“A typical PCR reaction contains a required concentration of dNTPs, Taq DNA polymerase, PCR primers, PCR buffer, template DNA and water. Each PCR reaction varies in terms of the concentration of ingredients. This guide helps you to prepare an effective reaction.”
PCR, short for Polymerase Chain Reaction, is an amazing and crucial discovery in genetics. No technique can fully replace PCR. The reason is, it solves the basic problem of genetics, the requirement of copies of the desired fragment.
It can produce millions of copies of a particular fragment. We can study a particular gene, fragment or DNA sequence, however, it was a tedious process in the past. Many excellent variants of PCR exist nowadays. But the success of any PCR solely depends on the reaction preparation.
PCR is a highly sensitive technique so reaction preparation is a crucial and important factor. Any minor fault fails the amplification. So you may wonder how to prepare an amazing and effective reaction for a PCR. Let’s find out.
Hey there, I’m Dr Tushar and an expert in molecular cytogenetics and techniques like PCR, sequencing and microarray. In this blog article, I will give you a step-by-step guide for preparing an effective PCR reaction.
Trust me, after applying this guide in your PCR lab, you can surely get positive results.
Stay tuned.
Key Topics:
How to prepare a PCR reaction?
I am hoping that you know the basics of PCR and I am directly stating the guide without wasting time.

Step 1: Prepare a template DNA:
The very first step in this guide is to prepare our PCR template DNA after extraction. Check the quality of DNA. It should be between 1.7 to 1.8 at 260/280 nm wavelength. If required perform alcohol purification to remove contaminants.
Quantify the DNA. We need 30 to 50 ng DNA for a 25 µL reaction. Store DNA at 4°C before use.
| Purity of DNA | 1.7 to 1.8 |
| Quantity of DNA | 30 to 50 µL |
Step 2: Prepare (or revive) PCR primers:
The concentration of PCR primers is indeed a critical factor to achieve excellent amplification. At a higher concentration, you will get more non-specific bands and primer-dimer while at a lower concentration the chances of amplification decrease.
The ideal concentration for primer is 10 pM. Our stock comes in 120, 100 or 115 pM, etc. You have to prepare the working solution of the primer by dilution. Suppose you have 100 pM stock,
Pipette 10 µL from stock and add 90 µL of nuclease-free water or TE buffer, mix it well and store it at -20°C temperature. Our 10 pM working is ready.
Primer checklist:
| Length | 18 to 22 nt |
| Annealing temperature | 50 to 65°C |
| GC content | 50 to 55% |
| Hairpin formation tendency | Less |
Note: 10 pM primer concentration is sufficient to amplify any DNA but I recommend optimizing the concentration as per the requirement of the assay.
These are the two requirements for PCR pre-preparation before preparing an actual reaction. Now moving to step 3.
Step 3: Prepare for the reaction:
- Wear gloves, a mouth cap and a lab coat.
- Sterilize the area or reaction preparation hood.
- Take all the reagents and thaw them gently.
- Followed by thawing, place all the ingredients on ice.
