In the touchdown PCR, “By sequentially decreasing the annealing temperature during each PCR cycle, the chance of the non-specific binding can be reduced.”
Every PCR technique is evolved to eliminate unwanted amplification during the PCR reaction. The unwanted amplification, a non-specific binding results in false-positive or false-negative results.
Also, the unwanted primer-dimer amplification results in the shorter non-specific bands which makes the primers unavailable for the target amplification.
Primer-dimers and non-specific binding are two major setbacks of any PCR reaction or we can say it is a kind of curse for any PCR reaction.
Non-Specific bindings are the amplicon other than the target sequence amplification. Hence it is necessary to stop any unwanted amplification in PCR reaction.
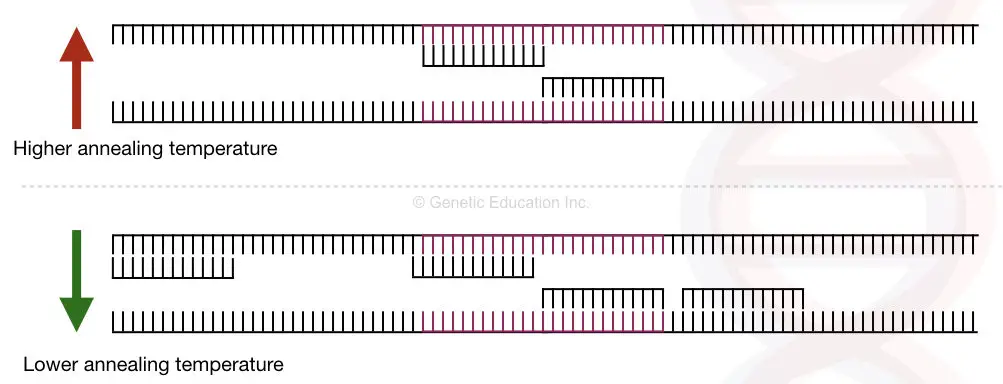
By increasing the annealing temperature, the primers cannot bind to other than its complementary sequence and result in specific amplification, however, it is not always the case.
Increasing the annealing temperature failure in PCR reaction can also be possible.
So what to do?
Is there any technique that avoids unwanted amplification and is still a success in the PCR reaction?
In this article, we will discuss touchdown PCR which facilitates high specificity in any PCR reaction. The content of the article is,
Key Topics:
What is touchdown PCR?
The touchdown PCR is the modification of the conventional PCR in which the high specificity of amplification is achieved by reducing the unwanted amplification on sequentially decreasing the annealing temperature after each PCR cycle.
For understanding the basic concept of the touchdown PCR, we have to understand the mechanism of how the primers anneal to the PCR template DNA.
The primers are the set of the short single-stranded DNA used in the PCR reaction which facilitates the amplification by providing the free 3’-OH ends to Taq DNA polymerase.
For more detail on the DNA primers, read the article: PCR primer design guidelines
For that, we have to design our target sequence-specific primers by using online tools. The primer must be between 18 to 22 nucleotides long, having 50% to 55% GC content and can not form the primer dimers.
The annealing temperature of the primers must be between 55°C to 65°C (ideally).
The annealing temperature of the primers is the temperature at which the primers bind to their specific complementary sequence on DNA.
Whereas, the melting temperature is the temperature at which the primer dissociates from the complementary DNA sequence.
Also, the melting temperature is defined as the temperature at which half of the DNA strand is dissociated or denatured.
Both definitions are right.
So ideally, the annealing temperature should be lower than the melting temperature, and then the primers can bind to its complementary sequence.
Generally, the annealing temperature is 5°C lower than the melting temperature but it is just an approximate assumption.
Hence it might possible that at a given annealing temperature the primer may bind to the sequence other than its complementary sequences.
It is hard to calculate the exact annealing temperature for impossible templates such as high GC-rich DNA.
All these problems result in non-specific unwanted amplification.
To overcome this problem, the touchdown PCR is designed.
