“Gene-editing is a technique to manipulate an organism’s genome or gene or DNA sequence which might be insertion or deletion. CRISPR-CAS9 system is one of the popular gene-editing techniques so far.”
Simply put, we can describe the gene-editing as editing the messages of the DNA. In gene editing, we are changing the sequences thereby changing the message driven by that particular DNA sequence.
Gene or genome-editing is a technique of genetic engineering used for treating genetic diseases and to developing economically important organisms. CRISPR- the DNA sequence, CAS9- a kind of nuclease and the special type of RNA are the key elements of the present system.
In this article, we will discuss gene editing or genome editing and related tools. The major focus of the article is
- What is gene editing?
- Definition
- History of gene-editing
- Need for gene-editing
- How gene editing is done?
- CRISPR-CAS9 gene editing system
- CRISPR-CAS9 in Bacteria
- CRISPR-CAS9 in genetic engineering
- Applications of CRISPR-CAS9
- Limitation of CRISPR-CAS9
- Ethical issues in gene transfer technique
Key Topics:
What is gene/genome editing?
Definition:
“ Manipulating the genetic composition of an organism artificially is called a gene-editing”.
History of Gene editing:
Some prefer to say it as genome editing but both the terms can be used, we can use any of it. The main point of editing DNA is to remove the abnormal or faulty DNA sequence.
The story for gene editing came to light in the late ’70s when Herb Boyer and Stanley Cohen discovered antibiotic-resistant genetically engineered bacteria. Until 2012, the idea for gene editing was just a fairy tale.
In 2012, the CRISPR gene editing system was discovered which changed the era of genetics. A group of scientists from the University of California discovered a so-called bacterial immune system that can be edit genes at the targeted location and named it a CRISPR-CAS9 system.
However, the idea was evolved in the year 1982 when human insulin-producing bacteria were discovered. This synthetic insulin becomes so successful that it was commercially available after FDA approval.
After the successful implementation of the synthetic insulin, A roadmap was created for the genetically modified Flvr shower tomato, bringer and some of the species of cotton. This is the short history and the milestone in gene-editing technology.
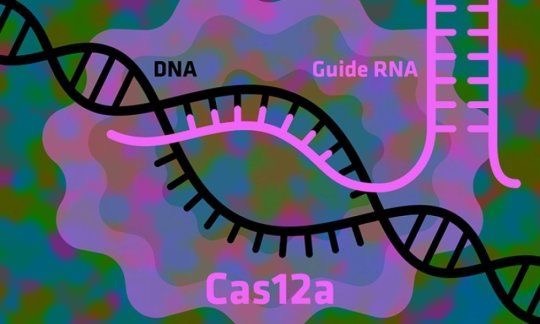
Editing genes of bacteria, yeast and mice are nowadays quite easy, however, not possible for the human genome. Actually, it is not impossible but there are some ethical issues associated with human embryo testing and research. We will discuss ethical issues in the last segment of this article.
The process of gene editing is divided into the following steps:
- Identification of target site into the genome of an organism.
- Designing a healthy copy of that particular targeted site.
- Designing the guided RNA.
- An enzymatic reaction to cut the faulty DNA.
- Insertion of new DNA
- Repairing the site of action.
The targeted site into the genome or in a gene is a place that is faulty or mutated and produces faulty/mutated protein. The specially designed “Guided molecules” (which are nucleic acids) binds to the specific target site into the genome with the help of the enzyme.
The enzyme is a nuclease, a bacterial nuclease also called a molecular scissor or engineered nuclease which helps in cutting the DNA, After binding of a guided molecule to the target site.
Once the guided molecule binds to the site of action the nuclease cuts that portion of the DNA from the genome and removes it. A nick is created at this place is later filled by the cell’s natural DNA repair mechanism.
The cell’s natural DNA repair mechanism finds any change or break into the genome and fill it with the complementary bases during the process of the replication. We can also insert the piece of DNA of our interest on that particular site as well.
Here the guided molecule is an RNA know as gRNA or sgRNA (single-stranded RNA). We have written an article on structure, function and designing of gRNA. Read it here: What is sgRNA? – Definition, Designing and Advantages.
By means of nuclease governed gene editing, we can delete or insert the DNA sequence of our interest. This is the simple explanation of gene editing. The pictorial representation of gene editing is shown in the figure below.
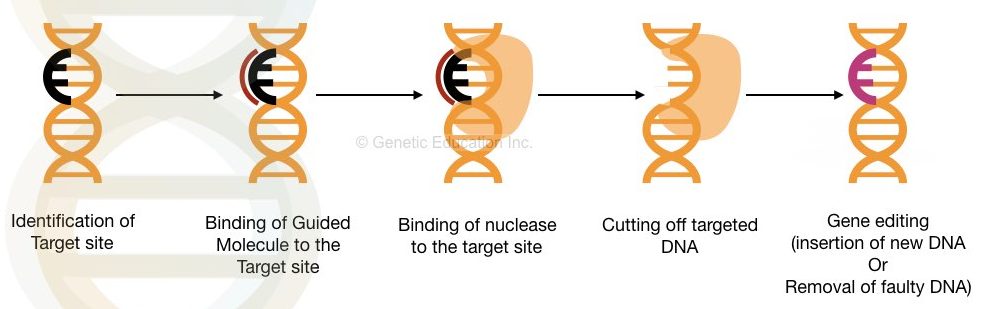
Glossary used into the article:
| CRISPR | Clustered Regularly Interspaced Short Palindromic Repeats |
| CAS9 | CRISPR associated protein 9 |
| gRNA | Guided RNA |
| crRNA | CRISPR RNA |
| DSB | Double-stranded break |
| ZFN | Zinc finger nuclease |
| TALEN | Transcriptional Activator-Like Effector Nuclease |
| NHEJ | Non-homologous end-joining |
| HDR | Homology direct repair |
| PAM | Photo-spacer adjacent motif |
What is a need for the gene-editing?
Genetic disorders are lethal and nearly incurable. After the discovery of DNA in 1953, the picture of the inheritance of disease becomes more clear. The chromosome carries the DNA that is inherited from one generation to another.
In this manner, the genetic material passes from parents to their offspring and so the mutations are! Any changes that occur in a genome can be pass-down to offspring. And a genetic abnormality happens. Gene editing can help to eliminate this mutated portion of the genome.
Importance of gene editing:
- It is used to edit the genome of any organism however, it is most applicable in the bacteria, yeast, mice and other model organisms.
- It is used to study the gene expression of organisms.
- The technique can remove mutated genes from the genome of an organism, also, it can be used to create new variations.
- One can change the characteristics of the organism.
- Economically important genetically modified organisms can be created too.
Types of gene-editing:
The molecular scissor, a genetically modified nuclease enzyme plays an important role in the gene-editing technique. Therefore, depending on the types of nuclease there are several techniques used for long in genetic engineering. Three major techniques are enlisted here:
- ZFNs (Zinc finger nuclease)
- TALEN (Transcriptional activator-like Effector based Nuclease)
- CRISPR-CAS9 (clustered regularly interspaced short palindromic repeats)
ZFNs and TALENs:
The difference between the ZFNs and TALENs is only because of the use of different molecules.
The ZFN technique was introduced in early 1991 by Palvetich and Pabo. The Zinc finger proteins are artificially engineered protein that binds to the specific location at the target site. The ZNFs bind to the 3 to 4 nucleotides away from the cleaving site.
Two copies of the ZNFs are required for gene editing. Once it binds to the target sequence, the bacterial nuclease Fok1 cuts the DNA sequences. Fok1 is the bacterial nuclease encoded by the DNA binding domains.
Related articles: what is gene therapy?
Here the Fokl recognizes both the end of the cleaved DNA. This can help to cleave, remove or insert DNA at that location.
The TALEN works similarly to ZFNs However, it can cut the larger sequence efficiently than ZNFs. In TALEN the Fok1 cuts the double-stranded DNA instead of single-stranded one.
