“RNA degradation is a common molecular lab problem caused by the fragile nature of RNA and the presence of the RNase enzyme predominantly in nature.”
Ribonucleic acid-RNA is a one of the common nucleic acid found on earth. It has crucial information in the form of a message regarding ‘in how much amount a particular gene will express’. The gene which is a piece of DNA actually has detail about what protein will form.
Whereas the RNA (the mRNA) has information about how much protein it manufactures at a tissue or cell level. Thus both nucleic acids have equally significant for a cell. So, at a technical level, by extraction of DNA and RNA one can study genes and gene expression, respectively. Note that DNA and RNA differ in many aspects. Read this article to know more:
DNA vs RNA: Differences And Similarities.
RNA extraction, however, is not as straightforward and easy as DNA extraction. There are a couple of valid reasons for that and occurs frequently during the extraction process. The assessment strength of any RNA assay solely relies on the quality, quantity and integrity of RNA.
With state-of-art instrumental development in nucleic acid extraction technology, the quality and quantity of RNA and even DNA can be governed, but integrity, especially for RNA is still a big hurdle in the same process. The integrity of RNA shows how much RNA is intact, usable and non-degraded.
Unfortunately, RNA degradation to “more or less extent” occurs during the extraction, assessment, estimation and testing. So what is RNA degradation and why is it a matter of concern? you may wonder, right!
In this article, I will explain the process of RNA degradation, how it occurs, causative factors and assays for detection. And at last, I will explain, technically, 5 parameters that show RNA degradation. The last segment will certainly boost your RNA extraction knowledge.
Stay tuned.
Key Topics:
What is RNA degradation?
RNA is acutely susceptible to degradation. As a geneticist, we know that RNA decay is a headache for us, but we have to understand why it is so? Why not DNA and why RNA is so vulnerable to degradation. And to better understand the fact, we need to jump into a cell and central dogma process, first.
mRNA is an intermediate transcript a cell forms, in fact in any cell, most often. This mRNA eventually forms the chain of amino acids. From tissue to tissue the protein requirements vary, so the degradation process destroys the mRNA, which is not required or less required and controls the gene expression process.
So first, degradation at a cellular level is significant for a controlled epigenetic profile. Second, it also protects a cell from microbial infection by degrading microbial RNA.
Any pathogen inserts its nucleic acid in the form of mRNA to the host cell and completes its life cycle there, but a cell’s natural defense system finds foreign mRNA or any RNA and degrades it.
Henceforth, to protect us, our cells and skins; nuclease in the form of RNase are everywhere, constantly looking for foreign RNA to degrade it. This seems beneficial to us but not for the extraction process.
Definition:
RNA degradation is a process in which, under the influence of external factors, RNase enzyme action or by any means the RNA digests, destroys or fragments. One common factor is the presence of the nuclease enzyme.
Now take a look at the common factors in this scenario.
Factors that cause RNA degradation:
Nuclease enzymes, temperature, chemicals and other external stimuli are common factors for RNA degradation. Take a look at each factor one by one.
Enzyme:
Nuclease as RNase is a natural RNA ‘killer’. RNase is present everywhere and we have discussed why! It is present on our hands, face, working bench, surface, atmosphere, and everywhere. So once our RNA gets in contact with RNase by any means, it deteriorates.
The active catalytic domain of the RNase, having a higher binding affinity to the RNA interacts with the RNA minor groove. It finds the phosphodiester bonds and breaks them. Read more about the RNase here: What is the function of RNase?
Heat:
Heat can fragment RNA rapidly. The temperature at or above 70°C, makes RNA more susceptible to degradation.
Other factors:
Acids, oxidation, UV light, other radiation and hydrolysis quickly destroy RNA.
How to avoid RNA degradation?
This topic I have discussed in my previous articles but as it is an important link in this article, I am given a brief overview.
- One of the easiest ways to prevent RNA degradation is to inactivate the RNase enzyme.
- Use certified RNase-free utilities.
- Maintain an extreme RNA-free environment.
- Use DEPC- water during the extraction.
- Autoclave utilities before extraction.
- Use RNA stabilizers.
