“In this article, Learn about the lysis buffer for DNA extraction, preparation, recipe and importance. Also, explore some of the practical optimization tips for effective lysis buffer preparation.”
DNA extraction is a crucial scheme to isolate DNA from a cell. It’s a complicated, yet sensitive technique that requires knowledge of DNA sample handling. It’s a multistep process governed by various chemical, mechanical and enzymatic procedures.
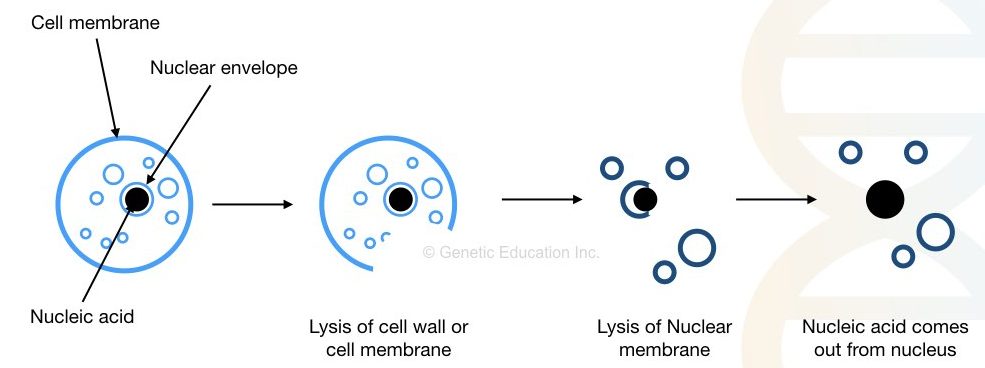
Cell wall/ membrane and nuclear membrane lysis, DNA stabilization, DNA collection and DNA rehydration are a few key steps in the DNA extraction process. The success rate of DNA extraction highly relies on the initiator step- Cell wall/ membrane lysis.
In the Cell wall/ membrane and nuclear membrane lysis, a cell’s outer layer, as well as the nuclear membrane, is disrupted and removed by the techniques mentioned.
Lysis buffer is a chemical combo that is used for this purpose- lysing the cell wall/membrane. But do you know what a lysis buffer is? What exactly is the composition of the lysis buffer? Why is it important? And how to prepare it?
In this article, I will explain the topic of a lysis buffer for DNA extraction. I will also give you standard recipes for various DNA extraction schemes. Let’s deep dive into the world of DNA extraction and understand the importance of a lysis buffer.
Stay tuned.
Key Topics:
What is a Lysis buffer?
A lysis buffer is a solution prepared using various chemicals, used in molecular and cell biology to disrupt or break the cell wall or membrane and release cellular components. It is used to isolate nucleic acid, protein and other molecules from the cell.
In DNA extraction, the cell lysis buffer is dedicatedly prepared and used to isolate the DNA only, from the cell. It’s a chemical lysis scheme that uses various chemicals to break down the cell wall/membrane, stabilize, and solubilize the DNA.
The lysis buffer is also known as cell lysis buffer, DNA extraction lysis buffer or DNA extraction buffer. So all these terms are the same, keep this in mind. The composition of a lysis buffer varies depending on the specific application and the DNA extraction scheme.
For example, the lysis buffer composition for blood DNA extraction is almost different from plant DNA extraction. However, the basic chemical foundation will remain the same.

Composition of Lysis Buffer:
Detergents, Buffering agents, salts, protease inhibitors and DNA stabilizers are common chemicals that should be present in the lysis buffer.
Buffering agents:
Buffering agents maintain a constant pH environment for enzymatic reactions. Hence, Buffering agents like Tris-HCl or phosphate buffer have been included in the lysis buffer for maintaining the constant pH range. Read about TE buffer: Importance of Tris-EDTA Buffer In DNA Extraction.
Detergents:
Detergents are a crucial component of a cell lysis buffer. A cell contains a lipid bilayer. Detergents can disrupt this layer, solubilize it and release cellular components in the solution. SDS, Triton X-100, Tween 20 and CTAB are common detergents used in the cell lysis buffer.
