An exclusive native PCR modification known as Asymmetric PCR has been applied to amplify only a single strand during the PCR.
PCR is a thermocycler, amplifies DNA. Meaning, it helps to synthesize DNA artificially which aids in genetic studies. It has many different variants to execute different purposes.
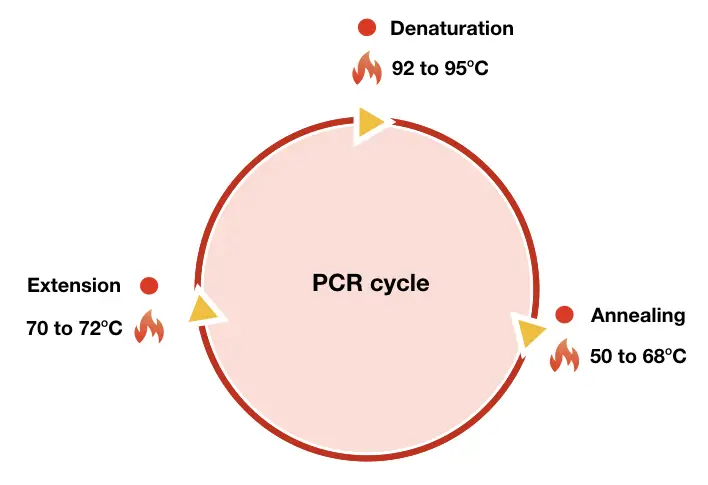
It’s a simple three-step, temperature-dependent enzymatic reaction including denaturation, annealing and extension. Each step performs a different function and synthesizes DNA using the Taq DNA polymerase.
To know more, you can read this article: Polymerase chain reaction.
Multiplex, PCR, RT- PCR, reverse transcription PCR, nested-PCR, ARMS-PCR are a few types quite known, however, researchers use techniques like asymmetric PCR in order to achieve some specific goals. The table below explains different types of PCR in brief you can click the link and read more.
| PCR types | Application |
| Gradient PCR | To optimize the reaction using different temperature zone. |
| rt-PCR | To real-time monitoring and quantifying the template. |
| Reverse transcriptase PCR | To study gene expression. |
| Nested PCR | To amplify the gene of interest using four sets of primers. |
| Inverse PCR | To amplify the region other than the target. |
| Allele-specific PCR | To amplify a specific allele in the PCR |
| Multiplex PCR | To amplify multiple templates in a single reaction |
| Hot star PCR | To increase the fidelity of the PCR |
| TB PCR | To quantity and amplify tuberculosis. |
If you are a regular reader of our blog, you may know that we are running the PCR series and exploring PCR optimization. In the present article, I will explain to you the basic concept of asymmetric PCR, applications and protocol.
I hope this article may add more value to your PCR knowledge and helps you in your research work.
Stay tuned,
Key Topics:
What is Asymmetric PCR?
The Asymmetric PCR is used to amplify a single strand of the template for various downstream applications.
Principle of Asymmetric PCR:
The use of an excessive amount of single-strand (target single-strand) specific primer amplifies only a single template ssDNA for DNA sequencing and hybridization probing.
It uses a 1:10 volume of primers in which a 10 times higher volume of target-specific primer is used. The use of the Linear-after-the-exponential process helps in deactivating the limited primer by higher annealing temperature to stop the amplification of dsDNA.
To optimize the asymmetric PCR four different parameters taken into considerations are the annealing temperature, the concentration of the primers, the concentration of template and the number of PCR cycles.
Explanation of Asymmetric PCR:
The asymmetric PCR has to face many problems during the reaction. Primer concentration has an important role here, the use of an excessive amount of a single primer creates various problems.
Modifying the melting temperature can overcome the problem. When using a
higher melting temperature for the limiting primer restrictions the dsDNA amplification. Secondly, once the limiting primers are used up, the reaction slows down, optimizing the PCR cycling condition defeats the problem. More PCR cycles are usually required here.