- Cryopreserve RNA in liquid nitrogen.
Related article: 11 Effective Ways to Use DEPC-Treated Water in RNA Isolation.
Assays to detect RNA degradation:
Gel electrophoresis of RNA is a classic way to investigate the RNA degradation or integrity but not RNA quantity, keep in mind. However, the Capillary electrophoresis-based RIN- RNA integrity number score is the most robust, accurate and rapid assay.
Also, by looking into the Real-time PCR results, RNA degradation can be estimated. We will discuss all the assays one by one.
Gel electrophoresis of RNA:
Electrophoresis for RNA is a classic, traditional and most trusted technique that not only shows RNA degradation but also the quality of RNA. Samples are run on an 0.8% to 1% gel under the influence of a constant electrical current.
The reagent preparation and RNA electrophoresis process remain somehow similar to the DNA gel electrophoresis. The banding pattern, quality of bands, smears and intensity are enough for the qualitative assessment of RNA. For RNA degradation, however, 18S rRNA and 28S rRNA banding patterns are closely examined.
Electrophoresis is an advantageous, handy and more practically suitable method. Although, it lacks a quantitative evaluation of RNA degradation. Meaning, we don’t know how much RNA is intact or degraded. Furthermore, the present technique is tedious, labor-expensive and time-consuming.
The use of hazardous chemicals also restricts its use.
RIN- RNA integrity number:
I wrote a whole article on this topic. I am giving a brief overview here. RIN is a capillary electrophoresis-based quantitative evaluation assay to estimate RNA integrity. The sample run in capillary provides a graph with peaks for every RNA species present in the sample.
Along with the 18S rRNA and 28S rRNA, other RNA species and factors are also taken into account by the software to depict the RIN value. RIN score >8 is excellent, and <5 is poor.
You can read more here: What is RIN? And why it is important.
RIN by CE analysis is a rapid, accurate and robust analysis assay. All the laboratories across the world use the RIN score for RNA-based assays. Bioanalyzer is an instrument used for RIN estimation.
RT-PCR analysis:
Real-time PCR-based analysis, though, is highly not advisable but can be used. The above two assays are primary and help in making decisions on whether to use the RNA for downstream processing or not. But RT-PCR assay for degradation study is different.
RT-PCR-based quantitative evaluation of mRNA is important for transcriptomic studies and RNA sequencing. The real-time amplification curve analysis demonstrates the value of RNA degradation and whether or not the same mRNA or RNA sample will be used further or not.
Unfortunately, it is a costly and time-consuming process.
5 interpretation Manifests RNA degradation:
- RIN < 6
- The size of mRNA < 1.5Kb.
- 28S:18S rRNA ratio more or less than 2:1
- Abnormal electropherogram areas other than 28S and 18S peaks
- Smear in a gel
Now, this segment is something important, more valuable, technically and helps actually while handling the RNA. Without quantitative analysis, we can’t determine DNA degradation or integrity, even the gel electrophoresis, which is a popular method for nucleic acid examination, needs expert eyesight to know about RNA degradation.
Still, it isn’t guaranteed!
However, here I’m enlisting several technical interpretations in which you can say that the RNA is degraded.
RIN < 6:
RIN, as aforementioned, is a quantitative tool for RNA integrity and degradation analysis. The standard RIN graph looks like this.