The graphical representation of both the techniques are given in the figure below,
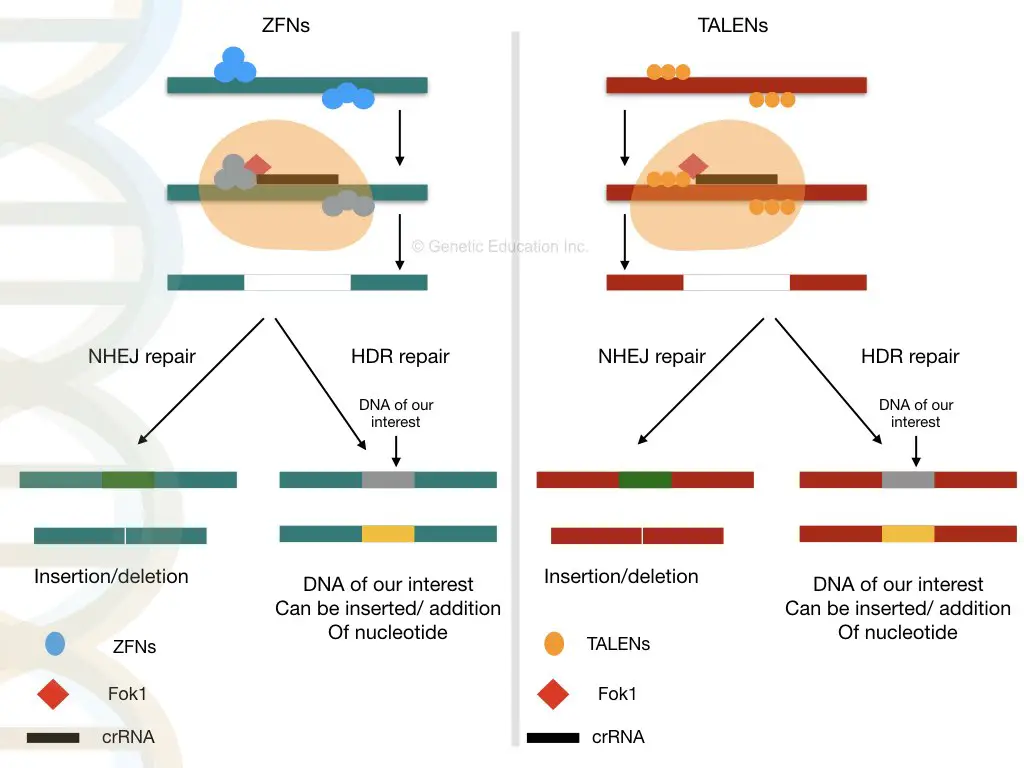
Zinc finger nuclease, Transcriptional activator-like effector-based nuclease and meganuclease are the traditional techniques that are not applicable in the recent day due to several limitations.
- These techniques are time-consuming and costly. More time and cost are required to engineer artificial proteins such as TALEN and ZFNs.
- The specificity of gene editing is also poor.
- The end result, the rate of gene edition is also low.
- Though the technique is sensitive enough and accuracy is less.
In the year 2012, a new method of gene editing was demonstrated by the group of scientists from the University of California named it CRISPR-CAS9. CRISPR-CAS9 is the most advanced and futuristic technique. It is fast, reliable, cost-effective, efficient and accurate.
CRISPR-CAS9 gene editing system:
CRISPR- Clustered Regularly Interspaced Short Palindromic Repeats are present in the bacteria and some archaea. It is the element of the bacterial natural defense mechanism. The CRISPR-derived RNA helps the bacteria to protect it from invading viruses and phages.
The nuclease, CAS9 protein cuts the DNA strand of the foreign pathogens and destroys them.
The CRISPR gene sequences were discovered in the year 1980 in E.Coli but their function was not known at that time. Later on, in the year 2007, Barrangou et al. reported their role in the adaptive immune system of the bacteria.
CRISPR-CAS9 in Bacteria:
During the viral infection, the virus injects its DNA into the Bacteria through the bacterial cell wall. The phage DNA is integrated into the genome of the host and transcribed and translated. The new viral protein forms from the bacterial genome.
To avoid phage infection, bacteria developed a unique defense mechanism called a CRISPR-CAS9 based defense mechanism.
During the viral invasion, the bacteria took some of the DNA sequences as a reference and inserted them into the palindromic repeats, that generate CRISPR. CRISPR is the combination of bacterial palindromic repeat sequences and the viral spacer DNA.
The spacer DNA is transcribed into the RNA known as crRNA which guides to binding at the specific location into the viral genome. The DNA-RNA heteroduplex is created at the site of the invasion.
Along with the crRNA, the bacteria also transcribed several short tracrRNA (trans-activating crRNA), partially complementary to the crRNA. The tracrRNA helps for the maturation of the crRNA from the multiple pre-crRNA.
The junction of crRNA and tracrRNA is called a gRNA or guided RNA which guides the nuclease for cutting the target sequence.
