Using the 3’ OH end of the primer, DNA polymerase adds nucleotides to the growing DNA strand for facilitating replication.
Different types of DNA polymerases help in different steps of replication.
Inheriting the information from one generation to another generation is a primer goal to any organism. By doing this, the organism can inherit characters or genotypes from one to another generation and hence do survive.
To fulfill this aim, cell division is must needed to create new tissues. And thus the DNA must be copied.
DNA replication facilitates DNA copying using DNA polymerase.
Using the RNA primer, the DNA polymerase synthesizes a new DNA strand from the existing one.
We have covered articles on DNA primase, helicase, single-stranded binding protein, ligase and topoisomerase. now it is time to move to the next step of replication, which is DNA synthesis.
We are here now,
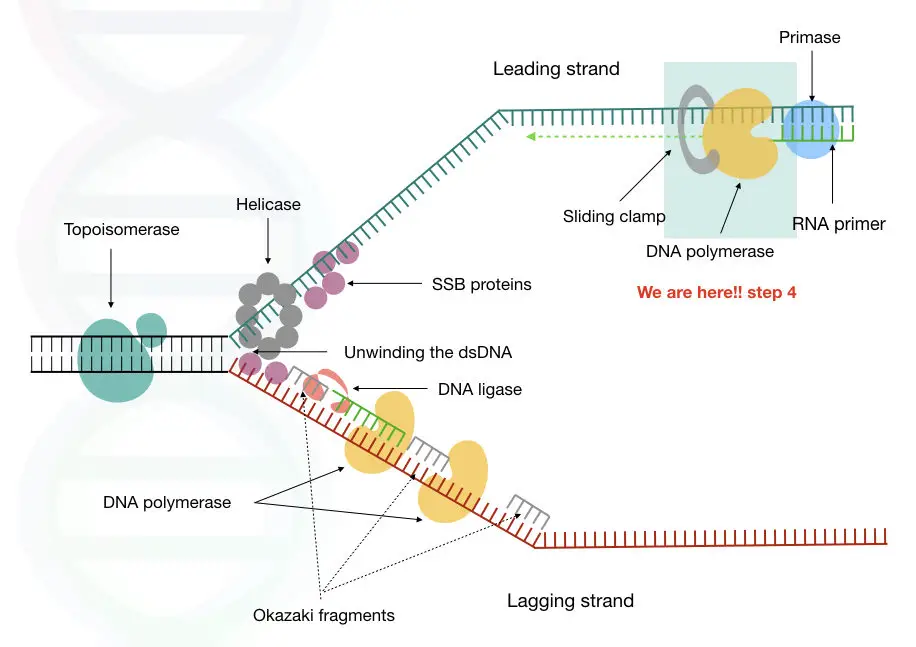
In 1958, Arthur Kornberg and colleagues discovered the first DNA polymerase from the E.coli and named it DNA pol I.
Interestingly, DNA polymerase II was discovered by Thomas Kornberg, son of A. Kornberg, who discovered type II polymerase working with Malcolm E. Gefter.
In the present article, we will discuss the main player of DNA replication, the polymerase. the content of the article is:
- The structure of DNA polymerase
- Types of DNA polymerase
- Eukaryotic DNA polymerase
- Prokaryotic DNA polymerase
- The function of DNA polymerase
- Conclusion
Key Topics:
Structure of DNA polymerase:
DNA replication is one of the important functions in all organisms starting from prokaryotic bacteria to the eukaryotes.
Hence, for a cell, it is very crucial to maintain the proper function of DNA polymerase in all the cells and in all the organisms.
The function of DNA polymerase is irreplaceable thus the structural domains of DNA polymerase are highly conserved in all organisms.
However, independent of its domain structure, the subunits may vary from species to species.
Hypothetically, the structure of the DNA polymerase is somehow similar to our hand. The structure contains three parts as like our right hand, the thumb, the palm and the fingers.
See the image below,
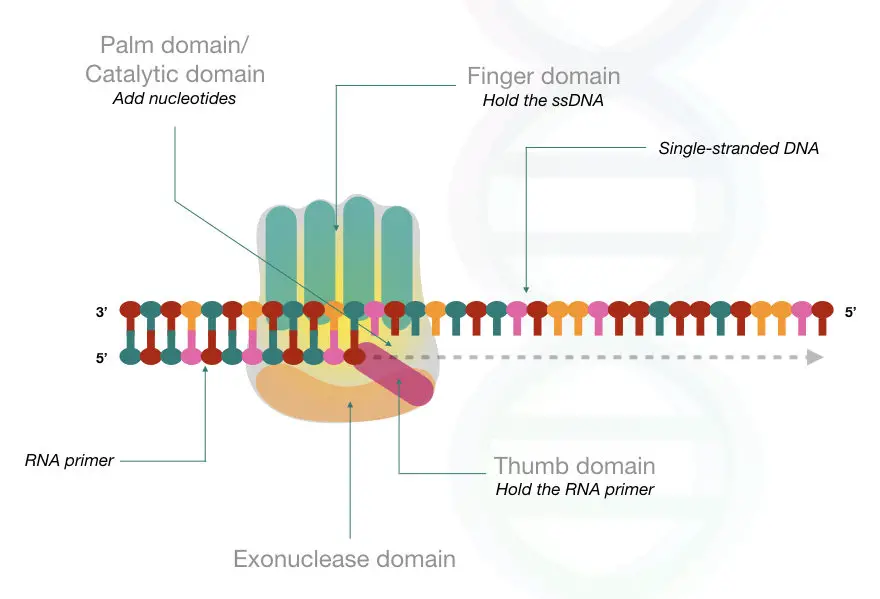
The thumb-like structure helps in the translocation of the enzyme once the nucleotide is added thus do the processivity.
The finger domain binds to the nucleoside triphosphate of the template single-stranded DNA.
The finger domains hold the DNA strand properly to synthesize the new strand.
The palm domain plays an important function by providing the catalytic domain for the enzyme to work.
By doing the phosphoryl reaction removes two phosphate groups from the nucleotide and creates phosphodiester bonds as well as the hydrogen bond.
See the image
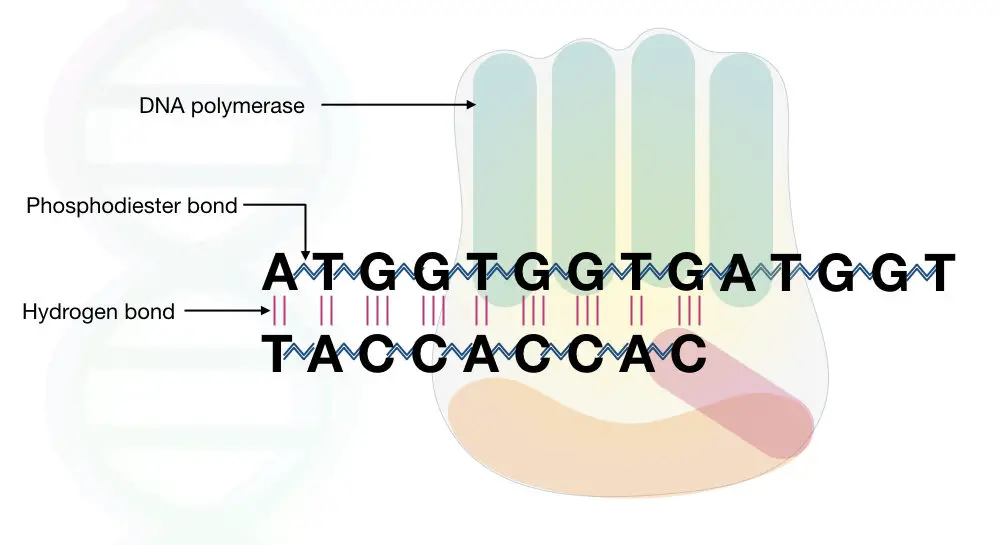
As we know that some special type of enzyme requires a metal ion to catalyze the reaction, the DNA polymerase uses two metal ions for doing DNA synthesis.
Some of the other proteins also play an important role in maintaining the activity of the DNA polymerase, named sliding clamp and sliding clamp loaders.
The sliding clamp is a ring-shaped structure that helps DNA polymerase to settle on a single strand. And thus prevents the dissociation of DNA polymerase.
The clamp loading is facilitated by the DNA clamp loader protein.
