“Over the past few years, advances in the field of genetics are revolutionizing our knowledge about many aspects of medicine. Such a big data are helping in the research and treatment of many complex diseses.
The development of powerful computational techniques and improvements in laboratory procedures have made it possible to obtain important new information on the origin and behavior of highly frequent complex diseases, such as Alzheimer’s disease, Parkinson’s disease, diabetes, and various types of cancer, among others. All this information about health can be accessible through DNA tests.
Complex diseases are multifactorial in origin and their development is influenced by both genetic and environmental factors to a greater or lesser extent. Therefore, knowing our genetic predisposition to a certain disease can help us to modify our habits and adapt our lifestyle according to our genetics.
But how does it all start?
Key topics
- Sample processing
- Results analysis
- Big Data and genetics
- Imputation method
- Rheumatoid arthritis
- Schizophrenia
- C-reactive protein
- Logical ability
Key Topics:
Sample processing
The user receives the DNA test kit, and then the sample is collected by depositing the saliva in the corresponding tube. The shipment is processed and sent to the laboratory where it will be analyzed.
The saliva sample arrives at the laboratory, which complies with the strict ISO 90001:2015 standards of the European Union. This analysis consists of DNA extraction, quantification, amplification with PCR techniques, and finally genotyping using array-based technology.
This last step uses Illumina® iScan technology and the most updated version of the Infinium Global Screening Array (GSA) genotyping chip, which consists of about 650,000 genetic markers distributed throughout the genome, to which around 100,000 more markers have been added in a personalized way. The reliability of this technology is greater than 99% and it contains internal markers that are used as part of the quality control.
Results analysis
Once the DNA sample has been genotyped, the results are processed in the genetic companies database, where the bioinformaticians, geneticists, and doctors proceed to their analysis.
The interpretation of results is carried out by combining genetic concepts and computational power including Big Data. Thanks to several updates, some companies like tellmeGen have managed to increase the number of analyzable genetic variants to tens of millions, being able to associate them with certain conditions or diseases.
Big Data and genetics
Genetic predisposition is determined by the presence of certain variants in the genome, which can be studied by powerful computational and bioinformatic technologies using big data-based genetic studies.
Genome-wide association studies (GWAS), which have been significantly developed in recent years, are allowing an increasingly better understanding of the relationship between genetics and the development of multiple conditions and can be used to determine the genetic risk of suffering them.
In these studies, thousands of individuals diagnosed with a given disease (cases) are compared with thousands of individuals who do not suffer from it (controls), making it possible to identify risk genetic variants (those found more frequently in the case group) and protective genetic variants (those which are more frequent in the control group).
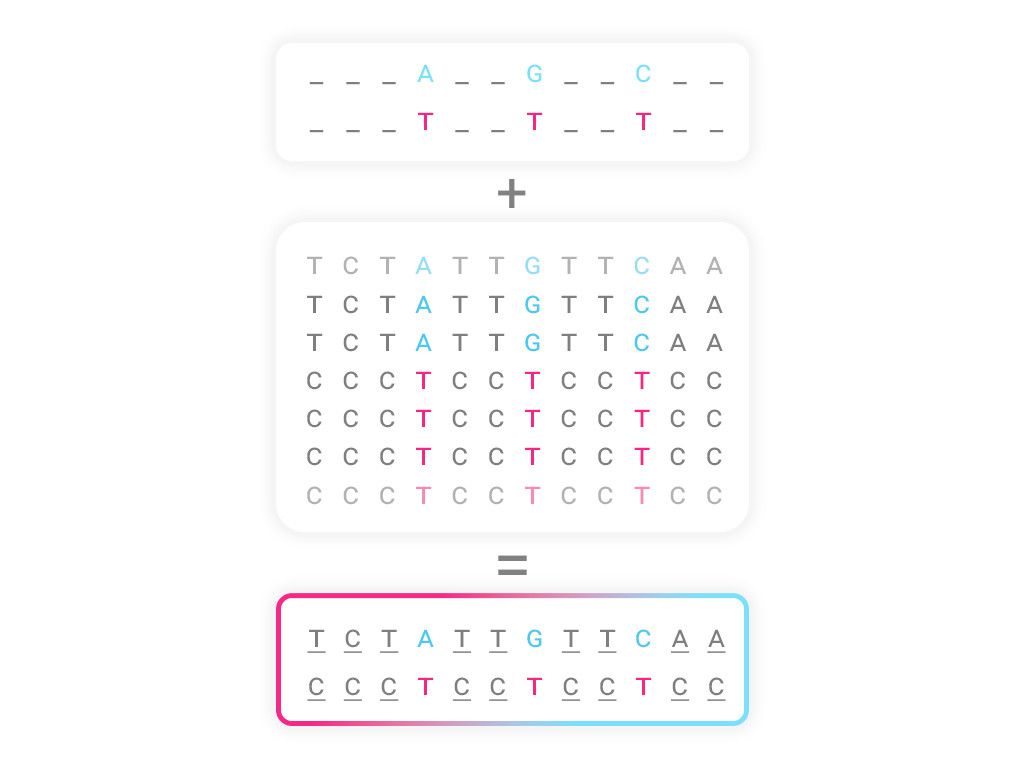