The CAS9 recognizes this gRNA and binds to the location of the heteroduplex. Once it is placed properly, two unrelated domains of a nuclease (CAS9 nuclease) are named as a RuvC- like domain and HNH domains lyses the phage dsDNA.
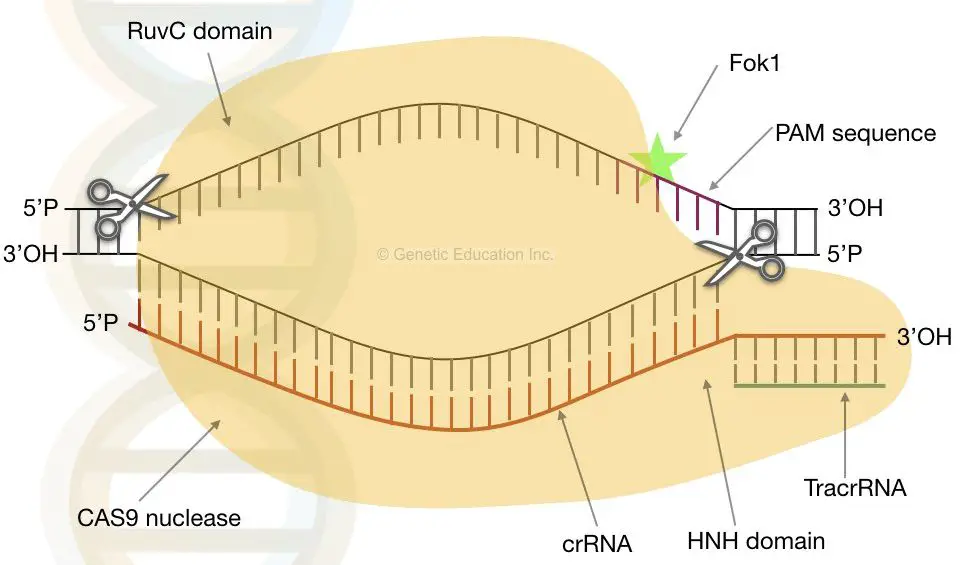
During this entire process, the CRISPR sequences work as a target for nuclease to cleave the DNA. The viral or phase DNA can’t replicate and destroy.
The RuvC domain cuts the non-homologous domain while the HNH cuts the homologous domain (homologous to gRNA).
The foreign pathogen or the virus is destroyed by the combined effort of CRISPR and CAS9 protein. One question arises after this explanation that, “does it cut its own DNA?” the answer is “Yes”.
The viral DNA sequence is present in the CRISPR as well. Therefore, it is might possible that bacteria CRISPR-CAS9 recognizes these spacer sequences as foreign DNA and edit them. Actually, it doesn’t happen, thanks to the motif called a PAM!
The photo spacer-associated motif (PAM) helps the CAS9 to recognize the gRNA. The nuclease locates near the PAM domain. It binds to the 3′ OH end of the gRNA.
This will give a signal to CAS9 for cleaving the DNA. CAS9 recognizes the PAM-gRNA complex on dsDNA and performs the catalytic reaction. Lack of PAM in bacteria’s own DNA protects them. It is a kind of built-in safety mechanism developed by the bacteria itself (How smart they are!).
PAM protects bacterial DNA from its own nuclease activity.
The CRISPR are the DNA sequences made up of the spacers and the repeats. The spacers are present between the short palindromic repeats throughout the CRISPR. Different spacer sequences are taken from different viruses that help the bacteria during the pathogenic attack in the future.
Bacteria use the information of these sequences to destroy pathogens. But how we can use it for our use?
We can design synthesis gRNA that leads the CAS9 to the target location we wish to study and that is how we can use it for doing different genome-editing experiments.
CRISPR-CAS9 in genetic engineering:
Guided RNA and the dimeric CAS9 protein are the only requirements in the gene-editing technique.