“Due to the proofreading mechanism armed in DNA polymerase, the error rate of DNA polymerase is very low, one in every billion nucleotide synthesized can be mistaken.”
Different types of DNA polymerases:
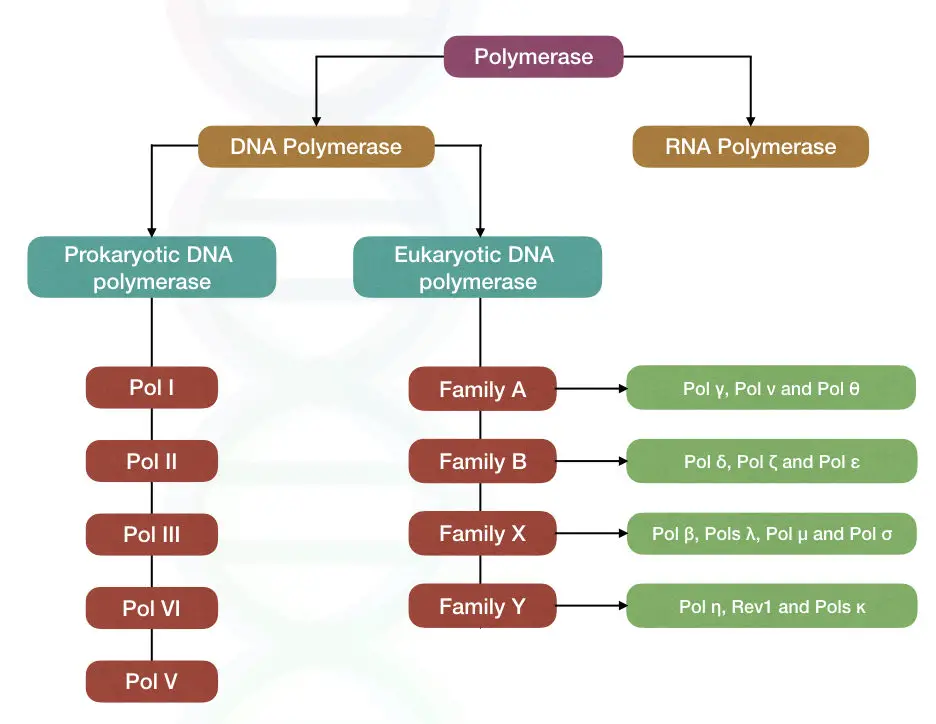
The DNA polymerases in prokaryotes as well eukaryotes are different and performs a different function. The common type of polymerases are enlisted below,
Eukaryotic DNA polymerase
Family A:
The polymerase categorized under this family is mainly involved in the Okazaki fragment synthesis and DNA repair.
Besides polymerization, the members of this family also have the 3’ to 5’ and 5’ to 3’ exonuclease activity. However, the exonuclease activity lacks in some.
The polymerases of Family A have a little role in DNA replication actually. But it plays some other functions too, such as antigen variability generation, DNA repair, TLS (translesion DNA synthesis) and somatic hypermutation.
Pol γ (gamma), Pol ν (nu) and Pol θ (theta) are examples of the family A DNA polymerases.
Interestingly, family-A DNA polymerases are similar to that of the DNA Pol I of E.coli. However, the major role of E.coli DNA Pol I is in DNA replication, it is not involved in other activities.
Family B:
The leading strand synthesis is one of the major functions to perform in DNA replication. The enzymes in the present family perform the function of DNA synthesis in replication and are thus called replicative enzymes.
Pol δ (delta) and Pol ε (epsilon) are members of Family B, adds billions of nucleotides to the growing DNA strand and hence they are the true polymerase functionally.
Although the Pol α (alpha) is another member of this family, it works by complexing with the primase.
Interestingly, family B polymerases have only 3’ to 5’ exonuclease activity.
The function of Pol ζ (xi) is not understood well.
Family X:
Pol β (beta), Pols λ (lambda), Pol μ (mu) and Pol σ are categorized in the Family X of the DNA polymerase. The present class of polymerases is different from the family B DNA polymerases.
Unlike the family B DNA polymerases which are involved in DNA replication, majorly; the Family X polymerases are involved in DNA repair and gap filling. And hence it is also named gap-filling DNA polymerase.
Structurally, the family X is monomeric in nature having an 8KDa DNA binding domain present on the NTD (N-terminal domain).
