CRISPRi or CRISPR activation is a process to upregulate gene expression or activate genes by effector activators, dcas9 and sgRNA.
In 2021, CRISPR have deep penetration in diverse scientific fields. It’s not restricted to gene therapy or gene editing only. Meaning, it can perform various functions ranging from monitoring the cell cycle to gene expression.
And as scientists explore further, more and more fruitful and worthwhile scientific approaches are developed. For example, one CRISPR can provide a history of a carcinogenic cell, how it behaved during various stress conditions.
Although CRISPR was basically derived from a bacterial defense system only to perform target-specific gene editing. In recent years, it has changed the scientific fraternity totally.
If you would like to learn more applications of CRISPR, read this article: Applications of CRISPR-CAS9.
As we discussed in our previous article- CRISPR interference, the technology can repress gene expressions of a cell. Not only a single but many genes’ expression at a time.
The CRISPRi (interference) concept is designed to repress the gene expression but do you know, one CRISPR technique can activate genes as well?
Yes, it’s possible.
Before the discovery of CRISPR activation, other traditional approaches used to activate a gene were tedious, costlier, less effective and can work on a single gene at a time.
CIRSPRa or CRISPR activation is designed to uplift gene expression or activate genes at multiple loci.
I am writing this article to explain to you what CRISPR activation is, its mechanism, applications, advantages and disadvantages.
So this article will boost your CRISPR knowledge and will make your concept more clear. Stay tuned.
We are exploring CRISPR and related technology. I have written 5 articles to date that you can read by revisiting this category:
Key Topics:
What is CIRSPRa or CRISPR activation?
The CRISPRa short for CRISPR activation has been designed to activate genes and upregulate gene expression using synthetically designed dCAS and sgRNA.
“Activation”, simply means, to activate- genes. And that’s what this technique can do; effectively, precisely and efficiently.
Gene activation by CRISPR activation was first reported in 2013 by Bicard et al., & Perez- Pinera et al., 2013.
Three important ingredients a CRISPRa needs besides the CRISPR itself are; activator (used to activate the target gene’s expression system), sgRNA and dCAS9. I will explain each one in the upcoming sections.
Let us first understand the mechanism of activation via CRISPR.
Important terminologie:
sgRNA: sgRNA is a single-stranded guided-RNA or guided RNA that navigates the CAS9 at the target location. It has two parts tracrRNA and crRNA.
CAS9: The CAS9 or CAS is an endonuclease enzyme that cleaves DNA at a target location.
dCA9: dCAS9 is a dead CAS protein that lacks endonuclease nucleophilic activity. It can’t cut DNA.
Mechanism of action:
When a synthetically designed and activator-linked sgRNA or dCAS9 is guided to the target gene, it triggers a gene expression system.
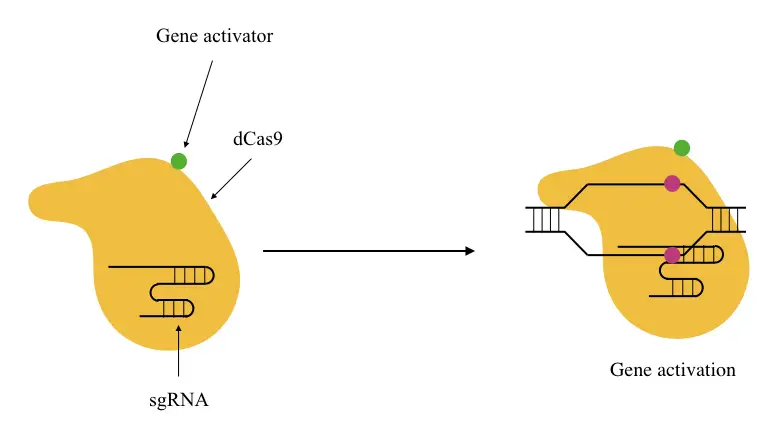
It binds upward to the promoter or near transcriptional start site (TSS) and recruits other enzymes or proteins to initiate gene activation (or transcription). The dCAS9 is a specially designed CAS that lacks endonuclease activity.
dCas9 based activator approach and sg-RNA-based activator approach are the two most popular ways the transcriptional activators can be transported to the activation site.
