“cDNA short of complementary DNA is a piece of DNA synthesized from the mRNA by the enzyme reverse transcriptase which has important applications in genetic research.”
Do you know that the only virus known as “retrovirus” can manufacture the cDNA from mRNA, whereas all other organisms on earth make mRNA from DNA?
DNA and RNA are two well-known and precisely-studied nucleic acids present on earth. Where most organisms have DNA, only a few organisms (viruses) have RNA as their genetic material.
However, why only a small group of viruses have the RNA as their genetic material is still a mystery but can do substantial damage when infects us. For example, HIV is an RNA virus.
The universal mechanism referred to as ‘central dogma’, which manufactures a protein by the process of replication, transcription and translation, sequentially, is DNA- mRNA and protein. The process is vital and unidirectional in all eukaryotes as well as prokaryotes (Except some viruses).
DNA to mRNA the process known as transcription gives further signals by removing non-coding sequences from the DNA and converting them into specialized RNA which is mRNA.
Broadly, the three types of RNA we have; are mRNA, tRNA and rRNA among all, the mRNA or messenger RNA give signals for tailoring the protein, whereas tRNA and rRNA help in the process of transcription.
But here we are talking about the reverse process, a process very specific and rare in nature that forms DNA from RNA, using a special type of enzyme. The reason behind studying it is its lucrative applications in proteomic studies, transcriptomic analysis and quantification.
But how exactly we can mimic this process in vitro? that’s what we are talking about in this article. Along with that, we will also explain the process of cDNA synthesis and cDNA library preparation.
Stay tuned.
Related article: Genomic DNA Library- Preparation and Applications.
Key Topics:
What is cDNA?
The cDNA acronym as complementary DNA is different from the total genomic DNA which has only coding DNA sequences.
Definition of cDNA:
cDNA is a functional coding segment of DNA made from the mRNA or transcript by using a specialized enzyme known as reverse transcriptase.
Principle of cDNA synthesis:
cDNA synthesis is a process, often referred to as reverse transcription, is governed by the enzyme reverse transcriptase and forms cDNA from the mRNA. The whole process is technically also named reverse genetics.
Naturally, it has been present in viruses like Lentivirus, HIV-1, Murine Leukemia Virus, Feline immunodeficiency virus, etc, broadly categorized into a retrovirus.
The polymerization property of the reverse transcriptase enzyme identifies each nucleotide from the mRNA and simultaneously forms the cDNA by using the primer and incoming nucleotides.
However, the in vitro process of cDNA synthesis, though somehow similar to the natural one but is a bit different in terms of technical processing. It is divided into the first and second-strand synthesis.
In the upcoming section, we will understand the in vitro process of complementary DNA synthesis.
Process of cDNA synthesis:
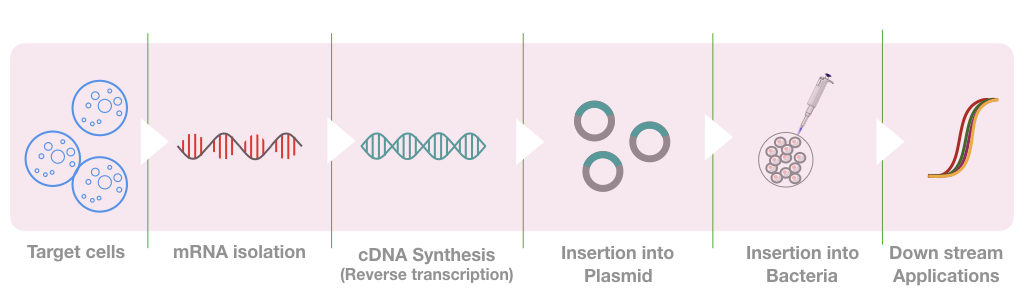
The whole process can be divided into steps like RNA isolation, mRNA isolation, mRNA purification, enzyme selection, reverse transcriptase PCR, Library preparation and library screening
Steps in cDNA synthesis:
mRNA isolation:
cDNA is a complementary sequence to our transcript, first, we have to isolate the whole RNA part from the sample and then the mRNA. A total RNA can be isolated using a standardized RNA isolation protocol or ready-to-use direct mRNA isolation kit.
A specialized isolation kit is required for mRNA isolation which can isolate only the mRNA from the rest of the RNA. dedicated mRNA isolation kits usually contain oligo (dT)-containing columns.
Once the mRNA is transcribed, the poly-A is added (artificial post-translational modification) to differentiate the messenger RNA from the rest of the RNA.
The poly-A tail of the mRNA remains bounded with the oligo dT containing column. After each round of washing, all the RNAs are washed off; only the mRNA remains in the column.