Principle of touchdown PCR:
In the touchdown PCR, the annealing temperature is selected 10°C higher than the melting temperature. Therefore primers can bind to only their specific and complementary sequence. After each cycle, the temperature is decreased by 1°C.
Even if the DNA is not amplified in the initial cycles due to a higher temperature (temperature higher than the melting temperature), it will be amplified in the successive cycles, because we are decreasing the temperature after each cycle.
The beauty behind the technique is that even if we are decreasing the temperature, non-specific bindings are not formed after the amplification of the first template.
Once the DNA sequence of our interest is amplified successfully, that amplified DNA is work as a template for further PCR cycles.
Read more:
Protocol for the touchdown PCR:
Now, let us understand the process by taking one simple example.
Suppose the annealing temperature for our PCR is 55°C.
Divide the PCR cycles into two stages.
In the first stage, decrease the annealing temperature by 1°C after each cycle.
Our annealing temperature is 55°C so, select the temperature of 66°C for the first round of amplification.
Now after each cycle the temperature decreases by 1°C hence for 15 cycles the temperature will decrease to 15°C from the original temperature.
So after the end of stage one, the annealing temperature of the reaction will be around 51°C.
In the second stage, apply this annealing temperature for the rest of the 20 to 25 cycles.
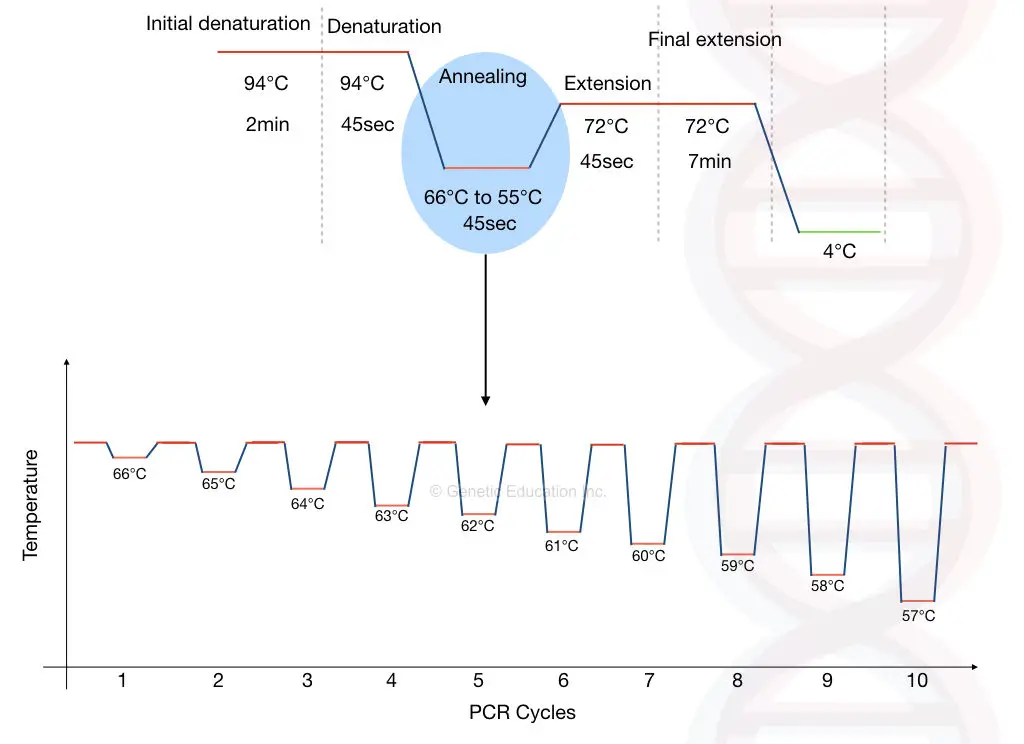
The protocol is shown in the figure below,
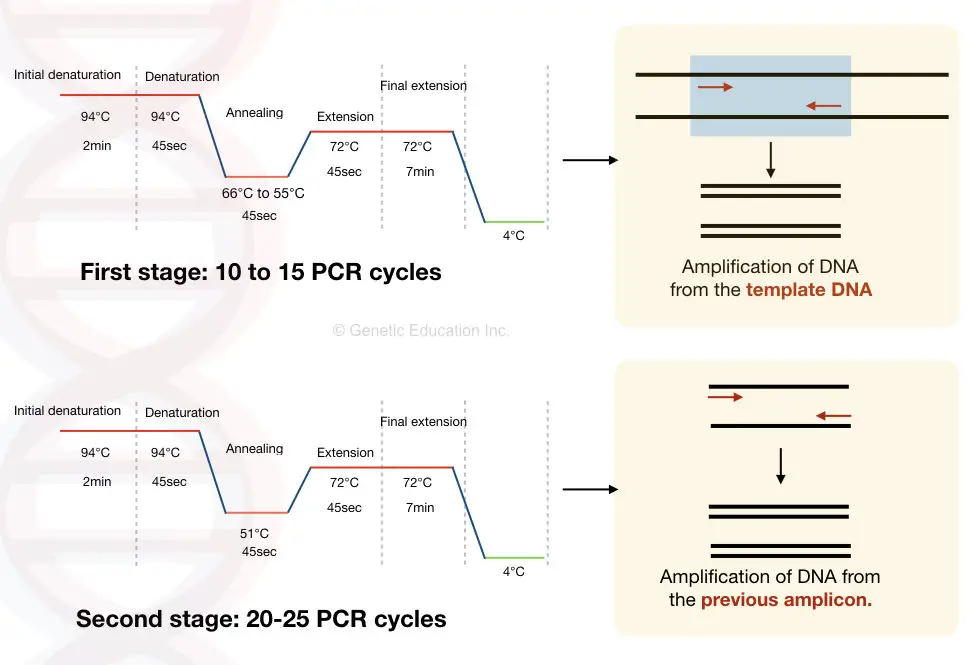
The combination of touchdown PCR with the hot start PCR makes this technique even more advanced.
If we combine an initial hot start step at the beginning of the PCR reaction at 96°C to 97°C for 2 to 3 minutes, it will make the template ready for the higher annealing temperature in the next step.
This combined method is more suitable for impossible templates such as GC-rich DNA.
As the main aim of the touchdown PCR is to decrease the non-specific binding, prepare all the reactions on the ice to avoid the early activity of the Taq.
Another best use of the touchdown PCR is its collaboration with multiplex PCR.
Want to know more about the multiplex PCR? Read this article: what is multiple PCR?
In the multiplex PCR, with the help of the multiple sets of primer, two or more target sites on a template DNA or on the multiple template DNA site can be amplified in a single reaction.
Obviously, it can save time.
But multiplex PCR has one major limitation,
Non-specific, unwanted and cross bindings of primers.
By combining the multiplex with the touchdown, the non-specific bindings can even minimize more in the multiplex PCR.
Here the annealing temperature is too high, each set of primer binds only when its optimum annealing temperature is provided.
Henceforth none of the primers can bind non-specifically.
Even,
The “hot start multiplex touchdown PCR” is more powerful.
“The annealing temperature can be altered as the composition of the PCR reaction buffer is altered, this will results in non-specific binding.”
Do not use more cycles for the touchdown PCR, this will again lead to non-specific binding.
Advantages of touchdown PCR:
The touchdown PCR reduces the primer-dimer formation capacity of primers.
It will also provide higher specificity by reducing the non-specific and unwanted bindings of the primer to the template DNA.
The technique is extremely useful in the templates having higher GC contents.
It also saves the time of the reaction.
Disadvantages of touchdown PCR:
Although it is highly specific, if not performed well the touchdown PCR also gives non-specific results.
Read more,
- CTAB DNA extraction buffer for plant DNA extraction
- Importance of Tris-EDTA (TE) buffer in DNA extraction
Wrapping up:
Touchdown PCR is also one of the best modifications of the conventional PCR method. the method is the best choice for the repetitive DNA sequence PCR. The ultimate goal of any PCR modification is to increase the yield and precision of the results, TD-PCR gives both.
Touchdown PCR is one of the best methods for increasing the specificity in the PCR reaction.