In the dCas9 based activator approach, the dCas9 protein is engineered to fuse with various transcriptional activators while in the sgRNA-based activator approach the sgRNA is designed to fuse with various activators.
Notedly, the sgRNA-based activator approach is quite complicated and less efficient in comparison to dCas9 based approach.
Components of CIRSPRa:
As we said, sgRNA, dCAS9 and activators are required to perform the artificial gene activation. Let’s talk about each one in detail.
sgRNA:
Original article: single-stranded guided RNA.
The sgRNA is the single-stranded guided RNA with spacer and scaffold regions. The spacer and scaffold regions have sequences complementary to the target location and the dCAS9, respectively.
The guided RNA usually has tracrRNA and crRNA parts. It’s synthetically designed to bind at a specific location. Separately synthesized, target-gene-specific sgRNA can be used for activating many genes.
The guided RNA for the CRISPR activation is designed to bind to its complementary sequence upstream to the transcriptional promoter or at TSS. However, designing the sgRNA for gene activation is a bit difficult and complex process.
The major problems arise due to the inaccessibility of transcriptional start sites (TSS) and the presence of more than one TSS which restricts the use of sgRNA mediated activator systems.
Advanced machine learning techniques and algorithms help to design specific and efficient sgRNA for CRISPR activation and thankfully are now available with many distributors.
Here for gene activation, an activator has been attached with the sgRNA which guides it along with the dCas9 to the target location. Some common synthetic activators such as SAM, Pumilio/FBF and riboswitches give excellent results when fused with the 3’ end of the sgRNA.
Do you know?
Tanenbaum et al., developed the SunTag system for gene activation in 2014.
dCAS9:
Original article: dead CAS9 protein.
The detailed importance, applications and how the dead Cas9 system is developed, is explained in our previous article. Anyway, let me give you a brief overview.
Technically the dCas9 or dead CAS is a type of nuclease that lacks endonuclease domain. So basically it can’t cleave DNA but the intact DNA binding domain facilitates target-specific binding.
Put simply it’s used as a vehicle to transport the activator effector molecule at the site of gene activation.
Much like the sgRNA, the dCAS9 can also locate activators at the site of gene activation. VP-64, SunTag and VPR are common activators that give excellent results when fused with the dCAS9.
Transcriptional activators:
Often known as Programmable transcription activators- Transcriptional activators are proteins that bind to DNA or a gene upstream to a promoter and induce transcription activity. It stimulates gene expression by activating and recruiting other proteins and enzymes.
The domain where the activator binds either on dCas9 or sgRNA is known as the activation domain having the power to locate various activators at the core promoter region of the gene.
The complex of various transcriptional factors, proteins and enzymes is known as the transcription preinitiation complex.
Take a look at some of the most popular TAs used in the CRISPRa.
VP64:
VP64 is a traditional and strong transcriptional activator, often fused with the dCAS9 (known as dCAS9-VP64) to elevate gene expression. It’s one of the primitive techniques that has a moderate level of up to 2- fold gene activation.
Therefore it’s considered as “first-generation CRISPR activators.” It can do its work during the initiation or elongation of transcription.