Salts:
Salts have many functions during cell lysis. They stabilize the DNA, preserve the integrity of the DNA, protect it from nuclease activity, maintain ionic balance and stabilize the solution.
KCl, NaCl and MgCl2 have been used in the DNA extraction lysis buffer for various purposes.
Proteinase K:
In some lysis buffers, a proteinase K enzyme is also added (separately) to make the cell lysis more effective. It is an enzyme that catalyzes the degradation of proteins from a cell and removes it from the DNA.
Note that it is added separately.
How to prepare a lysis buffer?
In this section, first I will give you a general scheme to prepare a universal lysis buffer. Later in this article, I will give you the scheme to prepare lysis buffers for different types of DNA extraction methods.
As aforementioned, an ideal DNA extraction buffer should have salts, buffering agents, detergents and DNA stabilizers. Thus, we will use Tris-HCl, EDTA, NaCl, MgCl2, KCl, SDS and proteinase K. The concentration of each chemical is given below.
- Tris-HCl: 10 to 50mM (pH 8.0)
- EDTA: 2 to 10mM
- NaCl: 100 to 500mM
- SDS: 0.5% (w/v)
- MgCl2: 10mM
- KCl: 10mM
Make up the final volume with distilled water.
Note: The concentration of ingredients may vary from scientist to scientist but mostly fall in this range.
First, prepare the stock solution of the TE buffer using the concentration given above and adjust the pH to 8.0 using a pH meter or indicator. Keep in mind that 8.0 pH is optimum and crucial for effective DNA isolation.
Weight other chemicals properly (MgCl2, KCl and NaCl) and add to the sterile flask containing the Tris-EDTA buffer and mix well.
Check the pH of the solution and make up the final volume with distilled water.
Autoclave the solution to remove impurities or nucleases. Cool it at room temperature and add 0.5% w/v SDS or other salts, as per the requirement.
Now prepare a stock solution of Proteinase K using the manufacturer’s protocol and store it at 4ºC.
Store the lysis buffer at 4ºC or at room temperature. Check the pH before using it.
Now let us prepare a lysis buffer for blood DNA, plant, bacteria and plasmid DNA extraction and their recipe.
Lysis buffer recipe for Blood DNA extraction:
DNA extraction from blood is an important procedure and is routinely used in diagnosis and research. I have a well-standardized lysis buffer recipe for blood DNA extraction, as I have a huge experience in this field.
We have divided the lysis buffer composition into two parts for effective isolation- solution 1 and solution 2.
Solution 1:
- 10mM Tris (0.061 gm)
- 10mM KCl (0.186 gm)
- 10mM MgCl2 (0.238 gm)
- Make-up final volume with D/W and set pH 7.6 to 8.0
Solution 2:
- 10mM Tris (0.061gm)
- 10mM KCl (0.037gm)
- 10mM MgCl2 (0.048gm)
- 0.5M NaCl(1.461gm)
- 2mM EDTA(0.037gm)
- Mix all ingredients in sterile D/W and set pH 7.6
- Autoclave it and wait to come to room temperature.
- Add 0.5% SDS (0.250gm)
Related article: How to Extract DNA from- Whole, Fresh, Froshen, Fried and Clotted blood.
Lysis Buffer Recipe for Plant DNA Extraction:
Plants contain a solid and pectin-rich cell wall which is difficult to lyse. So the lysis buffer for plant DNA isolation requires several additional chemicals for effective cell wall lysis.
Additional chemicals required in this scheme are CTAB, Beta-mercaptoethanol, ascorbic acid and PVP. Check out the recipe.
Solution A:
- 2% CTAB (4.0 g)
- 100 mM Tris (pH 8.0) (20 ml)
- 20 mM EDTA (2 ml)
- 1.4 M NaCl (16.4 g)
- 4% polyvinylpyrrolidone (PVP) (8.0 g)
- 0.1% ascorbic acid (0.2 g)
- 10 mM β-mercaptoethanol (140 µL)
Solution B:
- 100 mM Tris-HCl (pH 8.0) (20 mL)
- 50 mM EDTA (10 mL)
- 100 mM NaCl (0.12 g)
- 10 mM β-mercaptoethanol (140 µL)
Related article: Plant DNA Extraction Using CTAB.
