“Microarray array is a powerful hybridization-based genetic testing technique. DNA microarray, SNP array, chromosomal microarray, Methylation microarray, exon microarray and CGH are a few variants. Learn about different types of microarray-based genetic testing techniques in this article.”
Genetic testing allows us to study genes, DNA, chromosomes and the genome and helps us unfold the mystery of our DNA. It allows us to understand genes, related pathways and how diseases occur. Microarray is one such testing technique scientists use.
Microarray genetic test work on the principle of DNA hybridization. Here, hybridization occurred between probes and the target sequence gives us a positive signal. The labeled fragment of DNA when bound with the probe, releases fluorescence which is recorded and evaluated.
Probes are usually immobilized on a solid glass chip. Millions of known sequence probes are ready to hybridize with the sample. This makes the technique powerful enough to screen thousands to millions of alterations in a single run.
The microarray can study genes, related alterations, SNP, indels, gene expression and other chromosomal polymorphisms from the DNA sample. Learn about the basic concept of microarray-based genetic testing and its types in this article.
Stay tuned.
Key Topics:
What is Microarray-based genetic testing?
Microarray-based genetic testing is a technique that can analyze sequence variations from the sample, effectively. It closely looks into a person’s DNA or genome and discovers variations or alterations associated with any disease or health condition.
It’s as we aforesaid, a hybridization-based technique with slight modification. Instead of probes, the template DNA is labeled with the fluorescent dye. The probes in huge numbers (thousands to millions depending upon the chip) are immobilized on a chip, each specific for each sequence/fragment/gene alteration.
How is Microarray-based genetic testing done?
The steps in the process are here.
- Sample collection: A blood or cheek swab sample is collected.
- DNA or RNA isolation: Depending on the requirement, either DNA or mRNA is isolated from the sample.
- DNA fragmentation: Using an artificial fragmentation technique, the sample nucleic acid is fragmented into smaller chunks.
- Fragment labeling: The fragments are labeled with the detectable fluorescent tag. Red for the sample and blue or green for normal.
- Hybridization: fragments are allowed to hybridize to probes present on the chip under strict hybridization conditions.
- Detection: A fluorescence detector records the signals which are processed using a computer program.
Hybridization Removes the tag on the fragment and releases fluorescence. The signals are detected and processed by a microarray scanner.
Using the sequence information, copy number variation data associated with diseases, health conditions or genes and SNP data, scientists prepare millions of probes and immobilize them on the solid chip.
A strong fluorescence signal indicates strong hybridization and the presence of a target sequence. The poor and weak signal indicates poor hybridization and the absence of the target sequence or variance.
Scientists or healthcare professionals can determine in which amount a particular sequence, variation, CNV or SNP on a gene is present in the sample through fluorescence signal analysis. Noteworthy, the entire process is fully automated, so additional analysis tools or programs are not required.
This technique has been used in the healthcare, diagnosis industry and clinical studies due to the fact that it enables us to study thousands to millions of variants associated with a gene or disease in a single run.
Scientists and healthcare professionals closely study the data regarding variations associated with a disease, determine genetic predisposition for a disease, detect various chromosomal anomalies, study drug response and identify drug markers.
A few popular types of microarray techniques used for genetic testing are discussed herein.
Related article: DNA Probes: Labeling, Types, Applications and Limitations.
Types of Microarray-based Genetic Testing:
DNA microarray, Chromosomal microarray, SNP microarray, Methylation microarray, exon microarray and CGH are some of the popular microarray techniques used. Each one is explained elaboratively here.
