The junk DNA part- the 80% mystery has an important biological function, claimed by the authors of ENCODE, the encyclopedia of non-coding DNA.
Because DNA makes proteins, it is important to us, a partial truth we all know, but that’s not DNA, those are genes- a definite part of the genome capable of making proteins such as enzymes, hormones, receptors and immunoglobulins etc.
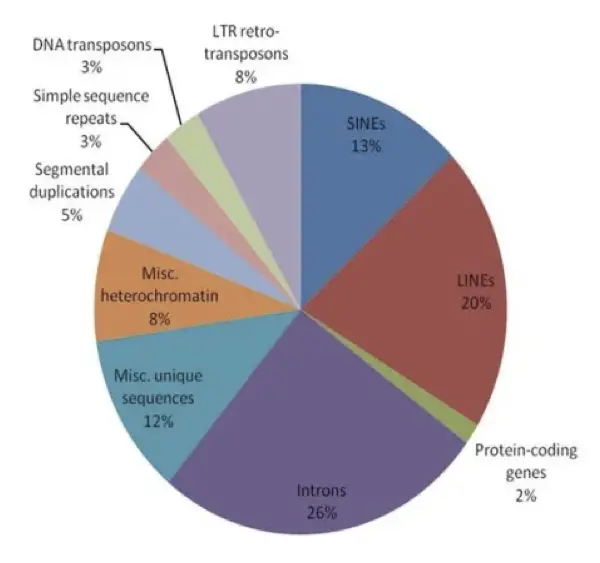
Interestingly, protein-encoding genes are only 3%, maximum. The genome comprises around 22,000 to 25,000 genes. A huge portion is junk DNA which is around 97% and is non-coding.
The genetic study mainly covers three things- genes, genome and chromosomes. Let me explain to you all things in a single paragraph.
Genes are the coding DNA sequences, making proteins and the genome is a whole set of an organism’s cell’s DNA. The entire genomic DNA is located on 23 pairs of chromosomes.
The DNA is nothing but the sequence of nucleotides only- a long polynucleotide chain made up of sugar, phosphate and nitrogenous bases. A, T, G and C are four types of nucleotides DNA has and are paired complementary. A paired with T and G paired with C.
If you wish to learn more you can search articles in the search box, we have so many articles on related topics. Now coming to the present topic. Why is so-called junk DNA important for us?
Key Topics:
Importance of Junk DNA:
Previously it was believed that the sequences which can’t make proteins are useless, meaning, are only junk and that’s why scientists named it as “Junk DNA”.
Unlike the coding genes, noncoding DNA has several definite functions in our genome which are more important than only encoding proteins, recent studies say.
Regulating gene expression, transcription and translation, making elements of chromosomes and providing regulatory elements during central dogma processes and many more are functions they have.
Even though it’s categorized as junk, is indeed useful, henceforth it’s super powerful.
Superpowers of Junk DNA:
This section of the article covers various functions of junk DNA in our genome.
Regulating gene expression is a vital function that must be performed correctly. In how much amount a gene makes a protein in different cells or tissue is referred to as gene expression. The below example makes things clear,
A gene for keratin has a significant role in hair and skin growth but doesn’t have any role in liver cells, RBC or WBC. So a gene for keratin expresses higher in skin and hair cells rather than liver cells, WBC or RBC, getting the point!
That is what the non-coding DNA does do, regulates the process of gene expression. Specialized sequences and genes are assigned to do so. Using many ways the process completes. Here I am explaining some.
Pseudogenes- a type of non-coding/ junk DNA has been assigned to produce a special kind of RNA. Meaning, it can transcribe into RNA (known as siRNA) but can’t make protein from it.
Smaller interfering RNAs and microRNAs are double-stranded short RNA sequences having a significant role in gene regulation. It binds to the target mRNA sequence and degrades it.
This means that a final protein product can’t be formed. The whole mechanism is known as RNA interference. RNAi is an entire process specially designed by a cell including pseudogenes, guided-RNA, RISC, Argo Protein and other elements to regulate gene expression.
Moreover, other RNA forms such as transfer RNA, regulatory RNA and ribosomal RNA are also formed by the noncoding DNA. Again, the tRNA and rRNA have a crucial function in protein synthesis.
So non-coding/ junk DNA controls the expression of every gene thereby it controls the expression of the whole genome. Transcriptomic studies rely on junk DNA analysis. It switches “On/Off” gene transcription by allowing and disallowing enzymes.
The whole set of transcripts can be studied and quantified in order to know how much junk DNA is active or not. Scientists study Histone modifications, interchromatin interactions, methylation patterns, chromatin remodeling and other protein DNA interactions to investigate junk DNAs.
Methylation PCR, ChIP-seq, ChIP- Chip and chromosome conformation capture are several techniques researchers employ to study and investigate how junk DNA controls gene expression.
Regulatory elements are noncoding sequences having critical functionality in transcription and translation. How does it work? See the image,
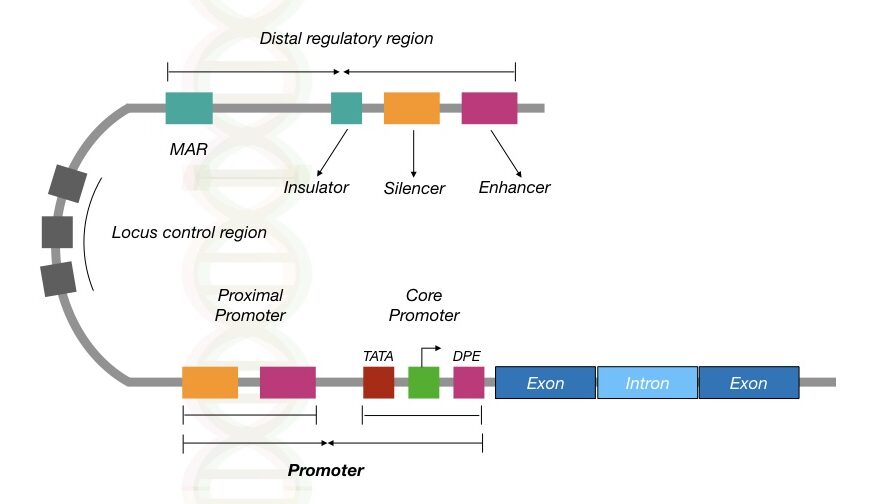
Regulatory elements are of two types, cis and trans-regulatory elements which are located near the gene, before a gene, away from the gene or at a location where enzymes bind.
Enzymes such as polymerase or acetylase recognize the regulatory element assigned and binds to it. Once it settles, it starts the catalytic reaction.
Replication, a process to synthesize DNA is initiated once enzymes recognize the origin of replication. Often known as “Ori” or “OriC” are the sequences present on multiple locations in the genome, recognizable to the polymerase.
After catching it, polymerase immediately starts the process of DNA synthesis. OriC, promoters, suppressors, insulators and regulators are elements that help in various biological processes.
Another important role Junk DNA has is forming structural chromosomal elements. Interaction between DNA and protein forms chromosomes having three structural elements- centromere, telomere and arms.
On the “p” and “q” arms vital genes are located. Genes must be expressed during translation while the telomere and centromere provide structural stability to chromosomes and protects genes from replication errors.
The centromere is a tightly wrapped protein-DNA structure that joins four chromatids together, located centrally. While the telomeres are located far away on the edges of each chromatid, cover arms and protects from degradation.
Both structures have non-coding, repetitive, GC-rich DNA sequences known as satellites.
Microsatellite and minisatellite are two types, have smaller and larger repeat DNA, respectively and have been used in many clinical practices. How satellite DNA employed in forensic science will possibly discuss later in this article.
Or this article may help you to learn more: DNA Fingerprinting- Definition, Steps, Methods and Applications.
Remember, we are discussing roles of “so-called” junk DNA or non-coding DNA, and it is yet not over, I am just reminding you. Ok moving ahead.
Scaffold or matrix attachment regions (S/MARs) separate chromatins into structural and functional domains thereby regulating gene expression. Technically, the S/MAR is a DNA region that makes a bridge between the nuclear matrix and chromosomal DNA.
Scaffold or matrix attachment regions are actually non-coding/ Junk DNA elements.
Triggering or suppressing genes is yet another function. If a gene doesn’t have a proper role, affecting other genes negatively, producing abnormal protein or not making any protein, those genes are worthless.
Junk DNA elements immediately inactivate those genes using one of the many available mechanisms. For example, a transposon moves and inserts in a gene, changes its structure and makes it unavailable for transcription or translation.
Similarly, it can also trigger functions of some nonfunctional genes. Again transposons can make it possible by moving away from the native gene. I am giving you a simple explanation to make you understand.
Yet another crucial function is to maintain genetic integrity.
The genome is located in the cell nucleus and uses it as a pocked, wrapping tightly inside. Interaction of non-coding DNA with proteins makes it possible to settle in the nucleus.
Studies conducted by Yamashita Y and coworkers on Drosophila melanogaster reveal these facts. When they have removed a specific protein that interacts with the DNA, the nucleus can’t form correctly and the cell dies.
This protein probably has a role in DNA packaging and with whom it interacts, are the non-coding sequences.
Junk DNA such as transposable elements has been assigned by our cells to control the DNA loss, especially the Junk DNA. Recent findings show that cells lose noncoding DNA gradually. DNA is fading faster than gaining.
For example, at the end of replication, some part of satellite DNA can’t be copied correctly. Resultantly, it can’t be inherited to new cells thereby gradually decreasing the length of satellite DNA.
