“Bacterial transposons are DNA transposons and help to spread antibiotic resistance. In this article, learn about bacterial transposons.”
Transposons are mobile genetic elements that can move within the genome of an organism. By doing so, it can spread into different locations. Both prokaryotes and eukaryotes carry them.
However, TEs are active in prokaryotes!
Tn family and IS elements are common transposons found in bacteria. Among these, IS, Tn5, Tn7, Tn10, Tn3 and Mu phage are some of the common and well-studied bacterial transposons.
Bacterial transposons carry a transposase gene that helps spread an antibiotic resistance gene. In this article, we will learn about bacterial transposable elements and their unique properties.
Stay tuned.
Disclaimer: The content presented herein has been compiled from reputable, peer-reviewed sources and is presented in an easy-to-understand manner for better comprehension. A comprehensive list of sources is provided after the article for reference.
| Type of transposons | Full name |
| LTR | Long Terminal Repeats |
| UTR | Untranslated Region |
| TE | Transposable elements |
| TSD | Target site duplication |
| TIR | Terminal Inverted Repeats |
| np | Nucleotide pairs |
| IS elements | Insertion sequence elements |
Key Topics:
Bacterial Transposons
Let’s first understand the basics of bacterial genetics.
Bacteria, in general, contain a circular bacterial chromosome (as a genome) and a few plasmid DNA in the cytoplasm. Although, circular chromosome is the main genetic constituent, plasmids also carry some crucial genes.
For instance, an antibiotic resistance gene. That’s present on a plasmid and can confirm resistance against antibiotics. Transposons, in bacteria, are present in the circular nuclear chromosome.
Interestingly, it can move between the nuclear chromosome and the plasmid. Tn family transposons- Tn3, Tn5, Tn7, Tn10, IS elements and Mu phages are commonly present in bacteria.
Bacterial transposons can be broadly divided into IS element transposons, Tn family and composite, and non-composite transposons. Let’s understand each, one by one.
IS elements
IS elements are Insertional Sequence elements. IS elements have been predominantly present in prokaryotes. In bacteria, it’s found in the circular genome and plasmids.
O.3% of the total bacterial genomic constituents carry the IS elements.
IS elements not only help in transposition but also regulate or control the process of transposition. That means it contains genes for transposase and resolvase.
Note that resolvase helps control the overall rate of transposition in bacteria. IS elements are compactly arranged, 1000 to 1500 nucleotides long TEs. Structurally, it contains three different parts– a transposition gene, ITRs and TSD.
A transposition gene is present in between the ITRs and is ~1500 bp long.
ITRs are inverted terminal repeats and present on both gene sides. Inverted Terminal Repeats (ITR) or Terminal Inverted Repeats (TIR) both are some. The length is between 8 to 40 nucleotides.
Noteworthy, ITRs are identical or nearly identical terminal repeat sequences. All the IS elements in all prokaryotes contain Inverted Terminal Repeats.
The last segment of the IS element is the Target Site Duplications (TSD) present after the ITRs. Upon insertion on the target location the IS elements copy (replicate) the target side and duplicate it and generate target site duplications on both ends.
Target-site duplications are direct repeat sequences. The whole process is explained in the figure. We already have written an article on this, you can click the link and read the whole process of TSD there.

Let me give you a brief overview of the duplication process for this article.
On the target location or DNA, the transposition produces staggered breaks on both sides. The TE is inserted in the target location. The gap then, filled with the replication process.
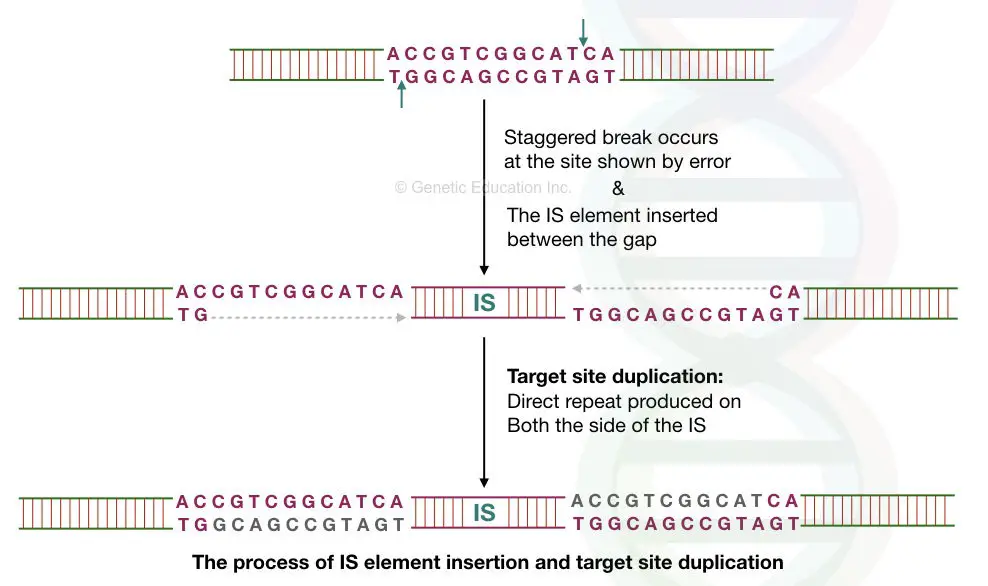
This produces direct repeats on both sides of the transposable elements. To understand this, carefully observe the above image.