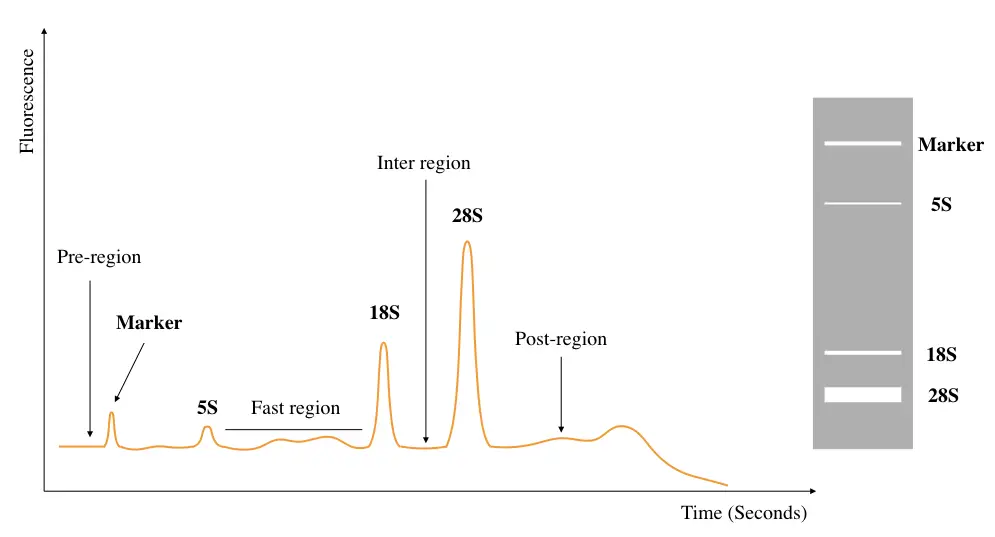
Now take a look at some of the real RIN electropherograms.
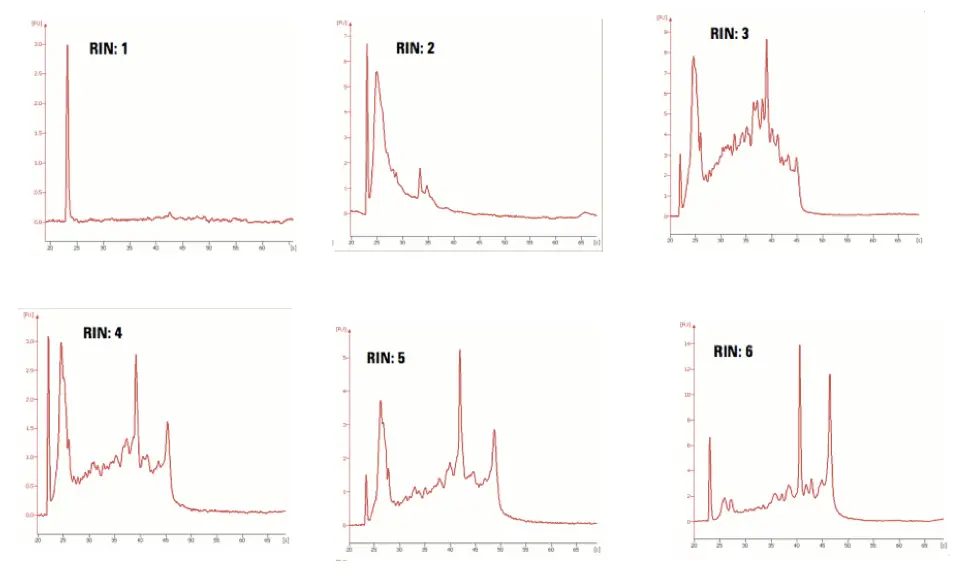
RIN value 1-6 shows substantial degradation and is not advisable for downstream processes. Keep your eye on the 18S and 28S peaks, which should be shorter and longer, respectively. Other parameters like 5S rRNA, marker region and other graph areas are taken into consideration too.
28S:18S rRNA ratio more or less than 2:1:
It should be noted that when we look into a gel or electropherogram of RNA, the ratio of two important RNA species 28S and 18S rRNA should be 2:1. Put simply, the peak of the 28S rRNA or the band must be doubled in height or more intense than the 18S rRNA, respectively.
A change in the ratio higher or lower signifies a decent amount of RNA degradation.