In the final step, the mRNA is collected in another tube by using the elution buffer.
Related article: What is Poly-A tail?- Definition, Structure and Function.
mRNA purification:
Purified mRNA must be required for the next step, for that, the mRNA is purified using the purification kit. Commonly oligo-dT complementary nucleotides are removed from the mRNA as not required further.
Gentle heating can do the job. The purified mRNA is then quantified and sent for reverse transcription PCR.
An important consideration for cDNA library preparation:
- Use DNase to remove DNA contaminants from the RNA.
- Perform the RNA isolation experiment at 4C to get good results.
- Use DEPC water to wash all the utilities during the experiment.
- Use autoclaved tips or tubes, do not reuse them.
Selection of enzyme- reverse transcriptase
A normal DNA polymerase can’t synthesize DNA from RNA. As we discussed, another polymerase known as reverse transcriptase is used, instead. A reverse transcriptase enzyme is a special type of polymerase isolated from the retroviruses having the power to synthesize cDNA from the mRNA.
The Avian Myeloblastosis virus reverse-transcriptase and Moloney Murine Leukemia Virus reverse transcriptase are two commercially available RTs used commonly in the cDNA library preparation.
Reverse transcriptase PCR:
Normal PCR is used for the synthesis of DNA from DNA while reverse transcriptase PCR is applicable for the synthesis of DNA from the RNA template using the reverse transcriptase enzyme.
DNA is synthesized back from the transcript or mRNA but the steps of the RT-PCR are almost similar to normal conventional PCR.
In the process, fist a single cDNA template is synthesized (single-trand synthesis) using which the ds(c)DNA is formed (second strand synthesis). The whole process, primers, probes and reaction preparation for reverse transcription PCR is explained in our previous article.
Read more on reverse transcriptase PCR: Reverse transcription PCR: Principle, Procedure, Applications, Advantages and Disadvantages.
Construction of library:
Now we have the amplicons of a cDNA, the cDNA is now inserted into the plasmid using the restriction digestion method.
The sticky ends are generated on a plasmid that binds complementary to the sticky ends of cDNA, these sticky ends are generated using the restriction digestion method.
Depending upon the size and type of cDNA, different types of plasmids are used for constructing different cDNA libraries. The plasmid with the cDNA is now inserted into the bacteria and grown using the nutrient media under aseptic conditions.
What we have now in our BAC (bacterial artificial chromosome) is known as a library of cDNA or cDNA library which has all the fragments of our interest. We can use it anytime. To use it in downstream applications, plasmid DNA is first isolated and processed as per protocol.
cDNA synthesis and cDNA library preparation are both topics that are actually a part of the whole process, so I think it does not make sense when discussed separately. So here in the upcoming section, I am explaining the cDNA library preparation process in-depth.
What is a cDNA library?
What is a library?
A collection of books or literature, more precisely, an organized collection of books or literature, right.
The cDNA library is similar to the original terminology, somehow, an organized collection of cDNA, in fragments, for a purpose. cDNA library is a great ‘tool’ I can say for transcriptomics, gene expression and proteomic studies.
For example, we can’t experiment on a real human subject, then how can we study a particular protein, its effect or optimization?
Through cDNA library.
The mRNA for the protein we wish to investigate is reverse transcribed into cDNA, fragmented and transferred into bacterial cells. Where it forms the exact same protein (that’s a simple explanation!).
cDNA library has some outstanding and lucrative applications to investigate individual transcript or whole transcriptomics.
