Explore the concept of data storage in DNA. Learn about the process, steps, applications and limitations of storing digital data in the DNA.
Ancient humans used to craft their art and daily events on stones and rocks. We are decoding that information now to understand their culture, living and lifestyles. Such information is a crucial part of human history.
Unfortunately, it has been continuously vanishing due to rain, water, wind and other factors. Resultantly, that part of our history is missed now! We and our coming generations would never know about it.
We evolved and found new ways to store data. The first hard drive was developed by IBM in 1956. After that, floppy disks, CDs, magnetic devices, hard drives and many other storage devices were developed.
MicroSD card is the latest and most compact physical device we have right now, for digital data storage.
Digital data is increasing every year. In 2012, the world’s total digital data was 2.7 ZB which will be 175 ZB by 2025 as per a recent study. Thus, we need more durable and high storage capacity units in the future.
Data is everything at this time. Physical data storing options have limited space and can be lost and broken. Although we have cloud storage, on the back end it is also a series of hard drives.
So the data storage options that we have, aren’t durable and long-lasting, and further, come with storage limitations. In addition, it’s costly and energy-consuming. We need more powerful options to store, transfer, retrieve and read the data.
See the picture below, it tells a thousand words, how the data storage units have evolved.
From a tons-heavy storage with a few MB size, we have a tiny mm-sized SD card with a few hundred GB storage. But what next? We are still running out of space, and data may still be lost.
Is there any secure and effective way to store data for a longer period of time? Can we store our historical data for thousands of years? Is there anything smaller than the micro-SD card?
The answer is DNA (Deoxyribose Nucleic acid)! Our own DNA can be used as a data storage unit. It can store a huge amount of data for hundreds to thousands of years without any degradation. Let us understand the concept of how digital data can be stored in the DNA, and the positive and negative sides of this technology.
Stay tuned.
“A gram of DNA can store 1 exabyte of data for as long as 2,000 years”.
Key Topics:
Introduction:
Nature equipped us with infinite data storage capacities. For millions of years, organisms have been storing their information in nucleic acid– particularly in the DNA. That information also transmits to the progenies and helps them to survive on Earth as well.
So DNA is the oldest data storage unit. It’s so huge that if we print all three billion nucleotides on paper, and stack all the pages, it will be larger than the Statue of Liberty. The human body contains approximately 30 trillion cells! Imagine the amount of DNA we have!
Let’s understand it by an example.
A recent study published in Nature collected and identified 38,000-year-old Neanderthal DNA. The study further estimated that modern humans and Neanderthals diverged from approximately 5,00,000 years ago.
Such lucrative information could be retrieved only by looking into some DNA sequences, which are not even visible to us. DNA store and retrieve so much information that we can’t even imagine.
DNA is a natural information storage unit and now it’s proven that it can be used to store digital data as well. And why not? After all, it’s a long sequence of As, Ts, Gs and Cs.
So instead of storing our data in the binary language of O and 1 which we are doing right now, we can store it into A, T, G and C bases of the DNA. Now, let’s understand the process of how to store digital data in the DNA.
Comparison of different data storing units :
|
Storage unit |
Storage capacity |
Storage density per mm3 |
|
Magnetic tap |
185 TB |
10GB/mm3 |
|
Optical disc |
1PB |
100GB/mm3 |
|
DNA sequence |
>1XB |
1EB/mm3 |
How to Store Digital Data Into DNA?
Encoding data:
DNA is a long polynucleotide chain. Each base in a nucleotide is A, G, T, and C. Now, we can encode our digital information in this format.
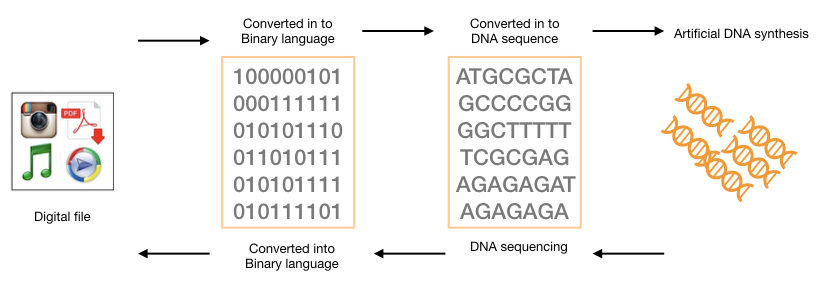
The computer works on the binary language of 0 and 1. So technically, all the information that we have, whether it’s our selfie, Instagram video or just a text file; on the backside is just a sequence of 0 and 1 numbers. Now, to encode the information into the DNA, the binary language 0 and 1 is translated into the DNA sequence.
For example, 00 – A, 01 – C, 10 – G and 11 – T. So if we have a file in a binary language of 00 11 10 10 10 00 10 01 11 10, it is translated into the DNA sequence as A T G G A G C T G.
This process is encoding/translating the binary data into the DNA. What we have after the end of this step is just a sequence of nucleotides.
Synthesizing the DNA strand:
Now, using the sequence information, a new DNA strand is synthesized in the genetic lab. The synthesizer reads every bit of information and creates a new DNA strand that is unique in the sense that it contains some digital information.
(the information that we have added to it.)
Storage and transport:
Now we have our digital data in the form of a DNA droplet. It’s collected in a sterile tube and stored at 4ºC temperature. For long-term DNA storage, the sample is processed and stored in liquid nitrogen at -198ºC.
This sample can remain stable for hundreds to thousands of years. As the sample is in the tube, it can easily be transported from one to another place without deteriorating the information.
