“It is believed that the heterochromatin is transcriptionally inactive sequences, and are further classified into constitutive heterochromatin and facultative heterochromatin.”
Highlight:
- Do you know some heterochromatin are still active and can form a protein? We will explain what later in this article.
- The heterochromatin comprises only 90% of the total genomic region, the rest of 10% are either euchromatin or mixed type of chromatins.
- Under the microscope, the heterochromatin regions appear as darkly stained, compact irregular patches near the nuclear membrane- periphery of the nucleus.
- It is mainly present in the telomeric and centromeric regions as satellite DNA and performs different functions.
You are on this article, which means you are in search of more information on heterochromatin, and probably you know the structure of the genome and DNA organization- how DNA is arranged on chromosomes.
Here is the walkthrough of the present topic in brief.
At different stages of the cell cycle, forms of DNA vary. It may appear as chromatin, chromosome, loosely packed chromosomes, and helices only! at different stages.
Chromatins are later on arranged on chromosomes by interacting with a special type of protein- histones.
What is the characteristic of transcriptionally inactive heterochromatin?
—–Methylation of histone 3 at lysine 9.
Chromatins are fragile, loose, irregularly arranged double helices DNA present in a cell nucleus, you can imagine a bowl full of noodles- bowl as a cell and noodles as chromatins.
Interestingly, we can’t arrange noodles in a bowl in specialized structures but a cell can! Cells allow DNA helices interaction with protein and manufacture euchromatin and heterochromatin regions- different in structure, function and arrangement.
At a molecular level, DNA interacts with histones like H1, H2A, H2B, H3 and H4 and forms a nucleosome. This, later on, manufactures bead-on-string appearing chromatin.
When the 30nm fiber of DNA is constructed from this complex, two distinct chromatids form chromosomes through centromeric attachment.
Euchromatin and heterochromatin regions are scattered on chromosomes having more heterochromatin on telomere and centromere regions. It can be further classified into two categories that we will discuss in the upcoming section.
Let me explain to you what heterochromatin is and its types in the present article. I will explain the function of both and their importance as well. In addition, we will also discuss the molecular structure of HC and other aspects as well.
Key Topics:
What is heterochromatin?
In 1928, Emil Heitz defined the concept of heterochromatin, abbreviated as HC.
Chromatin is the strand of dsDNA interacting with proteins to organize on chromosomes. Heterochromatin and euchromatin are two common types in which heterochromatin are transcriptionally inactive.
Around 90% of the human genome comprises this region in which 6.5% are constitutive while the rest are facultative.
Milestones:
1928- Emil Heits coined the term heterochromatin
1951- B McClintock discovered transposons
1974: Kornberg and Thomas stated the post-translational histone modification.
1975- Holliday and PUgh; Riggs explained gene inactivation by DNA methylation.
2001; 2006 and 2013- Nakayama et al., Boyer et al., and Li Zhou studied heterochromatin molecular profile using the ChIP technology.
Definition:
“A darkly stained, tightly wrapped, inactive region mainly located in the telomere and centromere region of the chromosome is known as heterochromatin region.”
Types of heterochromatin:
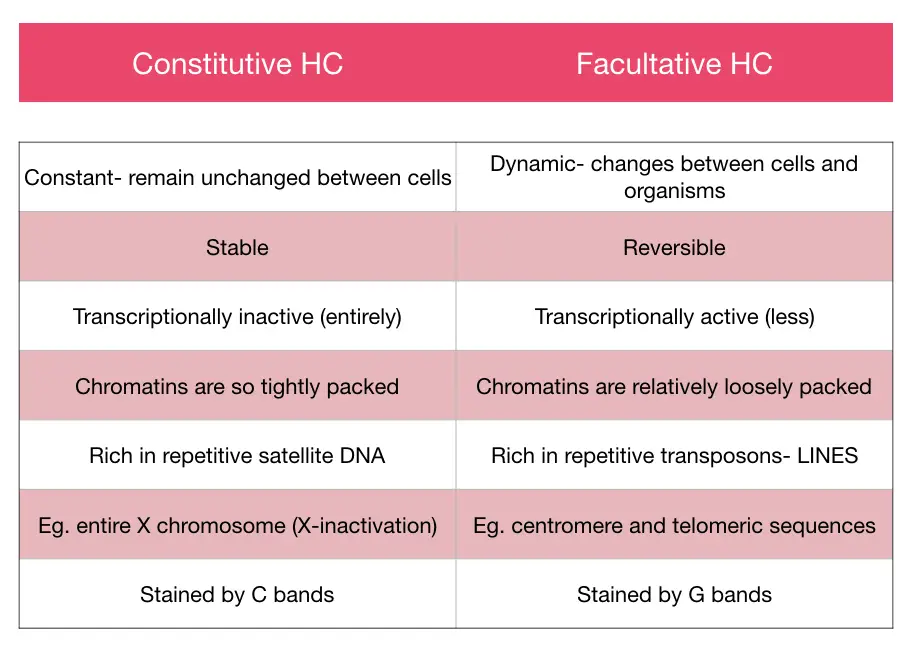
Previously scientists believe that the whole region is transcriptionally inactive, although it is not true. The entire heterochromatin region can be divided into two broad categories based on transcriptional activity.