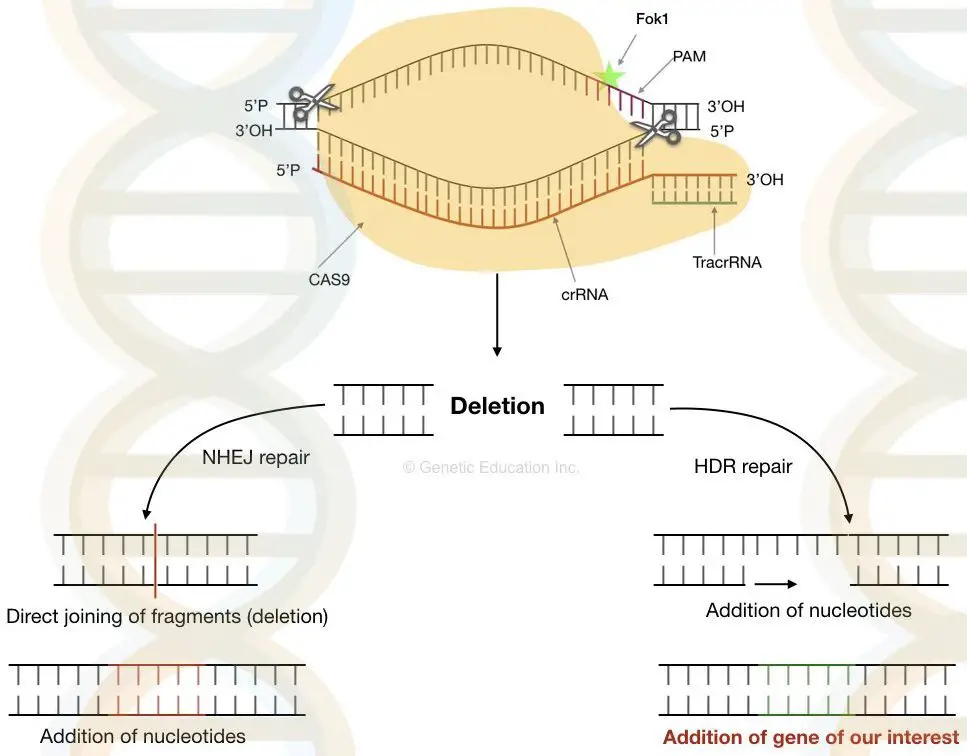
Once the nuclease cuts the DNA at the specific location, it creates the double-stranded cut into the DNA. Now, the natural DNA repair mechanism of the cell repairs this gap in one of the two ways either non-homologous end-joining or Homologous direct repair.
Through the process of double-stranded break repair, the whole mechanism works.
Non-homologous end-joining: in this type of DNA repair mechanism, the gap is directly filled by the joining of two opposite non-homologous strands. Here, in NHEJ several important pieces of genetic information might be lost because the entire fragment of DNA is lost.
This DNA fragment will never be present in the next cell division, will create a mutation into the genome. If some of the important gene for the organism is knocked out, the condition becomes lethal for it.
Homologous direct repair: the gaps are not filled directly in homologous repair. The Homologous direct repair follows the mechanism of recombination. Here the nucleotides are inserted at the site of the cut based on the information of the previous replication.
For doing this, the cell uses single-stranded DNA as a template for the addition of new nucleotides. Now, this is the time we can insert a single-stranded DNA of our interest which is synthesized naturally in consecutive cell division by the cell itself.
We can knock out the faulty DNA by non-homologous end joining while DNA can be inserted by the homologous direct repair.
In molecular genetics or in genetic engineering the CRISPR-CAS9 is used as a gene-editing tool. Based on the activity of the CAS9, three different gene-editing systems are postulated. All three types of systems are enlisted below:
- CAS9 with only cleavage activity
- CAS9 nuclease full activity
- CAS without the cleavage activity.
In the first system, the cas9D10A mutant is developed from the CAS9 encoding gene. This mutant can’t cleave both strands of the DNA. Instead, cas9D10A can recognize the gRNA and cleaves only a single strand of the target DNA.
What is the benefit of this mutant? Well, it increases the precision of gene editing. It allows only high-fidelity homologous direct repair of the single-stranded gap reduces the deletion-insertion mutation chance into the gnome.
In short, the cas9D10A variant does not allow the non-homologous end-joining.
The second system is the normal function of CRISPR-CAS9. None of the sequences is mutated for CAS9 proteins. We had already discussed the CRISPR-CAS9 system and how normal CAS9 works.
The third system depends on dCas9. The dCas9 is called nuclease deficient CAS9 or dead Cas. Some mutations are artificially introduced into the RuvC and HNH domains lead to suppressing the function of nuclease.