DNA microarray:
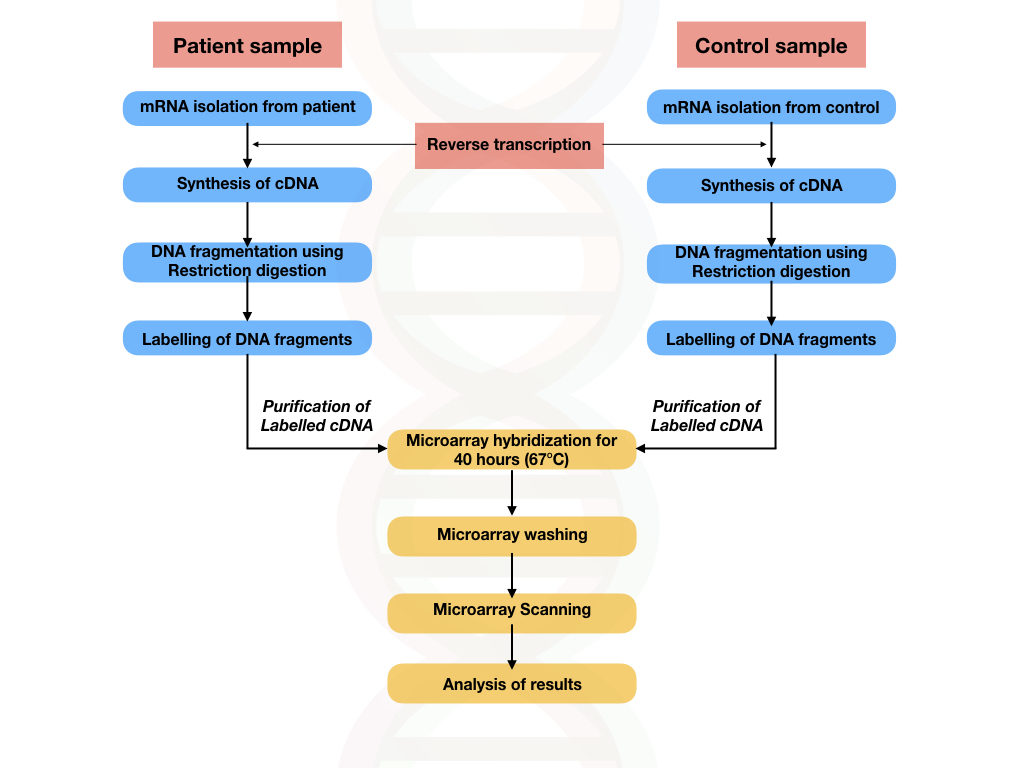
DNA microarray is also known as gene expression microarray or expression array. It can determine the expression of a gene. Meaning, the number of genes. A few hundred to a few thousand genes’ expression can be determined in a single DNA microarray using the same hybridization principle.
Here, instead of probes specific to a particular mutation or alteration, gene-specific probes are immobilized on the chip which upon hybridization gives us a signal regarding the expression.
A strong signal indicates higher expression and vice versa. Gene expression microarray is extensively used in tissue-specific gene expression studies, to understand genetic regulation, identify novel gene markers and their expression and help understand disease mechanisms.
However, in the gene expression microarray studies, instead of DNA, mRNA is isolated, reverse transcribed into cDNA, fragmented, labeled and used for hybridization.
Chromosomal microarray:
Chromosomal microarray is extensively used as a primary diagnostic tool in the healthcare setup. It can identify chromosomal alterations associated with developmental delay, autism, intellectual disability, cancer and other complex diseases.
Here, chromosomal region-specific probes are immobilized on the solid surface. It can detect deletion, duplication or insertion from an individual and all chromosomes from a DNA sample.
It has many thousand-fold excellent resolution and detection range than conventional karyotyping or FISH. Another amazing benefit of chromosomal microarray is that it doesn’t require extensive and tedious cell culture and chromosome harvesting processes.
Chromosomal microarray is also known as array comparative genomic hybridization. It is popularly used in preimplantation genetic screening and screening of birth defects. Note that it can detect copy number variations larger than a few kilobase pairs not less than that.