Now, coming back to the IS elements.
The insertion event is led by the gene present in the middle of the IS element. Interestingly, the IS elements are categorized into different categories. And the number in each category can vary in the genome.
For instance, E.coli IS elements are classified into IS-1, IS-2, IS-3, IS-4 and IS-5. The example given below explains the variations in the IS element copy number in bacteria.
| IS elements | Size in (bp) | No. of copies/ genome |
| IS1 | 768 | 8 |
| IS2 | 1327 | 5 |
| IS3 | 1300 | 1 |
| IS4 | 1426 | 1 |
Composite Transposons
As the name suggests, it’s made up of the combination of both different transposons. For instance, when two different IS elements fuse together, it creates composite transposons.
Examples of composite transposons are Tn5, Tn9 and Tn10.
Composite transposons are constituted when two similar types of IS elements are fused. This fusion flanks a particular gene(s). Those are single or more genes that encode specific proteins.
Studies depict that these are mostly antibiotic resistance genes, carried by the composite transposons. Note that it follows a conservative mode of transposition.
For instance, the Tn5 contains two flanking IS elements on the left and right sides, named IS50L and IS50R, respectively. In the middle, an antibiotic-resistance gene is present.
Notedly, the flanked IS elements are the same here but not identical. That’s one of the unique properties here.
Moving further with the example.
The IS50L lacks the transposase gene while the IS50R contains the transposase gene. That means, the IS50R can move within the genome. However, the transpositive activity isn’t random, it is controlled by the protein named resolvase.
Resolvase is another protein that is encoded by the transposons that reduce the production of the transposase gene and control the transposition activities. Notedly, this is another unique property of composite transposons.
Much like the other transposons, composite TEs also contain terminal repeat sequences, as it is flanked by the IS elements. Remember, IS elements already contain terminal repeats!
Two other examples of composite transposons are Tn9 and Tn10. In conclusion, Tn5, Tn9 and Tn10 carry kanamycin, chloramphenicol and tetracycline resistance genes, respectively.
The structure of Tn9 and Tn10 composite transposons are given below.
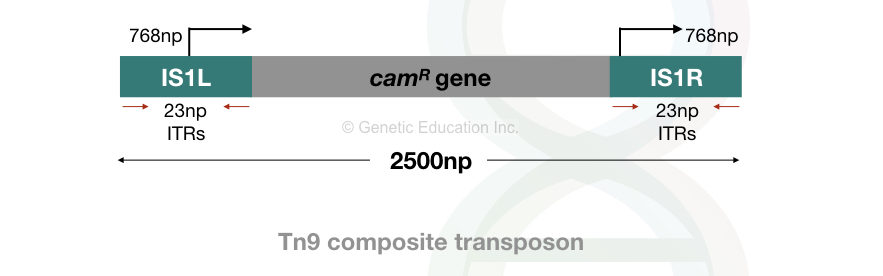


Non-composite Transposons
Like the composite and normal transposons, the non-composite transposons also contain transposons-specific elements (a coding gene, terminal repeats) but lack IS elements on both sides.
So as it is not made up of the composition of two IS elements, it’s referred to as non-composite bacterial transposons. For instance, Tn3.

Note that, these transposons also carry genes for transposons and resolvase proteins. Thus, the transposition activities are highly regulated here as well.
Tn3 elements are ~5000 bp long. On both sides, 30-40 nt long inverted terminal repeats are present. It contains tnpA, tnpB and bla genes and encodes transposase, beta-lactamase and resolvase proteins, respectively.
Note that beta-lactamase provides resistance against the antibiotic ampicillin. Lastly, the rest of the two genes help in transposition and regulation.
