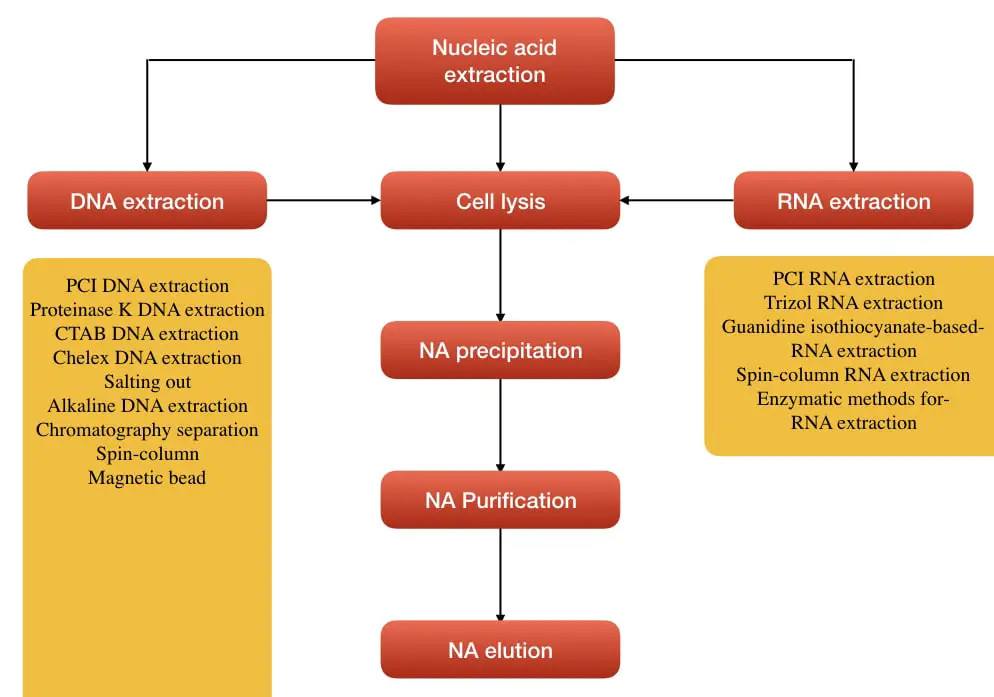
“Breaking of the cell wall or cell membrane and nuclear membrane, precipitating and removing proteins and other cell debris, purifying the nucleic acid, precipitation and elution. Note that the process remains almost similar for both DNA and RNA isolation.”
While selecting a technique for NA extraction there are several things one should keep in mind that eventually help for better isolation.
Things to be considered for NA isolation:
Selection of isolation technique- if the technique is capable of powerful isolation or not.
Effective cell wall or cell membrane disruption- if the technique can effectively break the cell membrane components or not.
Nuclear membrane disruption- If chemicals of the technique can effectively break the nuclear membrane or not.
Removal of cell debris- if the solutions and chemicals used in the separation protocol are capable enough to remove all other cellular debris or not.
NA purification- If solution or chemicals used in the protocol can do effective purification or not.
NA precipitation or elution- If the protocol comprises solution or chemical for NA precipitation or elution or not.
NA damaging properties- If the technique comprises any solution or chemicals that degrade or damage our NA or not.
Module 1: DNA extraction
List of DNA extraction techniques:
- Phenol chloroform DNA extraction
- Proteinase K DNA extraction
- CTAB DNA extraction
- Salting out method
- Silica gel-based DNA extraction
- Alkaline extraction
- Paper DNA extraction
- Magnetic bead-based DNA extraction
- CsCl density gradient method
- Chromatography separation
- Chelex-100 extraction
DNA extraction:
DNA extraction is a process to separate DNA only, from biological material. Usually, we need genomic DNA for routine lab analysis, however, sometimes additional optimizations are required to isolate cytoplasmic DNA such as mitochondrial DNA or chloroplast DNA.
It has the same principle and experimental setup as the NA isolation. This article, I, have dedicatedly written for only this topic:
Learning objectives- what is DNA extraction, history, review on various methods, purpose, the importance of DNA extraction, optimization tips and role of chemicals.
Though I have already explained everything and every technique of DNA extraction in the previous article, I separately prepared three articles on three common and popular techniques: Phenol chloroform and isoamyl alcohol, CTAB DNA extraction and Proteinase K DNA extraction.
PCI DNA extraction:
Phenol: chloroform and isoamyl alcohol-based separation technique is, however, traditional but more effective and routinely used in a genetic lab. These three ingredients are powerful enough for DNA separation. It is suitable for the extraction of DNA from a blood sample, animal tissue or any other body sample.
Read this article to learn more about the PCI method, how to perform it, advantages and limitations.
Learning objectives- Basic understanding of PCI method, the role of each chemical in DNA extraction (viz. Phenol, chloroform and isoamyl alcohol), saturation of phenol, preparation of PCI, a protocol for PCI, advantages and limitations.
Proteinase K DNA extraction:
The Proteinase K DNA extraction method is yet an important and popular method for DNA isolation. It’s more powerful than conventional chemical-based techniques. Proteinase K digests proteins part of the cell more effectively. Thus it provides highly pure and high-yield DNA.
Learning objectives- Role of proteinase K, preparation of proteinase K solution, standard protocol, Applications, limitations and possible modifications.