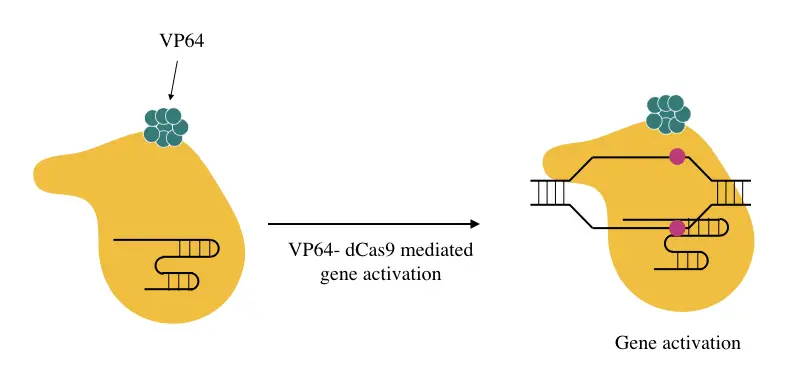
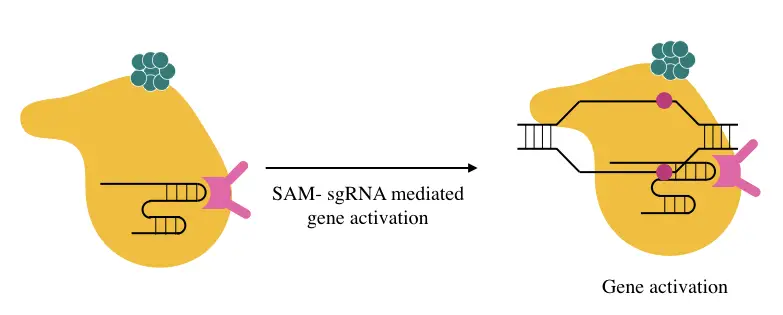
SAM:
SAM, short for synergistic activation mediator, is yet another important transcriptional activator that requires a specifically designed sgRNA to fuse and perform transcription.
It has more accuracy and targets to activate and is therefore widely used. The MS2 loops of SAM fuse with the 3’ end of the sgRNA which later on requires more activators like HS1 and p65 domains.
SAM greatly increases transcription activity for a single gene, however, SunTag and VPR perform excellently great for multiple gene loci.
SunTag:
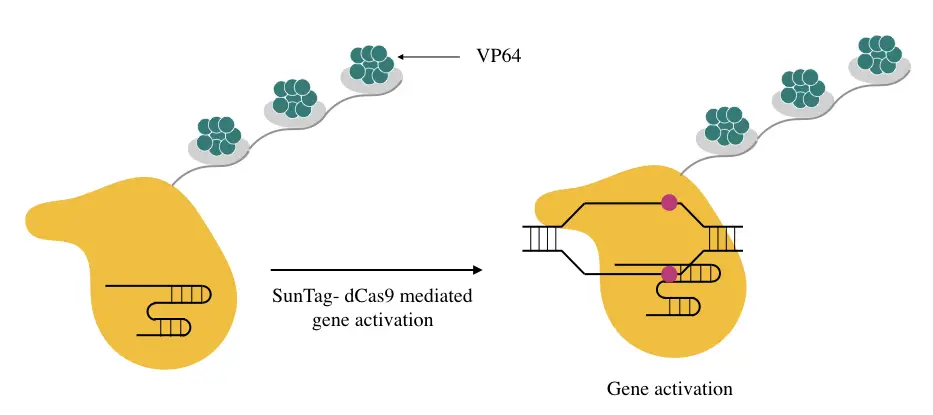
The SunTag is by far one of the most excellent activator systems which use multiple units of VP64 to recruit transcription machinery at target loci. The repetitive peptide units join various VP64 proteins and fuse them with the dCAS9.
The present activator works consistently excellent for multiple gene activation but has a lower activation index than the SAM for a single gene.
Nonetheless, the tedious construct-making process and use of antibodies in the process is the biggest shortcoming that restricts its use.
VPR:
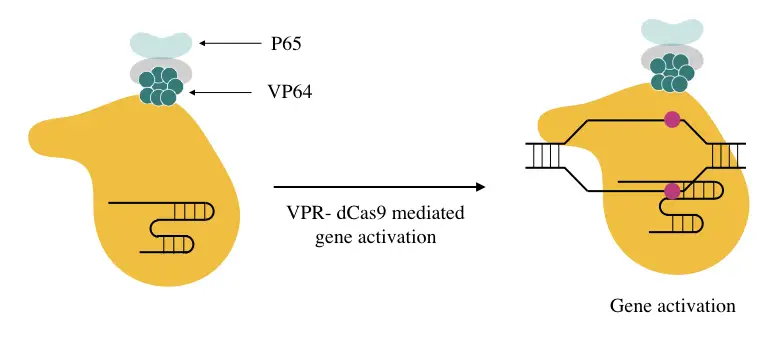
The VPR activation construct forms when the other two proteins/activators like p65 and Rta are fused with the dCAS9- VP64 complex. So basically, it’s a higher and updated version of the native dCAS9- VP64 complex, showing greater activity than the native one due to the presence of three different gene activators- VP64, p65 and Rta.