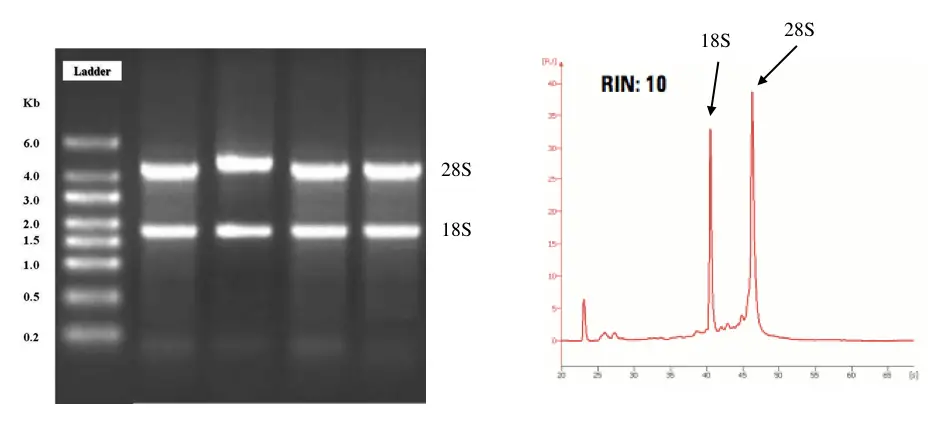
Abnormal electropherogram areas other than 28S and 18S peaks:
Newbies always rely on the computational results and only 18S and 28S rRNA peaks. But that’s not enough. Other electropherogram areas should be observed to assist the overall integrity of RNA.
Every other area- the peak of 5S, marker region, pre, post, inter and fast region should be correctly situated in the graph. Change in any of the properties shows a substantial amount of RNA degradation. For example, too large marker region, higher 5S rRNA peak, too large fast region, etc.
Experts suggest that all the electropherogram areas/properties must be observed for RIN evaluation viz RNA integrity assessment.
Read more: What is Electropherogram? How to Read it?
Smears in a gel:
Gel electrophoresis has been a conventional way to determine the quality and integrity of the RNA, in fact, DNA as well. Although it lacks quantitative interpretation, still expert eyesight and experienced personnel in electrophoresis can interpret the results.
Ideally, RNA is observed as intact, intense and thick bands, and as it has many RNA species, different banding patterns are observed. So if such banding characteristics aren’t seen, it eventually depicts RNA degradation.
The common observation showing a decent amount of RNA degradation is “smear”. When RNA degrades, it forms a long tail-like smear in the gel. Take a look at the image.
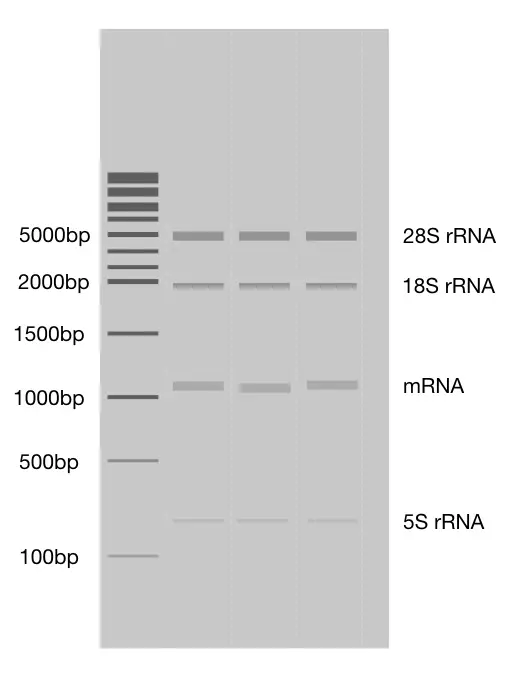

If the smear is too small and still bands are visible, purification can help to remove the fragmented RNA and one can collect pure and intact RNA once again. However, re-purification also increases the chances of further degradation.
The average size of mRNA <1.5 Kb in a gel:
28S and 18S rRNA are two important markers for RNA degradation studies. However, other RNA species’ sizes should be considered for this purpose. The mRNA size should be less than ~1.5Kb. Notedly, it lacks a clear band and thus looks like a slight smear in the gel.
Keep note that the mRNA thick smear must be <1.5kb, ideally 1.1Kb. Size difference shows RNA loss.
Wrapping up:
Intact RNA is very important for gene expression and transcriptomic studies. Degradation fakes the results or sometimes fails the assay. Nonetheless, techniques like conventional gel electrophoresis or RIN by CE are very useful for RNA analysis.
RNA stabilizers should be added before storing the extracted RNA and stored under appropriate conditions. Take necessary precautions, training and experience before processing RNA.