Composite vs non-composite transposons
| Feature | Composite | Non-composite |
| Definition | Two IS elements flanking a central region with genes. | Inverted repeats at both ends with no IS elements. |
| Structure | Contains two IS elements and a central coding region. | Contains a central region flanked by inverted repeats. |
| Examples | Tn5, Tn9, Tn10 | Tn3, Tn7 |
| Mechanism | Non-replicative (Cut-and-paste) and replicative using IS elements. | Non-replicative (Cut-and-paste) and replicative using IS elements. |
| Antibiotic resistance | Commonly carries antibiotic resistance genes. | Also carries antibiotic resistance genes. |
| Mobility | Moves between plasmid and chromosomal DNA. | Integrates into various sites in a genome. |
| Insertion | Target site duplication | Target site duplication |
Mu Phage
Mutator phage is short for Mu phage. These TEs are also known as transposable phages. It can jump and integrate into the bacterial genome randomly. Interestingly, it is a type of replicative transposon.
That means, it jumps to another location and leaves a copy behind in the previous position. The Mu phage is larger than IS elements with the size of ~36Kb. However, it can have some similarities with the IS elements.
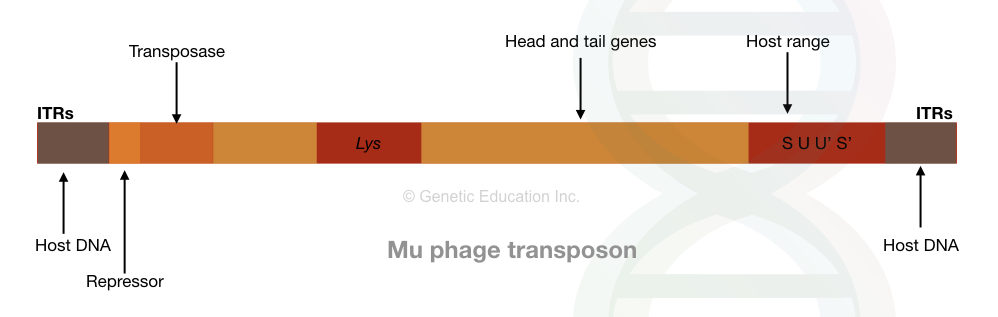
Structure, much like the other transposons, Mu phage also contains terminal inverted repeats and target site duplication sequences from the previous location.
It also carries genes for replication, coat protein and lysis. Noteworthy, two unique properties of the Mu phage that make them different from the IS elements are,
The target site duplication can not be transposed to a new location and it’s not necessarily located on the terminal ends.
Another unique property of the Mu phage is that two different Mu phages can combine together and carry a gene in between for transposition. Bacterial F-factors, markers and antibiotic resistance genes can be transposed between plasmid and genome by Mu phage.
Summary:
| Feature | IS elements | Composite TE | Non-composite TE | Mu Phage |
| Definition | Simple mobile genetic elements with inverted repeats. | Two IS elements flanking a central region with genes. | Transposons with inverted repeats, no IS elements. | Transposable phage integrating randomly into genomes. |
| Structure | Contains only a transposase gene with terminal repeats. | Contains IS elements on both ends and a central coding region. | Central region with inverted repeats on both ends. | Larger phage with inverted repeats and replication genes. |
| Mechanism | Replicative tranposition. | Non-replicative (Cut-and-paste) and replicative using IS elements. | Non-replicative (Cut-and-paste) and replicative using IS elements. | Integrates into the host genome by “copy-and-paste” method. |
| Antibiotic resistance | Does not typically carry antibiotic resistance genes. | Carries antibiotic resistance genes. | Carries antibiotic resistance genes. | Can carry resistance genes between bacterial genomes. |
| Function | Facilitates genetic recombination and mutations. | Transfers genes like antibiotic resistance between DNA. | Transfers genes like antibiotic resistance within genomes. | Transfers DNA and phage genes to host cells. |
| Example | IS1, IS2, IS3 | Tn5, Tn9, Tn10 | Tn3 | Mu phage |
Wrapping up:
In conclusion, transposons in bacteria help in spreading antibiotic resistance by transposing the antibiotic resistance gene. We will discuss this in our next article. Unlike the eukaryotic transposons, bacterial transposons are active.
These jumping genomic elements provide the necessary diversity to the bacterial genome and plasmid to survive against phages and antibiotics.
I hope you like this article. Do share it and subscribe to our blog for more genetic content.
Sources:
Babakhani S, Oloomi M. Transposons: the agents of antibiotic resistance in bacteria. J Basic Microbiol. 2018 Nov;58(11):905-917. doi: 10.1002/jobm.201800204. Epub 2018 Aug 16. PMID: 30113080.
Ooka T, Ogura Y, Asadulghani M, Ohnishi M, Nakayama K, Terajima J, Watanabe H, Hayashi T. Inference of the impact of insertion sequence (IS) elements on bacterial genome diversification through analysis of small-size structural polymorphisms in Escherichia coli O157 genomes. Genome Res. 2009 Oct;19(10):1809-16. doi: 10.1101/gr.089615.108. Epub 2009 Jun 29. PMID: 19564451; PMCID: PMC2765283.