SNP microarray:
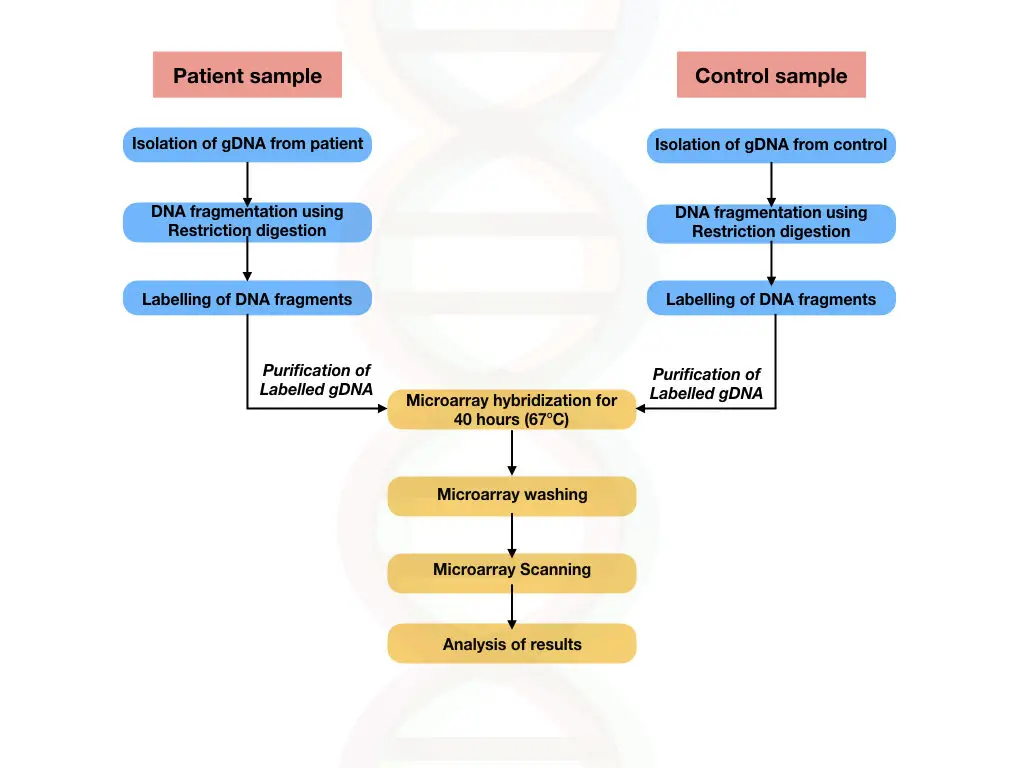
An SNP (Single Nucleotide Polymorphism) microarray is also known as genotyping microarray and is used for the detection of single nucleotide polymorphisms from a gene or genome (depending upon the requirement of the experiment).
It’s a tool largely used in genome-wide association studies and population genetic screening as it determines various genotypes or SNPs associated with a disease, present in the genome or particular population.
SNP microarray helps researchers to understand the large-scale effect of SNP and thereby they can classify SNPs as pathogenic, non-pathogenic, rare and common SNPs for a particular population.
Such studies help to study genetic alterations associated with diseases, complex traits and phenotypes and drug response at a population level.
Exon microarray:
Exon microarray is a specialized version of the conventional microarray that can only study exons from the genome. Exons are coding regions of a gene that directly participate in protein synthesis.
By only looking into the exons, scientists can understand the impact of exonic alterations on protein synthesis. They can also study splicing events, transcript isoforms, exonic variants, etc.
Again the instrumental, assay requirements and process will remain the same, like other microarrays but only chip content will vary.
Methylation microarray:
Methylation is a type of epigenetic modification which regulates gene expressions. Methylation microarrays focus on the highly methylation-specific regions viz, CpG island of the genome.
Probes specific to such regions can hybridize and give us signals regarding the pattern of methylation thereby the pattern of gene expression. Methylation microarrays have applications in epigenetic studies.
Notedly, the methylation microarray is a bit different process in which an additional bisulfite treatment step is added. In bisulfite treatment, the unmethylated cytosines are converted into the Uracil and help distinguish methylated and unmethylated regions effectively.
The rest of the steps of fragmentation and labeling will remain the same. Methylation microarray helps to study the epigenetic modification or methylation associated with disease, aging and developmental process.
CGH microarray:
CGH is a comparative genomic hybridization microarray which is a specialized technique used in comparative studies. Two different DNA samples, for example, tumor DNA or normal DNA, are used for competitive hybridization.
In the comparative microarray, the expression pattern from both samples are analyzed to determine the difference. CGH is one type of chromosomal microarray we can say, as it largely determines copy number variations.
Wrapping up:
Microarray evolution certainly revolutionized the healthcare and diagnostic industry with its extraordinary precision, wide range and power to study millions of genomic loci, However, it can only determine known genomic alterations only, that’s its only limitation.
DNA, chromosomal, exon, methylation and SNP microarrays are extensively used in research and diagnosis. Notwithstanding, highly-customized microarray arrays specific to diseases or genes are now available in the market.
For example, the microarray for breast cancer detection covers all the known, common and pathogenic variants from the BRCA1 and BRCA2 gene. Rather than focusing on SNPs or indels, it covers all the alterations and their expression profile.
I hope this article makes sense for you. I will try to explain each type of microarray genetic test in detail in each separate article. Till then read our previous articles, bookmark this page and subscribe to our blog.