In the past few years, due to improvements in whole-genome sequencing technologies, population reference panels have been created. They are large databases containing the complete genetic information of hundreds of thousands of individuals. This has enabled the development of the imputation technique: on the basis of the information obtained with genotyping chips, such as the one used in these types of companies, this technique allows to increase the number of variants analyzed from hundreds of thousands to millions.
As shown in the table, with the imputation method the number of analyzable markers increases dramatically from 750,000 to more than 13 million. In addition, reliability is another aspect to be taken into account, as this technique guarantees a high degree of accuracy in the analysis.
Due to a large amount of information to be processed, it is essential that the computational resources applied in the imputation method are extremely powerful, increasing the sample processing time at the present time. It is important to note that, as a result of the improvements in technology, it is estimated that the sample analysis time will be reduced in the coming years.
| Without Imputation Analysis | With Imputation Analysis | |
| Number of analyzable markers | 750.000 SNPs | > 13.000.000 SNPs |
| Reliability | incomplete analysis | complete analysis |
| Computational resources | medium | high |
| Analysis duration | a few hours | 1 day |
The technical and medical basis for some of the items that are analyzed using the imputation method is provided below.
Complex diseases
They describe the genetic predisposition to a number of known diseases compared to the average risk of individuals of the same age, race and sex in a given geographic area.
Rheumatoid arthritis:
Rheumatoid arthritis (RA) is an autoimmune disease based on prolonged inflammation and characterized by pain, swelling, and joint deformity. The most commonly affected joints are those of the wrists and hands.
Through the participation of almost 23.000 positive cases, one of the latest GWAS studies carried out on this subject estimates a 60% genetic contribution, finding more than 70 loci significantly associated with RA. They were preferentially located in binding sites of various transcription factors related to CD4+ T cell biology and, to a lesser extent, other cells of the immune system.
Schizophrenia
Schizophrenia is a chronic, severe psychiatric disorder that can cause hallucinations, delusions, and severe disturbances of thought and behavior.
One of the most important studies carried out with more than 33.000 patients shows that an imbalance in neurotransmitter levels occurs due to the result of brain physiology.
The importance of the genetic factor was also determined, with an estimated contribution of 50%, and 90 susceptibility loci associated with the condition were identified.
Among them, the one that showed the strongest association was PGBD1, which encodes for a transposase highly expressed in the brain, but of unknown function. Genes like CACNA1C and CNNM2 which have proven association with brain connectivity also stood out.
Personal traits
In the same vein, with the imputation method, it is possible to analyze and learn the genetic impact of many complex traits that we wouldn’t have imagined genetics could be responsible for.
C-reactive protein:
C-reactive protein (CRP) is a protein synthesized by the liver, the level of which increases in response to inflammation. It is commonly used to determine the degree of effectiveness of treatment because of its connection to the immune system and it plays a part in the defense system against infections.
Its levels, like many other biomarkers commonly examined in urine, are affected by both genetic and environmental factors.