This extra domain unlike other groups of polymerases is mainly helping in the recognition of a gapped substrate for polymerization. And thus it has DNA binding activity as well.
Interestingly, the Pol β, Pols λ, Pol μ and Pol σ are also present in some prokaryotes as well.
Broadly the function of other polymerases of this family are:
- The Pol β is involved in the base excision repair.
- The Pol σ is involved in the sister chromatid cohesion repair.
- Pol λ and Pol μ are involved in the repair of double-strand end joining (non-homologous end joining).
Besides this, the members of this family are also involved in the V(D)J recombination process and antigen diversity.
Family Y:
Pol η (eta), Rev1 and Pol κ (kappa) are the members of this unusual family of the DNA polymerase.
Why this family of polymerase is unusual?
Because it has lower or poor polymerase activity and does not have the exonuclease activity too, unlike the family A, B and X.
Furthermore, the polymerases classified in this family are neither involved in the replication nor in the DNA repair pathways.
The present group of polymerases works in the process called translesion synthesis.
During DNA replication some of the DNA lesions interfere in the process of DNA synthesis and thus slowing down the process.
To maintain the constant polymerization, these DNA lesions are bypassed by the cell’s natural replication mechanism which is later synthesized by the Y family polymerases.
The DNA polymerase works in a pair for synthesizing the DNA.
Prokaryotic DNA polymerases
Pol I:
The DNA Pol I is encoded by the PolA gene having both the polymerase and exonuclease activity.
The prokaryotic Pol I has both 5’ to 3’ and 3’ to 5’ exonuclease activity and a majority of the DNA synthesis function (95%) are done by the DNA Pol I alone. Though, it is replaceable.
The processivity rate of DNA Pol I is very poor, as it can add 15 to 20 add nucleotides per second.
As it is the main player of prokaryotic replication it synthesizes DNA at three places.
First, during the leading strand synthesis, here it starts adding nucleotides once the DNA: RNA hybrid once the OriC is formed.
Second, at the lagging strand, during the synthesis of Okazaki fragments.
Third, during the base excision repair. Here it performs polymerization as well as exonuclease activity to repair the mismatch.
Further…
Our best friend in PCR, the Taq DNA polymerase is a member of this family hence it belongs to the prokaryotic DNA Pol I family.
But……
It does not have exonuclease activity.
Read more on the Taq DNA polymerase: Function of Taq DNA polymerase in PCR
DNA Pol III:
The DNA pol III is categorized in family C polymerases having three different subunits called α (alpha), δ (delta) and θ (theta) subunit.
The alpha subunit has dedicated polymerase activity while the delta subunit has the exonuclease proofreading activity.
Interestingly, the theta subunit helps in stabilizing the complex during the catalytic activity.
DNA Pol III is a kind of holoenzyme. It is encoded by the polC gene.
DNA Pol II:
Encoded by the polB gene, the DNA Pol II has polymerase as well as the 3’ to 5’ exonuclease activity which is mainly involved in the DNA repair pathway.
Another function of Pol II is to support the activity of the DNA Pol III. Once the activity of Pol III is blocked ( in some cases), the DNA Pol II works in place of it.
Further…
Another friend of ours, the high fidelity DNA polymerase used in the PCR, Pfu polymerase is a member of the DNA pol II family hence family B member.
DNA Pol IV:
Pol IV is a family Y polymerase encoded by the dinB gene, an error-prone DNA polymerase involved in non-targeted mutagenesis.
If the dinB gene lacks, it increases the risk of mutations that can not be repaired further due to the lack of pol IV.
The mechanism of action of Pol IV is complex, but remember one thing, it is expressed during the SOS responses of bacteria.
DNA Pol V:
The prokaryotic Pol V is also a Y family DNA polymerase which is functions during the SOS response as well.
Further to this, it is also involved in the translesion repair pathway too.
Apart from all the DNA polymerases, two other types of polymerase are also present in nature which are very essentially required for the survival of an organism.
Reverse transcriptase:
The reverse transcriptase enzyme is an RNA-dependent DNA polymerase that has the ability to synthesize the DNA strand from the RNA.
Although it is not found in the eukaryotes, it is present only in retroviruses.