Note that care must be taken while DNA sample handling, processing and transporting the DNA.
Information retrieval:
Now, to retrieve the information, the sequence can be decoded. A state-of-art sequencer machine reads the sample sequence and creates a readable file of the sequence.
The sequence file is again sent for data decoding.
Decoding the information:
Afterward, the genetic code is decrypted back into the binary code and into the digital file using computational software. The software uses the mapping information that we have given in the first step to decrypt the information.
This is a simple-looking complex process of how digital information can be stored in the DNA and retrieved back into a digital format.
Advantages:
Here are a few amazing advantages of data storage in DNA.
Stability:
DNA can remain stable for thousands of years if adequate conditions are provided. The oldest DNA sample we have is 7,00,000 years old horse DNA and still, we can retrieve information from it.
Exceptional data storage density:
DNA is a very tiny molecule in which the information is stored in four nucleotides. It can store >1XB of data in 1EB/mm3.
High data storage capacity:
DNA can store huge amounts of information which is right now technically impossible to determine. We can store more than Xeta bytes of data in a single gram of DNA.
Data protection:
The three-dimensional structure of the DNA provides not only stability but also protection to the nucleotide strands. So DNA can’t be easily broken. Henceforth, the data remains secure under normal physiological conditions.
Copied easily:
Using techniques like natural replication or polymerase chain reaction, DNA can be copied easily, meaning, the set of information can be copied millions of times. Therefore, huge amounts of digital information can be stored in various copies.
Maintenance and transportation:
DNA can be easily transported and maintained, and therefore any data that is stored in it can be transported and maintained easily.
No additional need:
DNA storage is a durable option and doesn’t require additional energy or electricity.
|
Durability |
Data storage unit |
Access time |
|
3 years |
Flash drive |
Mili second |
|
5 years |
HDD (hard disk) |
10 second |
|
Up to 30 years |
Magnetic tape |
1 minute |
|
More than 100 years |
DNA storage |
More than 12 hours |
Applications:
Digital Heritage Preservation:
Right now! The best application for storing digital data in DNA is for long-term digital data storage, for instance– digital heritage preservation. DNA can be stored for a very long time.
Historical information, cultural artifacts, digital arts, historical data and other crucial information related to evolution, history or heritage can be stored in the DNA. Thus, future generations have access to such rich human history.
Miscellaneous files:
Miscellaneous files are difficult to store as it is in huge amount but mostly useless. Storing such miscellaneous files like medical records, legal documents, formal records and government documents in a DNA will be advantageous as we can store a huge amount of data in just a µL of DNA.
Creating archives:
DNA data storage is an amazing option for the government or organizations that are maintaining huge archives. They can store all their data in the DNA and save it for many years.
Scientific data storage:
Scientific data is so huge and organizations or companies have to face problems regarding data storage. For example, the data from space exploration or genomic research is enormous and valuable as well.
Storing these types of data into the DNA will help the organization or company in terms of data storing capacity, security and cost.
Did you know that the world’s entire digital data can be stored in a DNA archive in a single room?
Limitations:
Despite having lucrative advantages and potential good applications, the present data storage technology has several limitations too.
- Firstly, it’s a costly process. Information input, DNA synthesis, DNA sequencing and information retrieval are very costly processes. As per one report, the cost of 50MB of data storage in DNA is 1,00,000 dollars.
- In addition, the entire process is time-consuming. It takes a week to complete this process from coding information to retrieving it.
- Realistically, we can’t use the present technology as we use a pen drive or a CD. It can only be used for larger and giant setups only.
- Technical limitations are also a major problem for us now.
- Sequencing introduces errors in the sequencing read. So every time we sequence the information-encoded DNA, some sequences may be lost and the message or information can not be accurately decoded.
- In addition, on a technical level, synthesizing as well as reading the long DNA sequences are difficult to process and prone to error. As the length increases, the chances of error also increase.
New research:
- Scientists from Microsoft in collaboration with the University of Washington developed the first fully automated system to store and retrieve information from DNA. In 2015, 250MB of data from different files were retrieved from the DNA.
- In 2012, Church G. et al., from Harvard University successfully stored a book, an audio clip and a Javascript program into the DNA for the very first time.
- Shakespeare’s sonnet was stored in DNA in 2013 by researchers at the European Bioinformatics Institute.
- In 2019, Microsoft purchased 100 million DNA strands to continue their research on DNA digital data storage.
Wrapping up:
I guess, after reading this article, your dopamine level lifts up– How fascinating this thing is! Although, it’s not as easy as it looks. A huge computational power and software input is needed to encode and decode the information.
We have hope for so many aspects of the digital data storage in the DNA, the challenges right now that we have are so difficult to overcome, for instance— the accuracy of sequencing, data retrieving time and overall process.
We can’t fully explore the present technology but I am pretty sure that we will have an excess in the future. Imagine your entire health data will be in a small tube, safe and secure.
Isn’t that amazing?
I hope you like this article. Share it and bookmark the page.
Sources:
Ahn T, Ban H, Park H. Storing Digital Information in the Long Read DNA. Genomics Inform. 2018 Dec;16(4):e30. doi: 10.5808/GI.2018.16.4.e30. Epub 2018 Dec 28. PMID: 30602091; PMCID: PMC6440670.
Buko T, Tuczko N, Ishikawa T. DNA Data Storage. BioTech. 2023; 12(2):44. https://doi.org/10.3390/biotech12020044.