One, which is inactive but can be activated or active in other cells, and another type is an entirely inactive junkyard of the genome. These two types are facultative and constitutive heterochromatin, respectively.
Facultative heterochromatin:
The facultative heterochromatin region is dynamic in nature, which means, if it is active in one cell type or organism, can be inactive in another. These regions though have the capacity to activate transcription, are considered as poorly expressed genes.
What does it mean!
It shows that in some cell type or organism it’s facultative while in others it is constitutive. The classic example of facultative heterochromatin is the inactivation of the X chromosome.
Let me explain to you in brief,
There are two X chromosomes in females, so one must be inactivated to regulate gene expression. Noteworthy, the process of X chromosome inactivation is random, sometimes maternal while the rest of the time the paternal X chromosome gets inactivated.
The X-chromosome is rich in facultative heterochromatin, having the potential to switch on or off gene expression. The presence or absence of the repetitive DNA decides the nature of chromatin.
The facultative HC can be formed by nuclear RNAs as well. For example, the XIST gene plays an important role in the mechanism of X- chromosome inactivation. The XIST gene forms a transcript- XIST mRNA, that helps to inactivate the X chromosome in somatic cells of female mammals.
Another example of flexible HC is the inactive sex-vesicles at the pachytene stage of male meiosis.
The facultative HC is reversible, can be converted into euchromatin, are repetitive/ satellite DNA poor region and is enriched with transposons, especially LINES.
In addition, the present region is less polymorphic in nature. But cytogenetically It can’t be stained by the C banding.
Constitutive heterochromatin:
These regions are constant and entirely inactive in all cells and all organisms of a species. These regions are structurally made up of short repetitive DNAs, are satellite DNA and settled on telomere and centromere regions.
Genes located in it are very poorly active transcriptionally means, can’t form a protein at all.
Functionally, the constitutive heterochromatin, though are non-transcriptional, provides structural stability and flexibility.
The repetitive centromeric regions seal two sister chromatids during the metaphase using a sealing protein and also binds to kinetochores too.
It helps in the proper segregation of chromosomes during cell division.
Contrary, the repetitive regions- the non-coding ones, present on the arms as telomeres protect the vital region of chromosomes which is the “gene-rich” arms.
Telomeric sequences prevent loss of genetic material from arms and also prevent replication errors for genes, genes remain intact.
Chromosome 1,9, 16 and Y contains more constitutive heterochromatin regions than other chromosomes and stained darker by C bands.
Folding of varied satellite DNA helps in gaining the compact structure of the constitutive DNA region, moreover, it remains inactive and unchanged during all cell division phases. Enriched repetitive DNA’s make it more prone to polymorphism.
The presence of constitutive heterochromatin functions as a cohesive or glue to join two sister chromatids. Protein HP1 and Swi6 are essential needs to do the same.

Note: heterochromatin is one among many epigenetic gene silencing factors.
Structure:
Structurally, as we mentioned, the HC is made up of the satellite DNA and repetitive sequences majorly are tandem repeats and transposons. So either type of non-coding sequences is crucial for the structural consistency of HC.
