“Gene mapping is a process or method of discovering the location of genes on a chromosome.”
Or we can say
“Gene mapping is a method used to determine the place of a gene, the distance between the genes or identification of the location of a gene in a genome or on a chromosome.”
Gene mapping is also known as “genome mapping” or “mapping of a genome” helps researchers navigate around the genome and create landmarks.
As with the landmarks for different addresses, here, the landmark for the different genes can be created using some genetic methods.
These landmarks help to find out regulatory sites, repeated DNA sequences or a gene, even it helps in discovering new genes related to some phenotype.
In the year 1911, Alfred H. Sturtevant created the first genetic map denoting the location of genes on a chromosome.
Working on the model organism Drosophila melanogaster, he found that the genes are arranged in a linear manner on a chromosome.
He also observed that the distance between any two genes is fixed if the genes are located on one particular chromosome and the length of the chromosome is the same in all organisms of that species.
Furthermore, he realized that the location of genes on a chromosome could be determined by calculating the frequency of crossing over.
In his final remarks, he had said that “the genes are close to one another are linked and are more likely to inherited together, contrary to this, the genes are far from each other are less likely to be inherited together and thus are not linked.”
The genes located on the same chromosome are said to be “linked” and the distance between both is called the linkage distance.
His findings can be concluded like this,
If two phenotypes are inherited together in a pedigree, both can be said “linked” and located nearer to one another on a chromosome.
The frequency of crossing over is low between distantly related genes and hence can not be inherited together in offspring.
The map constructed by Alfred H. Sturtevant is called the linkage map, interestingly, the first human genome map was developed based on the linkage map method. However, the first genetic map of using an RFLP marker was published in 1987, 400 different RFLPs (restriction enzymes) markers are applied in constructing the entire map of the human genome.
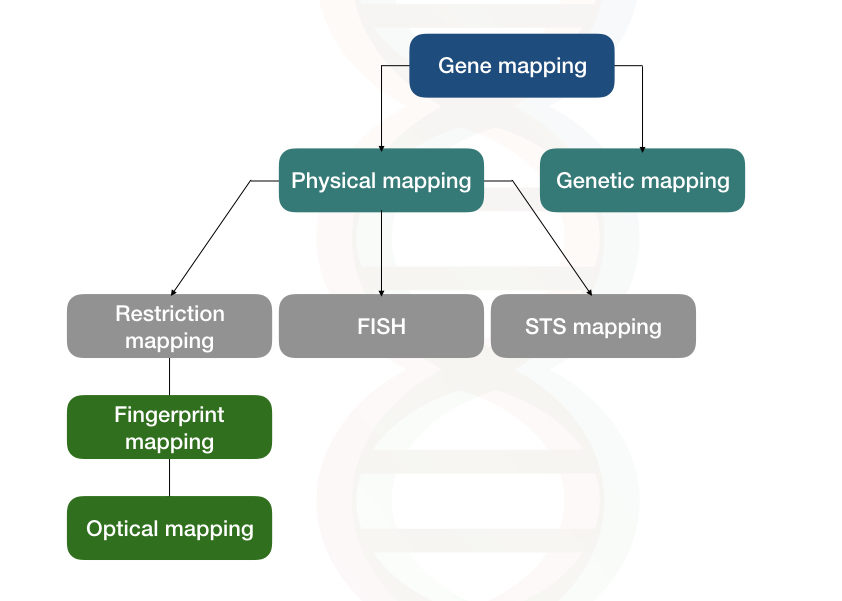
Two types of genome mapping/gene techniques are generally used by scientists; Genetic mapping and physical mapping.
Related article: 3 Of The Best Genome Sequencing Methods.
Key Topics:
Physical mapping
In the physical mapping, the exact location of a gene or a disease gene or a DNA sequence we wish to examine can be determined.
Three types of physical mapping techniques are used so often:
- Restriction mapping
- FISH mapping
- STS mapping
1. Restriction mapping:
Using the specific set of restriction enzymes one can create a genetic map. The method or genetic marker called RFLP, restriction fragment length polymorphism is used to do so.
However, the present mapping method is limited to the polymorphic restriction sites only.
In the genome, very few sites are highly polymorphic, thus it is very difficult to construct an entire map of our genome using RFLP.
The unknown DNA fragments can be mapped using the restriction endonuclease, subsequently, the digested DNA fragments are separated depending on their size using gel electrophoresis.