The VPR can work for a single and multi-gene locus much like the SunTag but has lower activation power than the first-generation CRISPR activator. Studies also suggest that the VPR is capable enough to activate multiple genes in the same cell types using separate sgRNA.
Besides these major activators which are widely used in the CRISPRa, SPH and CBP are other two activators, researchers are looking for future applications. The SPH is the combination of SAM and SunTag that increase activation manyfold in comparison to individual activators.
On the other hand, the CBP is a histone acetyltransferase domain fused with the dead CAS that activates transcription at the chromatin structure level by facilitating rearrangements.
Comparative studies on human, mouse and drosophila cells suggest that the SAM, SunTag and PVR activators deliver excellent results when fused with the dCAS9, instead of sgRNA.
Related article: CRISPR-Cpf1(Cas12a) Gene-Editing System- Explained.
Advantages of CRISPR(a) activation
- More effective, efficient, rapid and simple.
- Perform gene activation on multiple gene loci.
- Only required synthetically designed sgRNA or dCAS9 only.
- Separate synthetically designed sgRNA is used for different gene loci.
Limitations of CRISPR(a) activation
- It is more expensive than traditional gene-editing methods.
- Various activators are required for various genes. However, every transcription activator has a different level of activity which makes it restricted to use.
- It requires separate sgRNA for each target gene.
- It’s also difficult to target sgRNA for various genes as some genes have more than one transcription start site.
Applications of CRISPR(a) activation
- Activating a target gene and increasing expression pattern many folds for a single or many genes.
- The use of CRISPR activators in in vivo systems for creating transgenic organisms.
- Genomics-wide screening of activated loci using the CRISPR activators.
CRISPR activation validation assays:
An experiment can’t be said as complete until validated, RT-qPCR, Western blotting and Immunofluorescence analysis has been three important methods used to validate the gene activation process by CRISPR.
Among all three techniques, western blotting works on the level of protein produced which is the most traditional method. The Immunofluorescence technique relies on the use of fluoro-conjugated antibodies.
Both techniques are less preferred in comparison with the RT-qPCR. The reason is that the quantitative real-time PCR quantifies every mRNA formed from a gene. Therefore is most accurate.
Quantitative PCR is faster and cost-effective as well. Two broad techniques used to quantify the nucleic acid present in the sample is the dye-based and probe-based hybridization method.
Validation confirms if the CRISPR modification worked or not.
Important outcomes:
Previous research suggests that some gene promoters have multiple regions to bind for the same type of gene activator or transcriptional factors. Techniques like VP65 and SAM are designed in such a way that it requires multiple TA to promoter regions.
This increases transcriptional activation manyfold.
The binding of TAs upstream to the transcriptional start site increases transcriptional activity dramatically to an unpredictably high.
The chromatin states around the promoter region have a significant effect on the CRISPR activation process, as the pool of the same TAs can’t work for all genes, equally.
Gene to gene the efficiency of the selected system varies, for example, some give excellent results using the PV64; but the same doesn’t give equally good results for other genes.
Wrapping up:
Though a few well-studied gene activation techniques exist, the CRISPR activation technique has been by far one of the most effective ones. However, Yet not approved for real-world use.
Research data suggests that scientists have collected amazing research data explaining various gene activation, multiple gene activation and tissue-specific gene activation in different types of cell lines.
The main base of CRISPR activation is designing the sgRNA and selecting the dCAS9 and activator for transcription. In recent times, many different effective activators are now commercially available.
Sources:
Casas-Mollano JA, Zinselmeier MH, Erickson SE, Smanski MJ. CRISPR-Cas Activators for Engineering Gene Expression in Higher Eukaryotes. CRISPR J. 2020;3(5):350-364. doi:10.1089/crispr.2020.0064.
Ho HI, Fang JR, Cheung J, Wang HH. Programmable CRISPR-Cas transcriptional activation in bacteria. Mol Syst Biol. 2020;16(7):e9427. doi:10.15252/msb.20199427.