CTAB DNA extraction methods:
CTAB DNA extraction is an important technique for plant DNA extraction, as plants have different cell wall setups. The difficulties in isolating plant DNA can be overcome by using the CTAB DNA extraction protocol. This article will help you to understand the present technique and how you can use it.
Learning objectives- Role of CTAB, how to extract DNA from plant tissue, preparation of CTAB solution, advantages, disadvantages and modifications.
Module 2: RNA extraction
List or RNA extraction methods:
- Phenol, chloroform and isoamyl alcohol RNA extraction
- Trizol RNA extraction
- Guanidine isothiocyanate-based RNA extraction
- Silica-gel-based RNA extraction
- Ready to use RNA extraction kits
RNA extraction:
RNA extraction is a bit different than DNA extraction, however, the basic foundation, principle and some chemicals remain the same. A special need for RNA extraction is to inactivate the RNase enzyme as it is commonly present everywhere in the atmosphere. Read this article to understand the basic concept.
Learning objectives- Basic principle of RNA extraction, problems, steps and process, function of chemicals, a basic protocol, recent advancements, importance.
Phenol: chloroform: isoamyl alcohol RNA extraction:
With slight modification, the PCI method is also employed for the RNA extraction too. However, it needs additional chemicals and an experimental setup as well but the basic principle remains the same.
I have explained the entire PCI protocol for RNA extraction, chemicals and instrumental requirements in the above-given article. You can read it here.
Trizol RNA Extraction:
Trizol is a commercially available chemical foundation, specifically prepared for RNA isolation. It commonly has guanidium isothiocyanate, phenol and a secret ingredient. Trizol or Tri reagent based RNA extraction is one of the most effective protocols. You can read more here:
Learning objectives- About the Trizol, qualities, chemical composition and protocol, Advantages and limitations of Trizol.
mRNA isolation:
mRNA is an important RNA species to study gene expression. mRNA- messenger RNA often known as a ‘transcript’ is an intermediate form of nucleic acid that translates into protein. As the RNA is a mixture of many RNA species through only mRNA, we can study gene expression.
So above all the NA isolation techniques, mRNA isolation is extremely different and requires immense expertise.
I have written a dedicated article on this. Read it here:
Learning objectives- concept, mRNA isolation techniques- spin-column, magnetic bead and chromatography, mRNA Post-processing and my suggestions.
Timely I will add more articles to this on RNA extraction and on other techniques of DNA extraction. Now let us discuss the process of cell lysis.
Module 3: Cell lysis
NA isolation needs adequate cell lysis. So the overall experimental success highly relies on how effective we lyse cells. The cell lysis buffer is an important entity for NA isolation and achieving excellent cell lysis. It comprises the composition of chemicals to do the job.
Many ready-to-use lysis buffers are available, but to make things according to our experiment we need to prepare our own composition of lysis buffers. Don’t worry, I have a secret recipe for this. Follow this article.
Learning objective- the importance of cell lysis buffer, lysis buffer for blood, plant, bacteria and plasmid DNA extraction and decoding the lysis buffer mystery.
Tissue homogenization:
Sometimes we need to isolate nucleic acid from solid tissues, for instance, plant tissue, cancer tissue, liver tissue, spleen tissue or other parafilm embedded solid tissue blocks. Any solid tissue has a different structural hierarchy and composition and thus creates problems in NA isolation.
However, still, we can certainly isolate DNA somehow by managing and optimizing the protocol but we need a totally different setup and requirements for RNA isolation. Tissue homogenization is a technique to prepare a homogenous tissue mixture. Homogenization greatly increases the yield of RNA.
Check out this article.
Learning objectives- Techniques- mortar & pestle, mechanical, vertexing, sonication and biological methods and tips and techniques.
Module 4: Nucleic acid precipitation
NA precipitation allows the physical separation of DNA or RNA, although the precipitation is more valuable for DNA only. There is a definite mechanism behind DNA precipitation which can be achieved by using alcohol. I have explained the precipitation process in this article for DNA.
Learning objectives- mechanism of DNA precipitation, importance, chemicals used and protocol.
Module 5: Nucleic acid purification
Isolation isn’t enough to get high-yield nucleic acid, purification is required as other cell contaminants are present. There are many techniques to purify the NA, however, the most effective method is by washing it using alcohol. Unfortunately, as the process of DNA and RNA purification using alcohol is similar, I have written only a single article on DNA only.
Learning objectives- Alcohol DNA purification, spin-column DNA purification and automated DNA purification and yield analysis.
Module 6: Nucleic acid elution
Elution or dissolving DNA is important as well. Usually, we use TE buffer or ultra-pure water to dissolve or elute the DNA. I have covered separate articles on this topic. You can read it here.
Learning objectives- Importance of Tris and EDTA, preparation of 10X and 1X TE buffer.
Learning objectives- what is nuclease-free water, importance and functions.
Wrapping up:
So I think this is enough for this topic. Nucleic acid extraction isn’t rocket science, but one has to understand the basic theory behind every step and how different chemicals work. If you understand all the concepts, it is also possible that you can prepare your own recipe for lysis buffer, DNA or RNA extraction.
I have also prepared an ebook which is kind of a basic lab manual to learn DNA extraction, gel electrophoresis and PCR. If you wish to learn more and get technical exposure to such techniques, please buy it and support my work.
Buy our ebook: From DNA extraction to PCR
Thank you.
I hope this guide/course kind of write-up will help you in your genetics learning.


