The linking number is a topological property of DNA. A linking number is a sum of twists and writhes. The number of times one strand of DNA turned around another strand is called a twist while inter-coiling of the double helix is termed as writhe. In short, writhe is a number of a time DNA double helix is crossed, coiled over each other or the number of times one strand wraps around another strand.
As per the Watson and Cricks model, eukaryotic DNA is right-handed and negatively supercoiled. It is very difficult to understand DNA topology in two-dimensional space because topology is a science that cannot be understood without three-dimensional property.
For understanding the linking number, We can assume the winding of the telephone wire model. The same pattern DNA follows during packaging or supercoiling. For more detail on DNA packaging read the article, DNA packaging in eukaryotes.
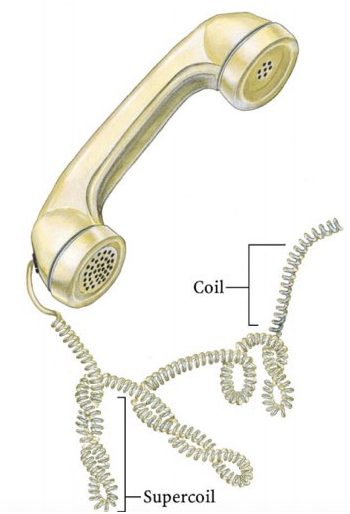
The image represents supercoiling in telephone wire.
DNA writhe is more important, as it will help in the arrangement of DNA. Here writhe can be interwound or overwound. Interwound writhe is plectonemic which means it wraps around each other or over the axis.
While over-wound writhe is spiral. The classic example of spiral winding is histone and DNA assembly, dsDNA is wrapped around histone in a spiral manner during nucleosome formation.
Key Topics:
DNA Linking Number
The sum of the total number of twists and writhes are called as a linking number.
If both ends of DNA are joined, the linking number remains unchanged, (the linking number is zero in the case of circular DNA molecule). So if we want to change the linking number, we have to break the DNA strand and that function is performed by DNA topoisomerase.
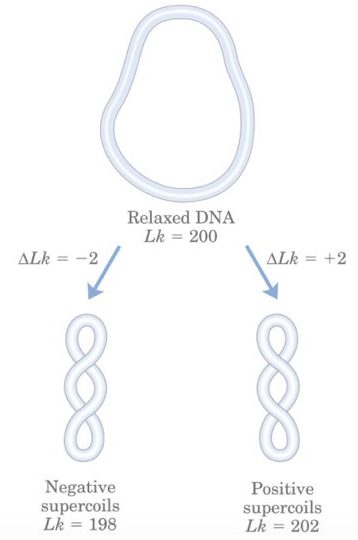
Importantly, twist and writhe can be interconvertible, in cccDNA under some distortion conditions some of the twists can be converted into writhe and some of the writhes can be converted into the twist. However, the total of both remains the same (because, Lk= Tw + Wr).
Eukaryotic DNA is B form, right-handed and under-wound which means it is negatively supercoiled. Left-handed DNA is positively supercoiled and overwound (not prevalent in nature). We will discuss later in this part why negative supercoiling is important.
In DNA supercoiling, writhe is also dependent on the twist. As we know the writhe is dependent on the axial stress of DNA, when the number of twist increase, it increases the torsional tension on the axis of DNA and ultimately the writhe will increase. This change makes DNA supercoiled and packed even tighter on the chromosome.
In cccDNA, writhes remain zero as there is no inter-coiling between dsDNA observed. here, if,
Lk= Tw + Wr
For cccDNA (writhe is equal to zero)
So, Lk= Tw.
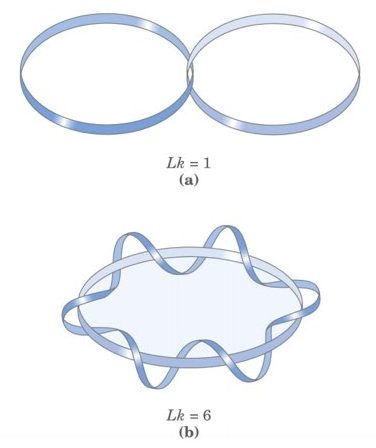
Now imagine that the cccDNA is fully relaxed which means it does not have any supercoiling, in this condition the linking number becomes zero hence supercoiling in cccDNA is denoted as Lk0 .