Role of tandem repeats:
Collectively the major portion of the HC region is made up of tandemly repetitive sequences such as STR or VNTR means, short tandem repeats and long tandem repeats.
Note that each one is used as a marker sequence and are non-coding DNAs. The STRs are 1 to 6 bp long microsatellites present 5 to 200 times in a genome.
The VNTRs, considered as minisatellites, are 10 to 60 bp long and present 100 to 1000 times in a genome.
The size and number of repeats vary among individuals of the same species.
Transposons:
The transposable elements are sequences that can jump from one place to another in a genome. And is specifically present in the facultative HC. LINES- long interspersed nucleotide elements are a type of transposons, present in facultative HC.
It’s a kind of retrotransposon if transcribed, can form a reverse transcriptase enzyme. Though present in eukaryotes, they are inactive for thousands of years. Note that LINES are also repetitive.
A distinct position of HC within the nuclear periphery and around the nucleolus is referred to as Lamina-Associated Domains and Nucleolus- Associated Domains, orderly.
A major heterochromatin portion in mammalian cells is the LADs comprising 30% portion, are gene-poor regions, indicating that LADs are gene less, a low gene expressed and fewer gene density regions.
Contrary, The NADs which are present around the nucleolus, are rich in satellite repetitive DNA sequences, inactive rDNAs and other repressed genes.
Note that the NADs and LADs can be converted in order to regulate gene expression.
As we explained, genes for some kind of rRNA are present in the HC, their expression is regulated by nucleophosmin 1 and nucleophosmin 2. Mutation in either of the proteins causes dysregulation of the rRNA gene expression and produces abnormal nucleolar morphology.
PEV- position effect variegation:
Heterochromatin is inactive itself, but nearby genes can also be inactivated or repressed, through the effect known as PEV- position effect variegation.
Characteristics:
There are some special characteristics only the inactive heterochromatin region has. Here they are:
- It is a repetitive DNA region.
When sequences are repetitive, indicating that present frequently at regular intervals.
- Located on centromere and telomere of chromosomes.
- Are satellite DNAs, note that transposable elements are also considered heterochromatin.
- When relaxed, located on the periphery or scattered near the nuclear membrane.
- Are methylated:
The entire heterochromatin region is rich in methylation, means, blocks access to enzymes, the constitutive HC is more methylated while the facultative one is less methylated and discrete.
Besides, another epigenetic factor, activation and deactivation of acetylation processes also influence chromatin activity. At post-transcriptional, heterochromatin region forms by less acetylation activity on the N-terminal lysine residue.
- Stain darker using Giemsa stain or G bands and C bands.
- Are mostly transcriptionally inactive through various mechanisms.
- RNAi- RNA interference, causes inactivation of mostly facultative heterochromatin.
- Genetic recombination can’t be possible in HC region.
- Render enzymatic activities.
One of the most important properties of the HC region is that it doesn’t allow access to any enzymes such as DNAse or restriction enzymes. That’s the reason it remains constant and unchanged.
- Are late replication.
The entire 90% portion of HC is late replication. There are two reasons for that, first, transcriptionally active genes must be replicated on priority to avoid replication error and second,
It doesn’t allow access to enzymes easily, and so for the polymerase too and therefore replicate at slow speed and rate.
- Gene less or gene-poor region:
As we explained, the entire HC region is either gene-less or gene-poor. The Facultative HC are gene-poor whilst the constitutive HC are gene-less portion.
- Most of the regions are conserved since evolution, within the species and between species.
- Forming centromere:
Studies on model organisms suggest that the heterochromatin, especially the constitutive one, can aggregate at the centromere region, interact with other ribonuclear proteins and forms the centromere at interphase.
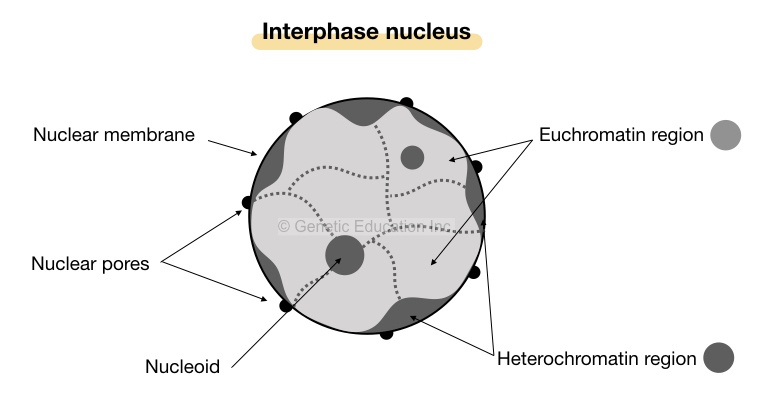
Note: the term heterochromatin was given due to its property of differential staining.