However, the telomerase somehow repairs it, but transposons have a more significant role in the overall regulation of the length. A study conducted by Josep M Comeron shows that since evolution, transposons move in the genome, switch on and off genes and also regulate the length of noncoding DNA.
The length of the telomere is supposed to be directly proportional to the age of an organism. After every cycle of replication, telomeres lose some amount of genetic material, gradually it shortens and the organism becomes aged.
However, as we said, the telomerase enzyme somehow adds several sequences and repair them. Telomeres are made up of satellite, non-coding/ junk DNA.
Noncoding DNA gives a signal where and when genes get expressed.
These all are existing functions currently the junk DNA performing in everyone’s genome, interestingly it does have an evolutionary role as well. Junk DNA has significant importance in the process of evolution, natural selection and mutation.
Let’s discuss that,
Role of junk DNA in evolution:
Sequences of the junk DNA are conserved since evolution which has a positive selection effect. In fact, some are conserved over millions of years. That’s how they preserve several biological functions.
“Genome sequencing data of humans, plants, nematodes and fruit flies postulates that the number of protein-coding genes between these organisms is more similar than expected.
Shockingly, the substantial differences have been shown in the amount of junk DNA, not protein-coding genes.” These results were published on Oct 20, 2005, in Nature by Andolfatto P and their colleague. Peter Andolfatto is an assistant professor of biology at UCSD.
Studies conducted on drosophila also show that this region is evolving slowly and having higher mutation resistance power than coding regions. Coding genes are more prone to mutation.
This implies that non-coding DNA has more powerful roles even if it could not make proteins and that is why many non-coding sequences are conserved between species, organisms and since evolution. Even several sequences have been conserved for millions of years in organisms.
Contrary to this, another portion of junk DNA is hypervariable too. Hypervariable regions have a variable number of repeats (not sequence) even between two individuals of the same species.
For example, the number of hypervariable regions varies even between two human individuals. Hypervariable regions, a variable number of repeats such as VNTR and STR though do not have a role in protein making but regulates gene expression.
Junk DNA and biodiversity:
According to scientists, junk DNA also provides biodiversity as well as protein diversity. Junk DNA has the power to “where and when to express genes.” Moreover, it also changes the structure, function or way a gene behaves.
Intervening sequences, introns, transposons and noncoding RNAs continuously affect a gene thereby changing either its structure, function or makes it nonfunctional.
Let us imagine a hypothetical situation in which some short of introns get lost, which results in altered gene structure. This may change the function or how the gene behaves.
Studies also show that due to the continuous effort of all noncoding DNA, biodiversity within the species and between the species are originated. Meaning, different gene sets, expression profile and varied proteins are formed.
Junk DNA has a role in cell defense too. Pseudogenes make RNAs having a role in defense. And are involved in the RNAi pathway- siRNA or miRNA. What it does do is finds the pathogenic mRNA binds to it, activates the pathway and degrade it.
Summary:
The functions of noncoding DNA:
- Regulating expression of genes
- Regulating genome expression
- Maintaining genome integrity
- Providing structural elements to chromosomes
- Work as a scaffold-matrix attachment region
- Helps in evolution
- Switch on and off genes
- Provides origins of replication
- Work as regulatory elements during transcription and translation
- Controls the process of mRNA and protein formation.
- Protects against foreign pathogens.
- Provides biodiversity in nature
- Prevent aging
Wrapping up:
We can’t wrap things here! Junk DNA is a huge, unexplored and understudy subject in genetics. Though we do not know more, it has many functions to perform and that’s why a huge portion is assigned in an organism’s genome.
So now we can say that the junk DNA isn’t actually junk, precisely it is only non-coding DNA that can not make protein.
A huge portion of the genome is repetitive and conserved, scientists still can’t rectify why it is there and why it’s repetitive. But everything in our genome has a definite role.
Change in the number of repeats can cause serious health issues, even though it isn’t coding. Myotonic dystrophy, Huntington’s disease, ataxia and Fragile X syndrome are conditions that occur by the change in the number of different repeats.
I think this article may have covered all the essential information related to junk DNA. We can now say this part is very important for us to survive on earth.
Sources:
Mariusz Nowacki, Brian P. Higgins, Genevieve M. Maquilan, Estienne C. Swart, Thomas G. Doak, and Laura F. Landweber. A Functional Role for Transposases in a Large Eukaryotic Genome. Science, 2009; 324 (5929): 935 DOI: 10.1126/science.1170023.
The University of California – San Diego. “UCSD Study Shows ‘Junk’ DNA Has Evolutionary Importance.” ScienceDaily. ScienceDaily, 20 October 2005.
Palazzo AF, Gregory TR. The case for junk DNA. PLoS Genet. 2014;10(5):e1004351. Published 2014 May 8. doi:10.1371/journal.pgen.1004351
Joseph M Comeron. What controls the length of noncoding DNA? Current Opinion in Genetics and Development 2001; 11(6): 652-659.