By means of one of the largest GWAS-type studies conducted to date, involving 355,000 individuals, 91 markers have been associated with variability in CRP levels. Among the most relevant genes are the CRP gene, which is responsible for the production of C-reactive protein in the liver, and the ubiquitously expressed HNF1A gene, whose mutations have also been linked to pathologies such as diabetes.
Logical ability:
Logical ability is the capacity that allows us to think and reason quickly when faced with a problem, without prior information and experience. Its measurement can be very useful in assessing a person’s psychological capacities.
It is influenced by genetics and other environmental variables such as neurophysiology. Also, growing up as a child in a nurturing environment promotes the development of neural connections in the regions of the brain associated with memory, learning, and spatial orientation.
One of the most recent GWAS-type studies conducted has identified 50 genetic markers associated with logical ability, including almost 120.000 people. The most important analyzed genes are SLC39A8, AFF3, and the ARTN gene, which are involved in transporting trace elements, neurodevelopment and intellectual disability, and neuron survival, respectively.
Wrapping up:
In summary, the implementation of this powerful and novel methodology has made it possible to reshape the estimation of genetic predisposition to develop complex diseases, personal traits and others, providing much more accurate and up-to-date results as new discoveries in genetic technology and research are published. And through the analysis of your DNA, all this information can be in your hands.
Bibliography:
Ha E, Bae SC, Kim K. Large-scale meta-analysis across East Asian and European populations updated genetic architecture and variant-driven biology of rheumatoid arthritis, identifying 11 novel susceptibility loci. Annals of the Rheumatic Diseases. 2021 May;80(5):558-565.
Giannini D, Antonucci M, Petrelli F, Bilia S, Alunno A, Puxeddu I. One year in review 2020: pathogenesis of rheumatoid arthritis. Clin Exp Rheumatol. 2020 May-Jun;38(3):387-397.
Smolen JS, Aletaha D, Barton A, Burmester GR, Emery P, Firestein GS, Kavanaugh A, McInnes IB, Solomon DH, Strand V, Yamamoto K. Rheumatoid arthritis. Nat Rev Dis Primers. 2018 Feb 8;4:18001.
Yao X, Glessner JT, Li J, et al. Integrative analysis of genome-wide association studies identifies novel loci associated with neuropsychiatric disorders. Translational Psychiatry. 2021 Jan;11(1):69.
Marder SR, Cannon TD. Schizophrenia. N Engl J Med. 2019 Oct 31;381(18):1753-1761.
Sinnott-Armstrong N., Tanigawa Y., et al. Genetics of 35 blood and urine biomarkers in the UK Biobank. Nature Genetics, 18 Jan 2021, 53(2):185-194
Nehring S.M., Goyal A., et al. C Reactive Protein. StatPearls
Sproston N.R. et Ashworth J.J. Role of C-Reactive Protein at Sites of Inflammation and Infection. Front Immunol. 2018; 9: 754.
UK Biobank Database. Fluid intelligence score, Data-Field 20016 [Version Jan-2022]
Kent P. Fluid intelligence: A brief history. Appl Neuropsychol Child. 2017 Jul-Sep;6(3):193-203. doi: 10.1080/21622965.2017.1317480. Epub 2017 May 16. PMID: 28506124.
Cochrane A, Simmering V, Green CS. Fluid intelligence is related to capacity in memory as well as attention: Evidence from middle childhood and adulthood. PLoS One. 2019;14(8):e0221353. Published 2019 Aug 22.
Li Y, Willer C, Sanna S, Abecasis G. Genotype imputation. Annu Rev Genomics Hum Genet. 2009;10:387-406. doi: 10.1146/annurev.genom.9.081307.164242. PMID: 19715440; PMCID: PMC2925172.
Naj AC. Genotype Imputation in Genome-Wide Association Studies. Curr Protoc Hum Genet. 2019 Jun;102(1):e84. doi: 10.1002/cphg.84. PMID: 31216114.
Davies RW, Kucka M, Su D, Shi S, Flanagan M, Cunniff CM, Chan YF, Myers S. Rapid genotype imputation from sequence with reference panels. Nat Genet. 2021 Jul;53(7):1104-1111. doi: 10.1038/s41588-021-00877-0. Epub 2021 Jun 3. PMID: 34083788; PMCID: PMC7611184.