The nuclease deficient CAS9 can only bind to the target sequence but can not cut it. This facilitates gene silencing or activation of a particular gene with the help of many effector domains.
Read further: Cas9 Protein: Structure, Function, Types and Importance.
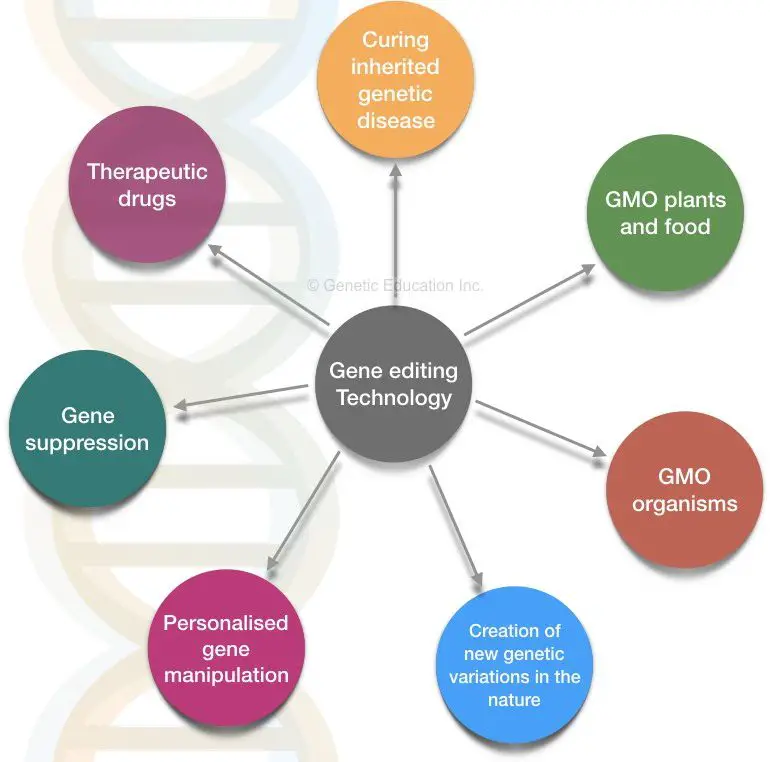
Application of CRISPR-CAS9:
The technique is reliable, cost-effective and fast.
The present system helps to create large deletions, rearrangements and inversion.
A point mutation can be induced with the help of single-stranded gRNA.
The gRNA can be easily synthesized and designed that is not possible in the case of ZFN and TALEN.
Gene silencing is achieved by the dCas9 system. Any mutated gene or gene of our interest can be suppressed by CRISPR-CAS9.
CRISPR-CAS9 is applicable in the creation of genetically modified organisms (also called genetically engineered organisms). Many different genetically engineered plant species can be developed through CRISPR-CAS9.
Furthermore, gene expression studies in mice, sea archaea, zebrafish and fungus are possible because of the CRISPR-CAS9. Knockout mice can be generated by CRISPR-CAS9 which helps in the study of the role of the particular gene.
The microscopic examination of specific gene loci is facilitated by CRISPR-CAS9.
Read more on the Applications of CRISPR-CAS9:
Do you know
We can visualize any gene with the help of CRISPR-CAS9?
The naked CAS9 (CAS9 without nuclease activity) can only bind to the specific gene loci. By introducing the mutation D10A into the RuvC domain and H840A into the HNH domain of CAS9 the activity of nuclease suppresses, however, it allows CAS9 binding to the gene of interest. The gene of interest does visualize microscopically.
EGFPs, Enhanced Green Fluorescent Protein fused with the dCAS9 was first used to visualize the repetitive sequences into the genome.
Limitations of CRISPR-CAS9:
CRISPR-CAS9 is the best tool in recent years. Nonetheless, only a single limitation makes it restrictive to use. The “off-target effect” introduces a mutation other than the target site. CRISPR-CAS9 cleaves off-target DNA too and reduces the efficiency of gene editing.
In some organisms such as zebrafish and sea archaea >70% efficiency of gene editing was reported (WY Hwang et al., 2013). In the contrary, only 3% to 4% efficiency was reported into human pluripotent cells (P Mali. et al., 2013).
Efficiency deviation in different cell types is the only limitation of the CRISPR-CAS9 tool. Still, we need a lot of optimization to use the present type of gene-editing system efficiently.
Ethical issues in gene editing?