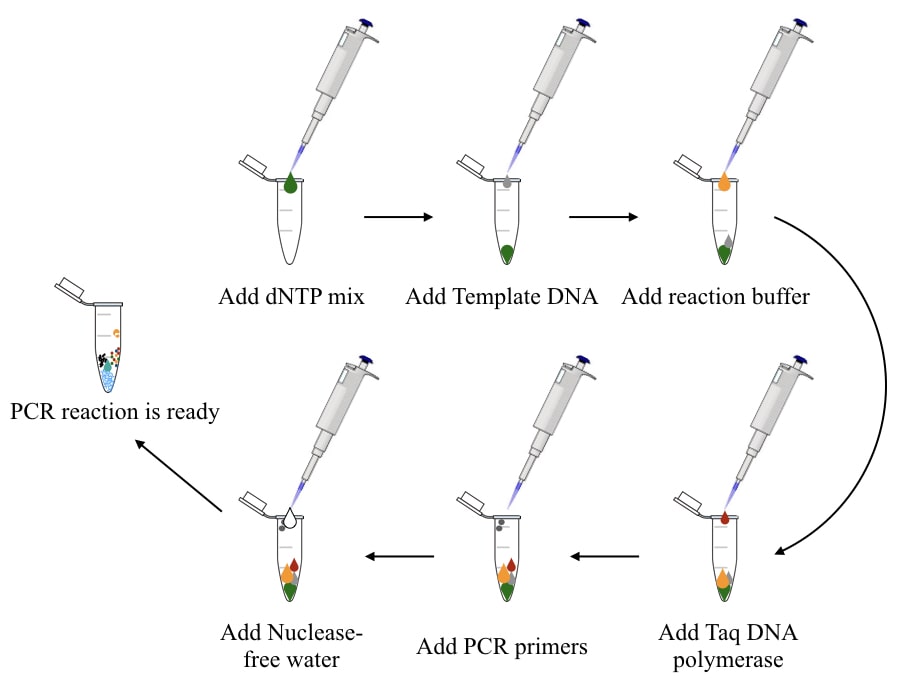
Step 4: Add dNTP mix:
Add each ingredient in the exact order in which I am enlisting.
dNTPs are free nucleotides that are added to the growing DNA template by the Taq DNA polymerase.
Add each dNTP or dNPT mix to the bottom of the tube. Add 50 µM each or 200 µM dNTP mix in the tube. Remember, we are preparing a 25 µL reaction.
Step 5: Add template DNA:
Template DNA provides a single-stranded substrate for the Taq DNA polymerase to perform the reaction. We already have explained the template preparation.
Add 30 to 50 ng template DNA to the tube.
Step 6: Add reaction buffer:
The reaction buffer contains PCR enhancers like MgCl2, KCl or albumin which increases the efficiency of the reaction. Note that if required, the MgCl2 can be added separately. To know more on the concentration and role of MgCl2 in PCR, read this article: What is the role of MgCl2 in PCR?
Add PCR reaction buffer as per the requirement. Usually 1X concentration is needed.
Step 7: Add Taq DNA polymerase:
Taq DNA polymerase is an enzyme that catalyzes the amplification reaction. It uses the cofactor, settles on the substrate and adds nucleotides to the growing DNA strands.
Add Taq DNA polymerase 1 to 1.5 U to the reaction.
Step 8: Add PCR primers:
Taq DNA polymerase lacks de novo synthesis property and thus requires a short DNA primer to initiate the synthesis. It uses the primer-DNA duplex junction as substrate to amplify the DNA.
Add 10 pM each primer to the reaction or put drops on the wall of the tube.
Step 9: Adjust the final volume:
To level up the reaction up to 25 µL, add the required amount of highly pure and nuclease-free water.
Prepare a final volume of 25 µL using nuclease-free water.
Step 10: Mix the reaction:
Mixing by gentle spinning or tapping the tube. It will mix all the ingredients equally in the reaction.
Spin the reaction gently and put the reaction tubes in the pre-set PCR protocol.
Note: perform all the steps on ice only.
25 µL PCR reaction preparation:
| PCR ingredient | Concentration | Volume |
| dNTPs mix | 200 µM (50 µM each) | 4 µL |
| Primer 1 | 10 pM | 1 µL |
| Primer 2 | 10 pM | 1 µL |
| Taq DNA polymerase | 1U | 1.5 µL |
| PCR reaction buffer | 1X | 10.5 µL |
| Template DNA | 30 to 50 ng | 3 µL |
| Nuclease-free water | – | 4 µL |
| Final volume | 25 µL |
Wrapping up:
This step-by-step guide hopefully solves your problem to prepare the reaction for PCR. However, for atypical reactions like long-range PCR and high GC-rich templates, I recommend taking expert’s advice.
Also separate PCR setup and reaction ingredients are required to make such reactions effectively. I have covered an article on this topic too. You can read it here: What is a Long range-PCR?
Related articles: