During each round of replicative transposition, a transposon is duplicated (replicated) and inserted in a new position. Both DNA and retrotransposons follow the replicative mechanism of transposition.Tn3, phage Mu are some of the examples of replicative transposons in bacteria.
Replication copies DNA. It also facilitates gap filling and sealing DNA breaks during the repair process. DNA polymerase- a multifunctional enzyme helps in this.
If you are on this article, meaning, you knew about the transposons right! If not, let me quickly introduce it,
The short-repetitive DNA sequence, considered as ‘Junk DNA’, that can’t make a protein but can jump from one location to another in our genome, is known as the transposons or transposable elements.
Henceforth, the whole process is referred to as transposition, having two types; replicative and non-replicative transposition.
In the present article, we will introduce one of the two common transposition processes, which is a replicative transposition.
Key Topics:
What is replicative transposition?
When a transposon replicates, makes a new copy and leaves the old copy behind, is considered as the replicative transposons while, when transposons move from one to another place by leaving a gap behind is considered as the non-replicative transposons.
Put simply, the process of replicative and non-replicative transposition is also known as copy-paste and cut-past transposition, respectively.
See the image below,
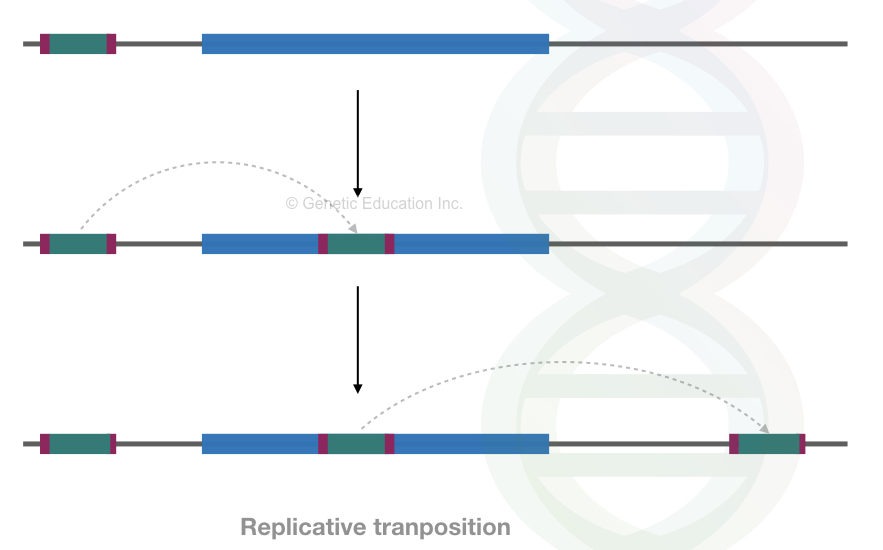
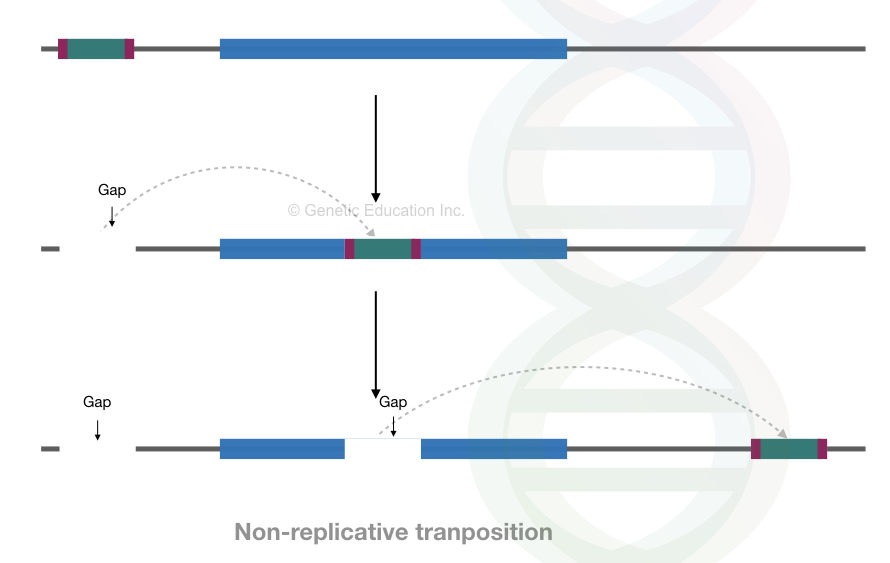
Transposition of the DNA transposons occurs via the strand transfer mechanism in the replicative transposition.
Replicative transposition of DNA transposons:
The explanation of the strand transfer mechanism for DNA transposon is given below,
Step 1:
At the beginning of the replicative transposition, the transposase enzyme assembles at the two ends of the transposable element.
This assembly migrates the entire transposon to another location.
This assembly creates the actual transposon, it helps in cutting the transposon at one location and inserting it to another location, simply called a “Cleaving and joining reaction.”
Step 2:
In the second step, the transposase protein catalyzes the reaction. The transposase protein cleaves the DNA at the two terminals of the TE itself and creates the free 3′ OH ends.
Furthermore, it also creates a gap in the target DNA sequences at which the transposon will insert.
The transposase proteins cleave the ends of the transposon as well as the host DNA. Overall, it creates 4 nicks, two at both the ends of transposons and two in the target DNA.
The nicking process in the transposons is ” sequence-specific nicking”, two nicks created on both ends of the transposons contain the same sequences because they are inverted terminal repeats.
The important event that happens here is, the transposase creates a free 3′ end at both ends of the transposon terminus.
These 3′ ends are very essential in the synthesis of DNA by a polymerase, meaning the gaps fill and new transposons will be created. However, in the cut-past mechanism, the nick remains as well.
Step 3:
In this step, a double branched DNA molecule is formed by the strand transfer mechanism.
The 3′ end generated in the previous step is now joined to the 5′ ends of the host DNA or the target DNA. The bond between them is covalent, contrary to this, the 5′ ends of the transposon sequence remain bound with the old DNA.
See the figure below,
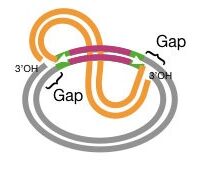
Step 4:
In the next step, the intermediate strand transfer complex creates two replication forks that facilitate a normal replication at the target DNA. Here, cleavage on the target DNA at the 3′ end accommodates a primer for the replication.
The replication machinery of the host cell assembled at the replication fork junction, the 3′ free end, serves as a primer, replication occurs from end to end.
Step 5:
In the last step, at the end of the replication, two DNA molecules each with the transposon are generated at the end of the process. The transposon is flanked by the short direct target site duplication.
These target site duplications will move with the transposons to another location and be inserted at a new location.
Note that the major limitation of the process of replicative transposition mechanism is frequent deletions and duplications that occur during.
The entire mechanism is shown in the figure below,
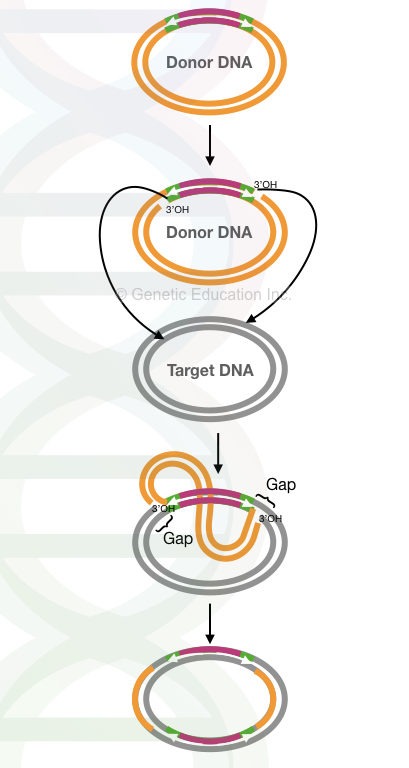
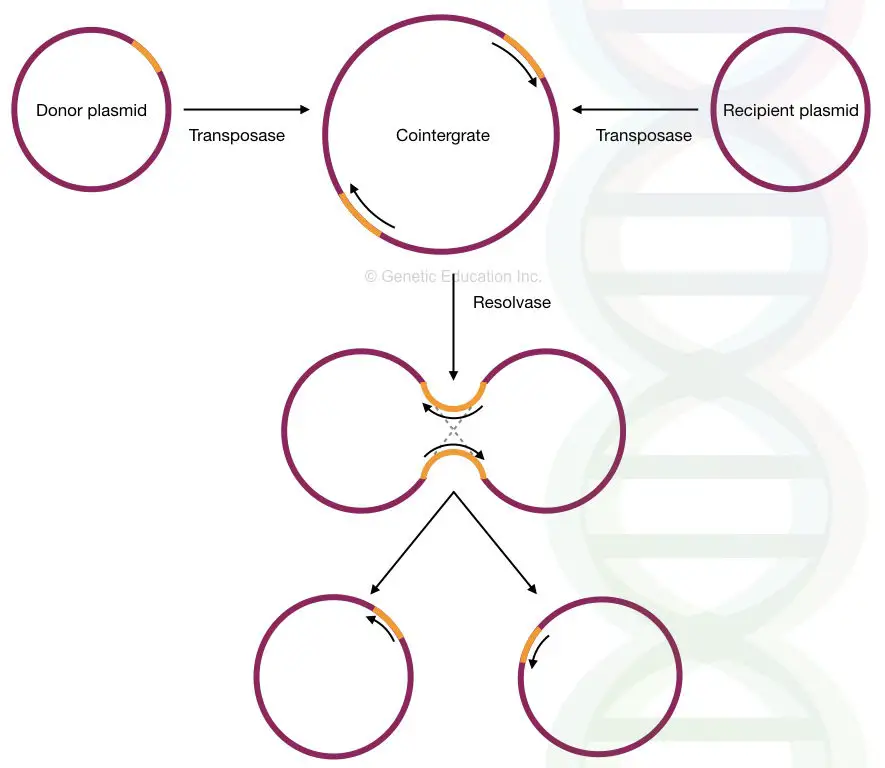
In the plasmid DNA, the replicative transposition occurred through the cointegrate structure.
The replicative transposition starts by the fusion of two plasmids, donor and recipient. It generates a structure known as “co-integrates” via transposase-mediated catalytic activity.
The transposon replicates in the cointegrate structure and creates another molecule of it in the same direction. During the process, in the next step, recombination occurs between both transposons. Transposon and resolvase govern the process at the “res” site.
The resolvase is another bacterial protein that suppresses the transposition process. After that, the two molecules are detached and two plasmids with two transposons are generated.
See the figure below,
Replicative transposition of Retrotransposon:
The retrotransposons also follow the replicative strand transfer mechanism of transposition.
Instead of direct replication, here the reverse transcription mediates the transposition of DNA, via the RNA intermediate.
In the very first step, using the cellular RNA polymerase, the RNA is synthesized from the DNA of the retrovirus or retrovirus-like particle through the mechanism of transcription.
Structurally, as we knew, the long terminal repeats of the transposons aren’t encoding any protein and also have a promoter region.
The transcriptional machinery forms the full-length RNA molecule from the one of the LTR. From the RNA, a new DNA is formed through the reverse transcription mechanism.
Here, the cDNA, or complementary DNA generated by the process doesn’t have any copy at the previous location.
In the next step,
The recombination occurs via the recognition of the integrase protein by the cDNA. Once the integrase is recognized by cDNA, it is assembled on both ends of the DNA.
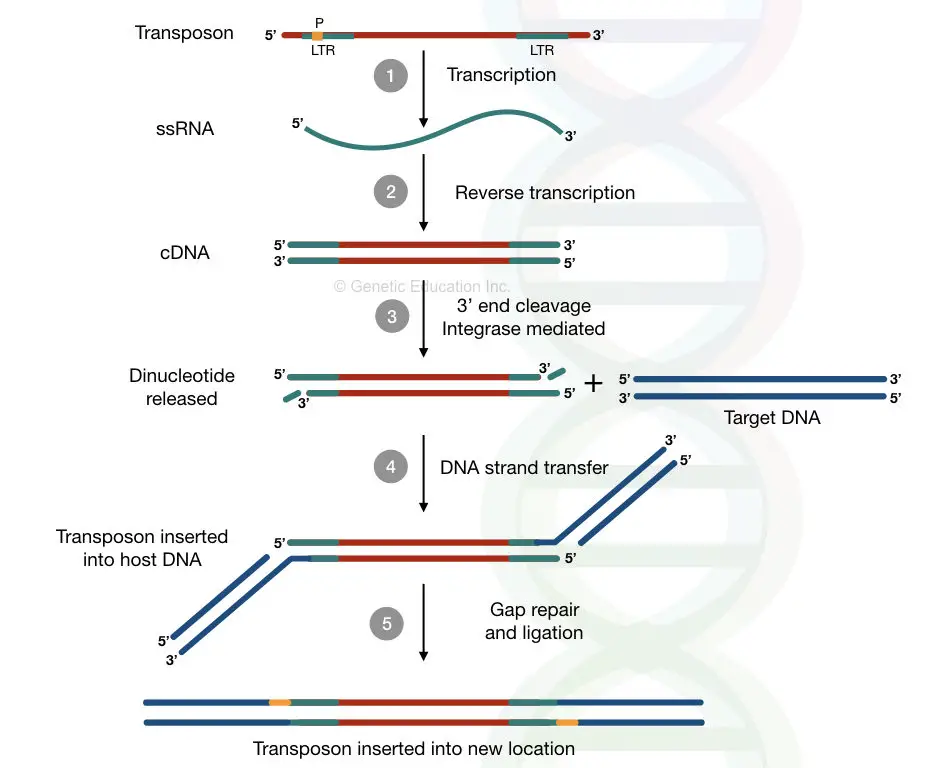
Likewise the strand transfer mechanism of the DNA transposition, the gap is generated at the 3′ end of each strand of the DNA by cleaving some of the nucleotides from this end by the integrase.
By using the DNA strand transfer mechanism, the integrase catalyzes the insertion of the modified DNA into the host genome DNA.
Once the gaps are filled by the DNA repair mechanism by a DNA repair enzyme, the process of recombination completes.
After the completion of recombination, the target site duplication is generated on both the ends of a new transposon possibly because of the same DNA repair mechanism on both sides.
This is a complete process of replicative transposition for DNA transposons and retrotransposons.
TY element in yeast and Copia element in Drosophila whereas Tn3, Tn5 and phage Mu follow the replicative mechanism for transposition.
Our additional resources related to this topic:
Transposons: A Jumping Entity and a Foe with Benefits
- History, overview, DNA transposons, retrotransposons.
Transposase, Transposons and Antibiotic Resistance in Bacteria
- Bacterial transposons, Is elements, composite transposons, transposons and antibiotic resistance.
- Transposons in Drosophila, transposons in maize, transposons in yeast.
Role of transposons in evolution
- How transposons are involved in the process of evolution.
Conclusion:
In a replicative transposition, a new copy of the transposon is inserted at a new location by leaving the original TE at its native position. Both the DNA and retrotransposons follow the replicative transposition. New alleles originated due to these types of transposition.
Replicative transposition played an important role in the process of evolution and the generation of new variations into nature.
Sources:
- Griffiths AJF, Gelbart WM, Miller JH, et al. Modern Genetic Analysis. New York: W. H. Freeman; 1999. Mechanism of Transposition. Available from: https://www.ncbi.nlm.nih.gov/books/NBK21287/
- Transposons By Nature publication.