Ethics is the moral principle of the individual. We have to take care of the moral principles of the organism while performing any scientific experiments on them. The prior concern is further required before performing any experiment on any organism.
Safety is the first ethical concern of any experiment. So the gene transfer technique must be safe enough to use for routine diagnostics.
It should be available for all. If it is too costly, only rich people can afford it which means it creates discrimination based on wealth. The gene transfer technique must be cheaper enough to be available for all.
Though the technique can be used in disease diagnosis but is not used for personal health care. One can use it for increasing athletic capabilities and beauty treatments. Therefore, the use of the technique is restricted for some meaningful purpose.
The gene-editing technique should not have any lethal side effects.
The risk level of the technique should be minimum otherwise it will adversely affect the organism as well as the environment.
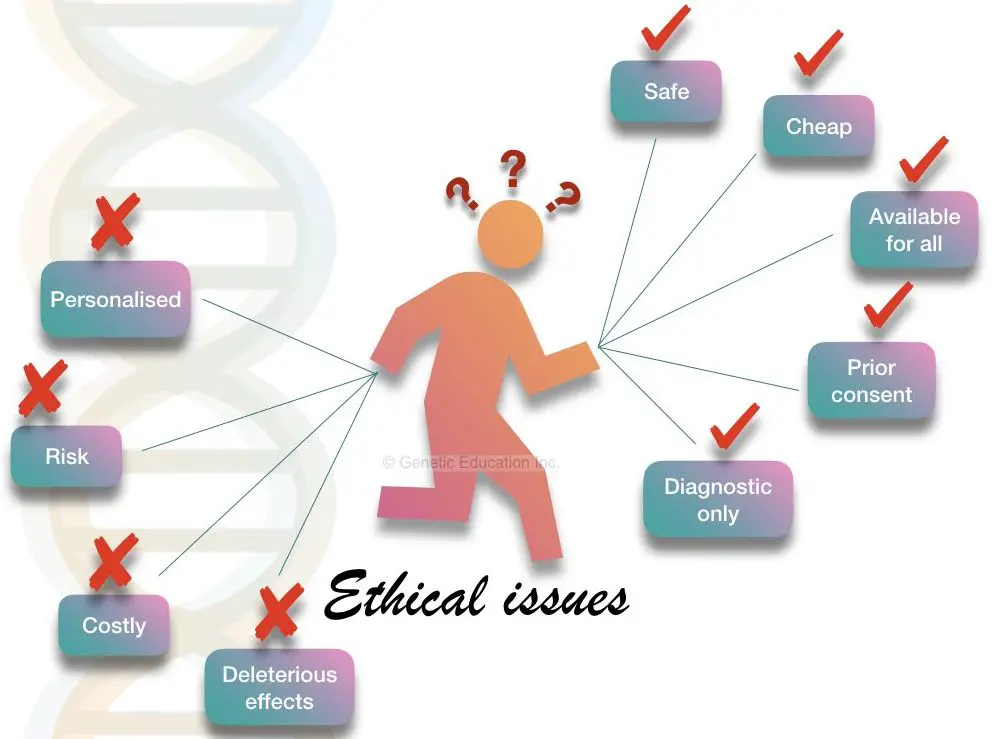
Gene editing should be avoided in the germline cells or into the developing embryo because it can create any new deleterious, lethal or harmful mutations in the population.
Creating, manipulating or destroying a human embryo is not a legal activity and prohibited by the USA, UK and other governments. After all, the social, religious or moral sentiments of any individual should not be affected.
The off-target effect is a big challenge in the CRISPR-CAS9 gene-editing technique. Many homologous DNA sequences are present in the genome of an organism. The gRNA can bind to any of this homologous DNA sequence.
This results in the induction of the unknown mutation into the genome. The deleterious effect of new mutation may be harmful to the cell or the organism.
Interesting articles to read:
- The Comet Assay to Detect DNA Damage.
- Paternity DNA Test- Process, Results, Legal Proceeding and Cost.
Conclusion:
Though the gene-editing technique is a revolutionary and emerging genetic technique, experiments on humans and other animals should be avoided. No doubt, CRISPR-CAS9 like gene editing tools will be available in medical science in the nearby future but manipulating the genetic composition of an organism can create serious problems for mankind.
We have to use gene manipulation wisely only to treat disease, not for personal use.