The retroviruses having an RNA as their genetic material required a special mechanism that synthesizes DNA from the RNA. The reverse transcriptase facilitates this function.
Telomerase:
Actually, the telomerase is not a true DNA polymerase, but it synthesizes the DNA sequences at the end of chromosomes.
The end of the chromosomes known as the telomerase has the 5’-TTAGGG-3’ repeated sequences at its ends.
These single-stranded overhang sequences can not be replicated by the normal DNA polymerase of a cell during the replication.
Ribonucleoprotein telomerase does this work on behalf of DNA polymerase.
However, it does not require the free 3’ OH group for starting the nucleotide addition, instead, the TERT subunit of it, works as a reverse transcriptase that uses RNA to synthesize the DNA and thus provides the DNA: RNA junction for telomerase to work.
After each cycle of replication, the telomere becomes shorten which affects the aging of an organism.
The function of DNA polymerase:
Obviously, the main function of the DNA polymerase is to synthesize the new DNA strand, notwithstanding, de novo synthesis can not be possible for DNA polymerase.
De novo DNA synthesis is a process in which the entirely new DNA is synthesized without using a primer.
The RNA polymerase only has the power to do so, no DNA polymerase is able to synthesize the DNA de novo.
Well then, how DNA polymerase does its action?
Any of the DNA polymerases required two things for starting the catalytic reaction, one is the single-stranded DNA and the second, the free 3’ OH end.
The enzyme called helicase fulfills the first requirement by unwinding the DNA along with the SSBP protein.
Read more on helicase and SSBP:
The second requirement, the free 3’ OH end is provided by the short DNA or RNA primer.
The enzyme called DNA primase synthesizes a short stretch of 10 to 15 nucleotide long primer which provides the free end for DNA polymerase to work.
Read more on primer: Meet DNA Primase: The Initiator Of DNA Replication
Now in the next step, the DNA polymerase settled on the single-stranded DNA/ primer junction and starts the catalytic reaction.
Its joints the free 3’OH end of the primer with the 5’ P end to another nucleotide, once the phosphodiester bond formed, it moves forward.
The direction of polymerase activity is thus from 5’ to 3′.
See the image below,
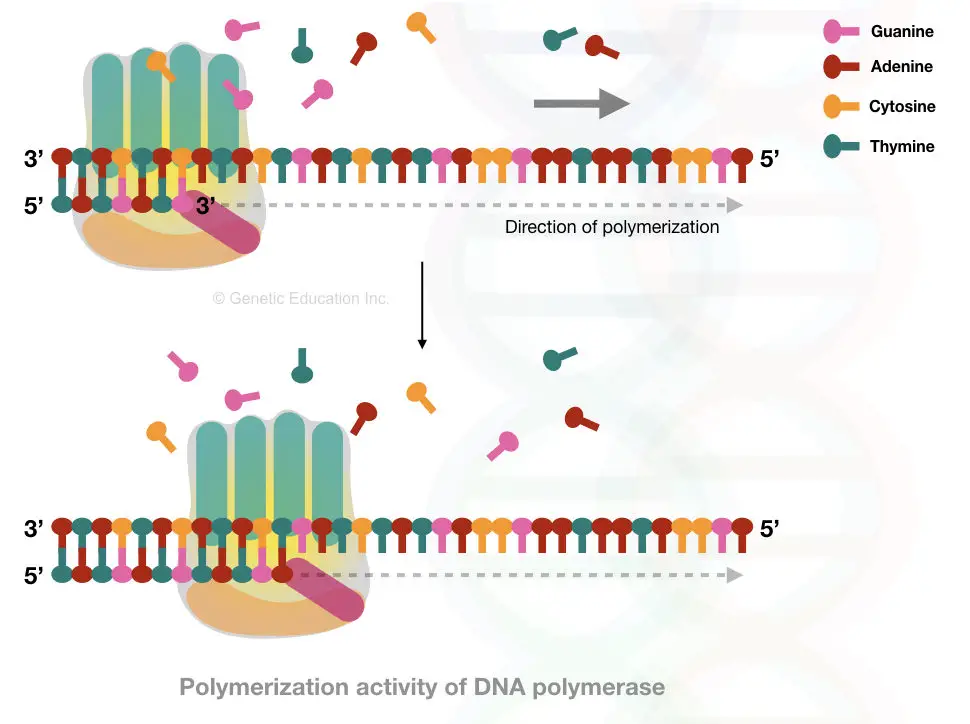
Note
The DNA polymerase can add nucleotides only on the preexisting nucleotide strand and only from 3′ OH end.