Functions of heterochromatin:
Transcriptionally inactive heterochromatin performs critical functions for our genome, even though considered as a junk one! Here are some:
Provides genetic or genomic instability.
Provides accurate chromosome segregation.
Provides structure to chromosomes.
Hold sister chromatids during chromosome construction.
Induce cell division.
Helps in transcription and translation
Functions as enzyme binding sites.
Note: Proteins that are involved in DNA packaging are categorized into nucleoprotein complexes.
How to study heterochromatin regions?
Before going inside the present topic, first, understand why to study heterochromatin. Sequences present in these regions perform varied functions to regulate gene expression.
And therefore in order to investigate gene expression or expression profile we should have to study it.
Many state-of-the-art and accurate techniques are available to study, however, karyotyping- a kind of cytogenetic technique is the first and widely used for doing so.
The whole principle of karyotyping depends on the differential banding pattern obtained by differential staining of euchromatin and heterochromatin. Ultimately, varied patterns of bands observed on chromosomes, make them distinguishable from each other.
Euchromatin stains lightly while heterochromatin stains denser due to the tight and compact DNA arrangement. GTG bands show darker blue bands as heterochromatin.
(let’s get in-depth, more technically)
Darker bands or regions indicate more heterochromatin. A specialized banding technique is known as constitutive banding or C bands performed in order to locate only constitutive heterochromatin regions, suggesting that 1, 9, 16 and Y possess larger constitutive heterochromatin rich chromosomes than others.
Centromeric regions of these chromosomes appear darker and thicker by C bands.
Noteworthy, no cumulative or quantitative information can be generated using the present technique. More extensive techniques based on the principle of “ChIP” are applicable to investigate HC.
ChIP is chromatin immunoprecipitation, used in two techniques, ChIP-chip- a microarray-based technique and ChIP-seq- a next-generation-based technique.
The objective of either technique is to investigate those chromatins associated with the proteins and are inactive.
This means it studies chromatin or DNAs only are tightly wrapped with histones or other nuclear proteins. Resultantly, information of DNAs which are transcriptionally inactive, collected.
Associated abnormalities:
To gene or chromosome mutation, epigenetic alteration also causes serious health issues, and HC too. Alteration in either constitutive or facultative heterochromatin causes varied problems.
In the facultative region, a mutation in the XIST gene causes an imbalance of the X chromosome inactivation, which means an expression of genes located on the X chromosome can be duplicated.
There are serious consequences of it in females. It leads to infertility and other mental or cognitive problems.
Another condition known as Roberts syndromes or ICF syndrome occurs by alteration in the constitutive region. It causes Immunodeficiency, Centromeric instability and Facial anomalies. It is a kind of recessive genetic condition.
The whole inactive X chromosome is an example of heterochromatin.
Conclusion:
Heterochromatin regions in a genome perform three critical and important functions such as to regulate gene expression, genome organization on a large scale and formation & structurization of the nucleolar region.
Heterochromatin is present in three regions commonly, are telomere, centromere and pericentric regions.
Sources:
Allshire, R., Madhani, H. Ten principles of heterochromatin formation and function. Nat Rev Mol Cell Biol 19, 229–244 (2018). https://doi.org/10.1038/nrm.2017.119.
Allshire RC, Madhani HD. Ten principles of heterochromatin formation and function. Nat Rev Mol Cell Biol. 2018;19(4):229-244. doi:10.1038/nrm.2017.119
Saksouk, N., Simboeck, E. & Déjardin, J. Constitutive heterochromatin formation and transcription in mammals. Epigenetics & Chromatin 8, 3 (2015). https://doi.org/10.1186/1756-8935-8-3