One of the biggest problems using asymmetric PCR is that it will give a very low amount of final ssDNA product. To optimize results, one has to use more starting materials and more PCR cycles.
The efficiency and yield of the asymmetric PCR are lower than the symmetric PCR. The LATE-PCR can increase the signal strength up to 80% when used for asymmetric PCR. It’s a Linear-After-The-Exponential PCR setup in which a limiting primer has been processed at a higher annealing temperature. Resultantly, it inactivates early during the reaction.
To get our final product as an ssDNA, DNA purification or targeted enzymatic degradation of the dsDNA is required.
J Aquiles Sanchez and E Kenneth et al has optimized the asymmetric PCR with the LATE-PCR technique for detecting the delta 508 and TSD 1278 alleles for the cystic fibrosis and Tay-Sachs disease from a single heterozygous cell.
Their results show an increment of the specificity and signal strength up to 250%
Paul Rutland from the Unversity College London gave a few optimization tips in his ResearchGate conversation and said to use 0.04 to 0.00004 micromol dilution of the limiting primer when using 0.05 to 1 micromol target or forward primer.
He also said that the technique needs more optimization and PCR rounds to reduce the dsDNA amplification and more PCR cycles to get a high yield of ssDNA.
Protocol for asymmetric PCR:
Make the 25µl PCR reaction for the first round of amplification as followed with fixed primer concentration. 0.5µl of 10mM dNTPs, 2.5µl of PCR buffer (10X), 1 µl of MgCl2 (25mM), 0.75 U of Taq DNA polymerase and 1µl of each primer.
Take some amplicons from the first PCR reaction.
If you don’t know how to prepare a PCR reaction, you can read this article, I explained everything stepwise: 10 secrets to prepare a PCR reaction.
The second round of 25 µl reaction consists of 0.5µl of 10mM dNTPs, 2.5µl of PCR buffer (10X), 1 µl of MgCl2 (25mM), 0.75 U of Taq DNA polymerase. Note that reaction needs various concentrations ratio of primers to optimize.
A couple of optimization goals are,
- Use 1:10 concentration of different primer sets with a fixed annealing temperature.
- Use different higher annealing temperature gradients for the limiting primers, make fix all other criteria.
- Use different numbers of PCR cycles to get more ssDNA amplicons, make fix all other criteria fixed.
Results of Asymmetric PCR:
The present technique can be performed for real-time PCR and for conventional PCR, so one can get results as either DNA bands or an amplification graph.
On a gel, results on the asymmetric PCR show two bands, one of the dsDNA and another of ssDNA which is our target. Real-time PCR shows a single prominent amplification graph of the ssDNA.
However, the dsDNA band or amplification signals should be weaker, if excessive amplification is obtained, do more optimization, increase PCR cycles and restrict more the limiting primer early.
Applications:
- The ssDNA has great value for DNA sequencing and hybridization studies.
- It is used in site-directed mutagenesis studies.
- It is also applied in allele discrimination and identifying various alleles.
Advantages:
- It facilitates single-strand amplification with a high precision rate.
- It works at a higher annealing temperature.
Disadvantages:
The process,
- Requires more PCR cycles and time.
- Time-consuming and tedious process.
- Less accurate and specific.
- Need additional optimization steps and procedures.
- More prone to contamination (with primary dsDNA).
- The yield of the experiment is too low.
Wrapping up:
To optimize and perform the asymmetric PCR, experience and expertise are needed. However, the technique has exclusive applications in some forensic analysis, diagnosis and genetic analysis.
We have written many articles on PCR, PCR types, PCR ingredients and PCR optimization, you can read it, here, by searching here.
Sources:
J Aquiles Sanchez and E Kenneth et al., Linear-After-The- Exponential (LATE)- PCR: An advanced method of asymmetric PCR and its uses in quantitative real-time PCR. Proceedings of the National Academy of Science Feb 2004, 101 (7).
Paul Ruthlands ResearchGate Thread.