Lysis Buffer Recipe For Bacterial DNA Extraction:
Bacteria contain a smooth cell wall which is easy to lyse compared to other cell types. However, the composition of the lysis buffer and protocol may vary depending on the requirements and the type of bacteria.
Most of the time, a simple composition including Tris and EDTA at pH 8.0 can do the job. Moreover, adding additional components like SDS or CTAB can give better yield and purity.
- Tris: 50mM (pH 8.0)
- EDTA: 25 mM
- SDS: 1% (w/v)
Or
- Tris: 100mM (pH 8.0)
- EDTA: 25mM
- NaCl: 1.5M
- CTAB: 2%
Either recipe can work effectively. However, learn about the cellular composition of the bacteria you are working on and choose chemicals accordingly.
Related article: How to Extract DNA From Bacteria? + Protocol.
Lysis Buffer Recipe For Plasmid DNA Extraction:
Plasmid is an extrachromosomal bacterial DNA and is present in the cytoplasm. It is commonly used in genetic engineering, gene therapy and recombinant DNA technology.
Note that a simple TE buffer can give us a good amount of plasmid DNA, however, I strongly recommend using a spin-column kit for plasmid DNA extraction for effective isolation. Or you can give it a try with this recipe.
- Tris: 50mM
- EDTA: 10mM
- RNase: 100mg/ml
Related article: A Comprehensive Guide to Plasmid DNA Isolation.
Importance of lysis buffer:
A DNA extraction buffer has been used in various DNA extraction schemes to maintain the pH, disrupt the cell, maximize the yield and improve the DNA quality. Here are a few key reasons why DNA extraction buffer is important.
DNA extraction buffer contains the ingredients that help maintain the pH of the solution, thereby providing a constant environment for various biological activities.
It contains the ingredients that effectively lyse the cell wall/ membrane and nuclear envelope by disrupting the constituents and solubilizing them using chemical treatment.
It helps remove cell wall constituents, proteins, enzymes and cellular organelles from DNA.
The cell lysis buffer also stabilizes the DNA by preventing DNA degradation. It separates DNA from other cellular debris and effectively isolates it.
A good quality and well-maintained lysis buffer improves DNA yields many folds. This purpose can be achieved by effective cell lysis, nuclear lysis and DNA stabilization.
Effective lysis also improves the quality of DNA samples. It prevents DNA by enzymatic degradation, removes DNA binding proteins and improves the quality.
In addition, the lysis buffer contains ingredients that do not interfere with downstream processing like PCR amplification, restriction digestion and DNA sequencing.
How to Optimize Lysis Buffer?
- Check the pH of your buffer periodically and ensure optimum pH (7.8 to 8.0) for DNA extraction. Do not use the buffer out of this pH range.
- The salt concentration is a crucial factor for good quality DNA. Weight each chemical correctly and use adequate concentration.
- Autoclave the solution before use. Keep in mind to add some ingredients separately only after autoclave, for example— SDS.
- Select the detergent as per the requirement of the protocol. For example— SDS for blood DNA extraction, CTAB for plant and bacterial DNA extraction and Triton X-100 or tween 20 for biological fluids.
- Read the literature to understand more regarding the use of detergents in DNA extraction and use each chemical, accordingly.
- Use additional technicals like heat or mechanical lysis along with the lysis buffer only if required. For example— heat for bacterial cells, homogenization and grinding for plant cells. Additional mechanical treatments can greatly improve the quality and quantity of the DNA.
- Use enzymes like proteinase K or RNase to improve the quality of DNA. Proteinase K and RNase remove proteins and RNA from the DNA sample. Note that, use enzymes separately.
Read more: How to Increase DNA Purity and Yield?
Wrapping up:
Lysis buffer for DNA extraction is a crucial ingredient of any DNA extraction scheme. It provides stability to DNA, disrupts the cell wall, removes impurities and improves the yield and quality of the DNA.
Although, outcomes highly depend on the expertise and experience of the performer in this field. You can also use ready-to-use DNA extraction buffers, available in the market.
I hope you like this article. To learn more about DNA extraction and related things I strongly recommend joining our e-course now. It contains video lectures, reading materials and protocols that help you in your DNA extraction endeavors.