As per the Watson and Crick model, the dsDNA has 10.5 bp per helix and is relaxed. The relaxed cccDNA has a total of 1050 bp so the total linking number is,
Linking number (Lk)= total number of base/basepair per helical turn
hence, Lk= 1050 bp / 10.5 (bp/turn)
Lk= +100 (because the DNA is other than B form and left-handed)
Elaborately, it is indicated that if our DNA is left-handed and has 10.5 bp per turn with 1050 bp length then one strand is crossed or over-wound (positive supercoiling) to another strand 100 times.
However, it is just an assumed example because fully relaxed DNA (or positively supercoiled DNA) does not exist in nature. When we repeatedly decrease the twists and writhes by cutting with an enzyme, gradually linking number decreases and becomes zero.
Therefore, the difference between the linking number (Lk) and zero linking (Lk0) is called as linking difference ΔLk,
So, ΔLk = Lk – Lk0
If the value of ΔLk is different from zero ( >0 or <0 ) then the DNA is called as supercoiled. Additionally, Lk is lower than Lk0 (the linking number is less than zero), ΔLk is also <0 and the value which we get is negative so the DNA is called negatively supercoiled.
Similarly, if Lk >Lk0 then ΔLk > 0, and the value obtained is positive so the DNA is called positively supercoiled. Here the value of ΔLk is dependent on the length of DNA. If the length increase, supercoiling will increase and vice versa.
Specific linking differences are another important property of supercoiling. It is also called superhelical density which is a difference between linking difference (ΔLk) and linking the number of cccDNA (Lk0). So,
Superhelical density= ΔLk /Lk0
Some Facts on Linking number
- Human DNA is B type and right-handed with negative supercoiling. Majorly, all the organisms (from bacteria to eukaryotes) on earth are negatively supercoiled, However, some of the thermophilic bacteria are positively supercoiled.
- Negatively supercoiling stores energy, which is utilized during the denaturation of dsDNA in replication or transcription. Positively supercoiling stores more energy hence even at higher temperatures DNA does not denature. Even the enzyme cannot recognize positively supercoiled DNA and is unable to cut it.
- The length of DNA from 46 chromosomes is approximate ~2 meters and it has to fit in ~10µM radium nucleus now imagine why supercoiling is important.
- If we stretch dsDNA from both end it is approx. ~100km long.
- Globally, the DNA of all living organisms is supercoiled on an average of approximately ~6%.
Identification of different forms of DNA
Agarose gel electrophoresis is a technique by which we can identify the different forms of DNA molecules.
If we have four DNA molecules of the same length with different topologies, each molecule migrates differently in agarose gel.
Suppose the first DNA is closely circular with no supercoiling, and the second DNA molecule has both strands twisted around each other which means linear double-stranded DNA. The third DNA molecule is coiled around each other with some writhes and the fourth one is supercoiled.
In each case, DNA molecules migrate differently. The more compactly supercoiled DNA migrates faster in the gel. Hence the fourth one (which is supercoiled and compact) migrates faster as compared with other DNA (Lane C and D: the band at the base).
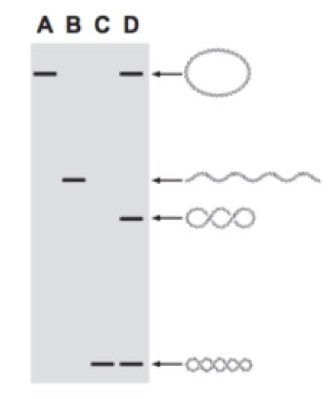
The first one, circular DNA does not have any supercoiling hence it migrates slower than any other DNA and remains last (lane A and D).
The more compact the DNA is, it can easily pass through agarose pores. So sequentially, circular DNA remains last, linear DNA cover more distance than circular DNA and coiled DNA covers more distance than linear DNA. At last supercoiled DNA cover more distance than any other DNA, and run faster.
Read more: What is rcDNA?- rcDNA vs cccDNA.
Topological properties especially the linking number of DNA are very important properties of DNA if it is not maintained properly in vivo, DNA cannot be packed properly on the chromosome which will result in epigenetic alteration and failure in replication.
The linking number of the DNA can be calculated using the formula given above, feel free to comment here and let me know how linking numbers can be useful for the diagnosis of disease.


I am lucky that I found this web blog, precisely the right information that I was searching for!
Thank you