cDNA library preparation:
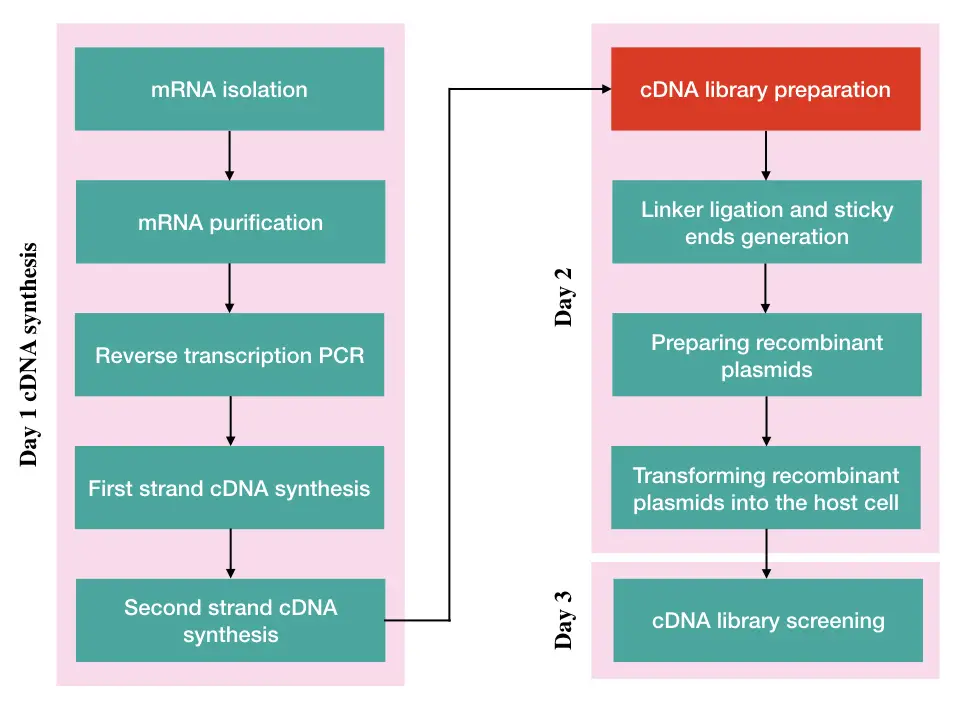
Steps in cDNA library preparation:
- mRNA isolation and purification
- cDNA synthesis
- Reverse transcription PCR
- Fragmentation, linker adaptation and sticky end generation
- Preparing recombinant plasmid vector
- Transforming recombinant plasmids into the host cell
- Screening cDNA library
cDNA library as we said is the collection of specific DNA fragments, and so the steps of cDNA synthesis for library preparation remain the same. The process starts with mRNA isolation, purification and reverses transcription PCR.
You can read this section above.
The actual process of library preparation is initiated after reverse transcription PCR. The free OH end generated by the use of oligo (dT) primers is trimmed off using the S1 nuclease. It is a type of restriction enzyme, isolated from East Asian fungus having an activity to cut rest or unamplified ssDNA from the dsDNA.
After this step, we have blunt-ended ds(c)DNA fragments which are inefficient to incorporate into the plasmid. To generate an effective cDNA fragment library, we need sticky ends. Although, randomly prepared sticky ends don’t work either.
Linker ligation and sticky end generation:
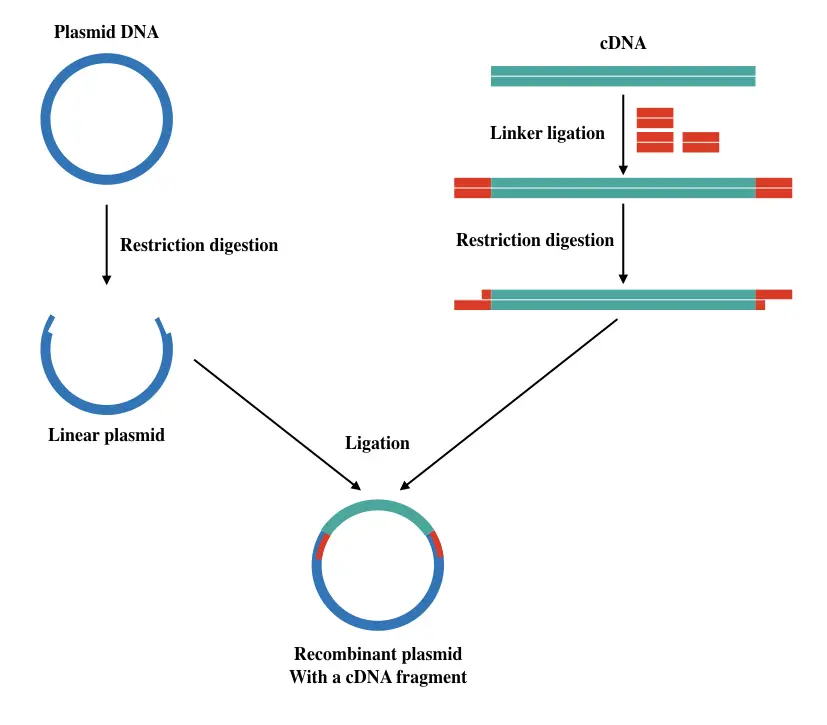
Sticky ends are produced by a special type of restriction endonuclease whose ends have a few base ssDNA. To generate a specific type of known sticky-ended cDNA, linker oligonucleotides are first added to the cDNA.
Enzyme ligase seals the gaps between linker and cDNA, followed by restriction digestion. Note that linkers are attached at ends of cDNA.
Preparing recombinant plasmid vector:
The plasmid is circular and double-stranded DNA that replicates into the bacteria. Plasmids are used as a vector or vehicle to transfer DNA into the host cell and are the effective ones.