As we discussed above, DNA polymerase also has a proofreading activity.
The proofreading activity lead by the exonuclease domain of DNA polymerase identifies the mismatch and moves back.
It moves back immediately, identifies the wrong nucleotide and removes it by the exonuclease activity, after that it adds the correct nucleotide and once again moves forward in its path.
The work is done by the specialized domain called the proofreading or exonuclease domains, The proofreading domain is present at the bottom of the enzyme.
See the image,
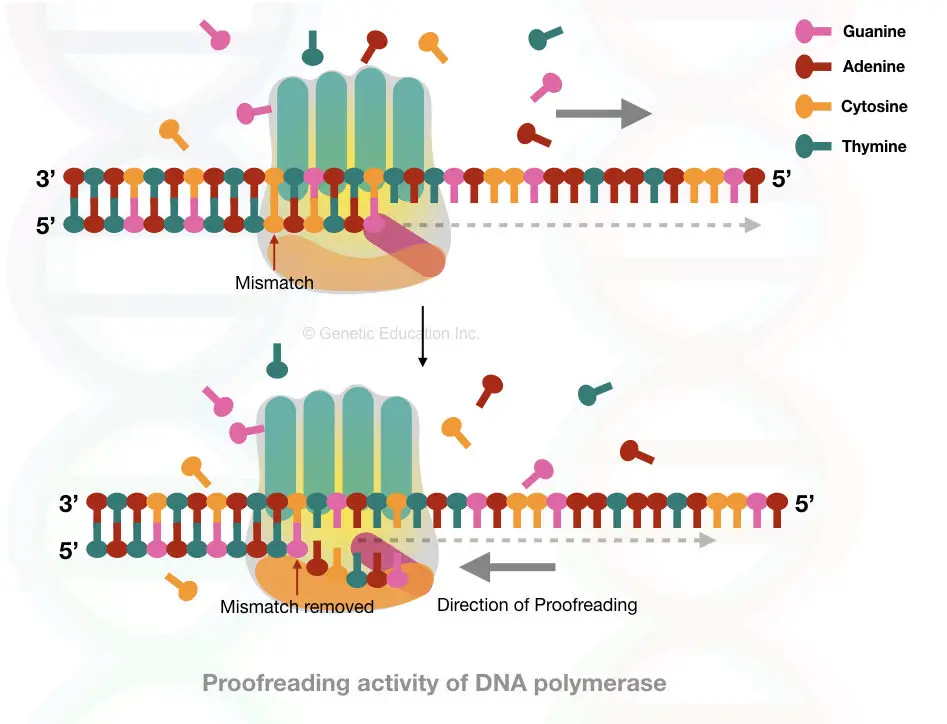
Besides its role in the DNA replication, the DNA polymerase is also involved in some other functions of DNA maintenance and DNA metabolism.
The DNA polymerase also functions in:
- DNA repair pathway
- Translesion activity
- Maintain the length of telomere
- Gap filling
- Somatic hypermutation
- Antigen diversity
- V(D)J segment recombination
The highest activity of DNA polymerase reported in the phage T4 at the rate of 749 nucleotides per second at 37°C temperature.
However, the normal rate of the nucleotide insertion is one nucleotide per second for a non-processive DNA polymerase.
The processivity of DNA polymerase refers to as the rate of addition of nucleotide per second during the process of replication.
Related articles:
Conclusion:
The DNA polymerase has unmatched utility in DNA synthesis in vivo as well as in vitro. The special type of thermostable DNA polymerases such as Taq DNA polymerase and pfu are used in the in vitro DNA synthesis.
The polymerase chain reaction is a technique used for the DNA synthesis for downstream applications.
Sources:
- Garcia-Diaz M, Bebenek K. Multiple functions of DNA polymerases. CRC Crit Rev Plant Sci. 2007;26(2):105–122. doi:10.1080/07352680701252817.
- What is DNA polymerase by News medical and life science: https://www.news-medical.net/life-sciences/What-is-DNA-Polymerase.aspx.