Once the plasmid starts replicating into the bacteria, the DNA or cDNA we incorporated also starts replicating. In the end, we get copies of our desired DNA fragment- the library.
To go ahead in the process, the circular plasmid is digested with the same restriction enzyme used to digest the cDNA + linker. The idea is to generate complementary sticky ends on both plasmid and cDNA.
Once the plasmid gets linearized, our cDNA is ligated using the ligase enzyme into the plasmid. To make the experiment screenable, an antibiotic gene as a marker is also added near to the cDNA.
Sometimes, a second marker gene is also added.
Now the recombinant plasmid is ready for the transformation.
Transforming recombinant plasmid into host cell:
Practically, we have mixed the solution of cDNA + linker, digested plasmids and ligase enzyme to allow recircularization of plasmid along with cDNA and marker gene, however, not all the plasmids become recombinant.
So to select those only having our cDNA, we have to transform and express it into bacterial cell culture. The recombinant plasmid is now transformed into bacteria cells and allowed to culture in a specific antibiotic-rich media.
Here the idea is to grow cells that only have the antibiotic-resistant gene, meaning transformed cells only. At the end of cell culture, we will get bacterial cell colonies with the recombinant plasmid.
The recombinant plasmid DNA is now isolated and stored. It is our cDNA library.
Screening of cDNA library:
The final step in the cDNA library preparation is screening, validating whether a correct cDNA and/or a correct plasmid is constructed and transformed or not.
The first layer or primary screening method is transformation itself and replica plating, helping us to select only transformed cells.
Other molecular genetic techniques such as different types of PCR are also used for screening purposes, which indeed is a more accurate and faster method.
For example, plasmid or cDNA-specific primers are used to amplify the recombinant plasmid DNA- the nested PCR approach.
Quantifying the amount of transformed recombinant plasmid- RT-PCR approach.
Evaluating only the cDNA- conventional PCR approach.
Once the screening process completes, our cDNA library for a specific mRNA is ready to use for mRNA sequencing, whole-exome sequencing, mRNA microarray or gene expression studies.
Optimizations and recent advancement:
cDNA synthesis and library preparation process is a tedious and time-consuming job (so is boring, sometimes!) Therefore scientists timely optimize protocols to overcome shortcomings. Remember cDNA synthesis and library preparation are crucial techniques in genetic and genomic studies.
Goldman S et al., 2015 also published another protocol for preparing cDNA libraries for high throughput DNA sequencing. They have modified the 5’-OH end of the cDNA to sequence each cDNA fragment.
Figueiredo A et al., 2007 published a protocol that eliminated the need for restriction digestion, cloning and subcloning steps and obtained high-quality cDNA libraries.
Both the protocols explained are effective and time-saving.
Advantages and Applications of cDNA library:
- It provides ease, accuracy and effectiveness during the experiment as noncoding and other nonrequired DNAs are removed.
- The cDNA can directly be expressed into bacteria to investigate protein function.
- It is useful in reverse genetic studies.
- One of the important applications of the present technique is to clone and study low copy number genes.
- The expression of a eukaryotic protein can be evaluated into bacteria cells and investigate its effect.
- The cDNA libraries are highly required for techniques in sequencing and microarray analysis.
Limitations of cDNA library:
One of the major limitations of the present library preparation technique is that we can only use the transcriptomics part of the genome. Meaning, It allows only mRNA, gene expression and proteomics studies.
Other genomic sequences can’t be investigated, for example, non-coding sequences, pseudogenes or other non-coding DNA sequences. We need to prepare a genomic library to investigate, study and evaluate the whole genome.
It is a time-consuming, costly and tedious process.
Wrapping up:
cDNA has significant importance through which a protein and its association with a DNA or gene can be studied. It is used in sequencing and as well as microarray analysis. The whole ‘transcriptomics’ technology relies on the cDNA synthesis and library.
Currently, rapid advancement in PCR technology allows us to prepare cDNA and libraries with ease and accuracy, though the technique is still time-consuming and costly. The original experiment consists of a lot of enzymatic reactions, PCR amplifications and chemical treatments.
Sources:
L. B. Klickstein, R. L. Neve, E. A. Golemis, J. Gyuris. Conversion of mRNA into double-stranded cDNA. Curr Protoc Mol Biol. 2001 May; Chapter: Unit5.5.
Vvedenskaya IO, Goldman SR, Nickels BE. Preparation of cDNA libraries for high-throughput RNA sequencing analysis of RNA 5′ ends. Methods Mol Biol. 2015;1276:211-228. doi:10.1007/978-1-4939-2392-2_12.