The restriction mapping generated two types of fragments; sticky ends or blunt ends, both types of restriction digestion is shown in the figure below,
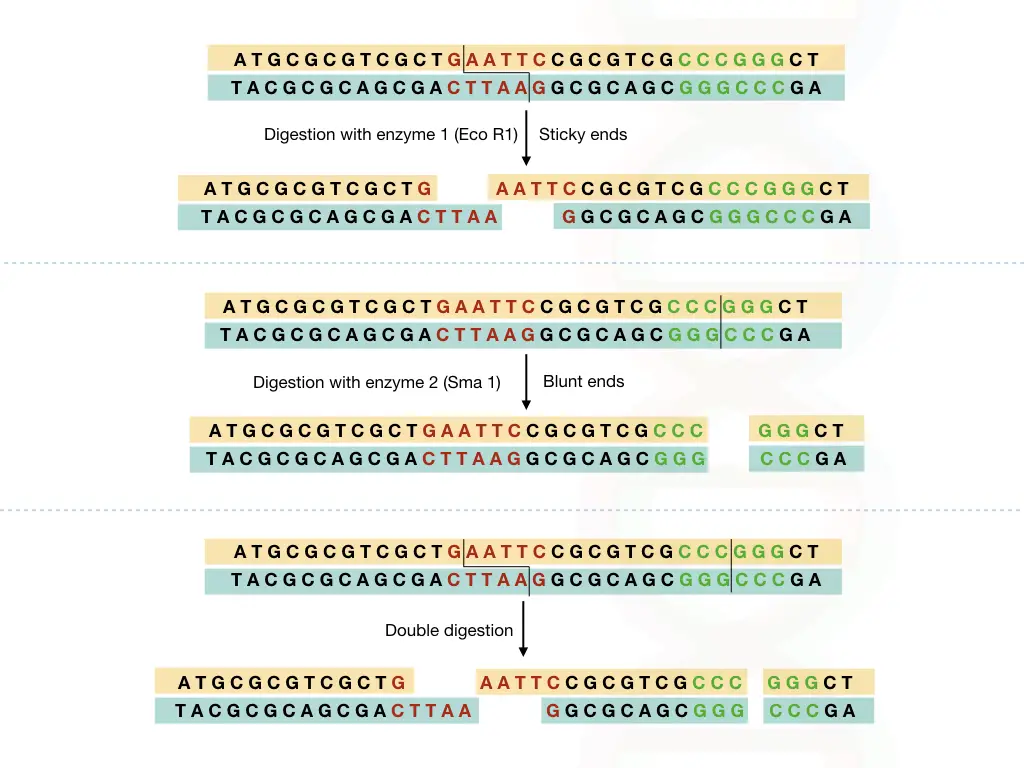
The DNA sample is extracted using the DNA extraction methods.
Using two different restriction endonucleases, the DNA sample is digested in three ways, that prepare three different aliquots of the sample.
Digest one aliquots with one restriction enzyme, second aliquots with seconds enzyme and double digest the third aliquots (See the above image).
After the digestion, each DNA fragment are separated based on their respective sizes, on agarose gel electrophoresis or PAGE (you can use any of the electrophoresis methods).
The restriction mapping method is used for the mapping of genes on a plasmid or shorter piece of DNA although mapping the entire genome using the restriction digestion or restriction method is a tedious, inaccurate and time-consuming method.
The restriction enzyme cuts DNA at its specific location of a short sequence called the restriction site.
Some of the restriction sites of some endonucleases are given in the table below,
| Restriction endonuclease | Recognition site |
| Eco R1 | 5’-G*AATTC-3’
3’-CTTAA*G-5’ |
| Bam H1 | 5’-G*GATCC-3’
3’-CCTAG*G-5’ |
| SAl I | 5’-G*TCGAC-3’
3’-CAGCT*C-5’ |
| Hae III | 5’-GG*CC-3’
3’-CC*GG-5’ |
| Hind III | 5’A*AGCTT-3’
3’-TTCGA*A-5’ |
| Sma I | 5’CCC*GGG-3’
3’GGG*CCC-5’ |
* indicates the cutting site.
The restriction map indicates or locates all the restriction sites from the entire genome or from the piece of DNA.
The restriction mapping method is further divided into two methods:
- Fingerprint gene mapping
- Optical gene mapping
Fingerprint gene mapping:
It is impossible to digest the entire genome with even one single restriction endonuclease because we can not distinguish all the fragments in a gel.
The EcoR1 has a 6bp recognition sequence which means if we digest the entire genome with only the EcoR1, it produces almost 80,0000 fragments. And those fragments are not distinguishable on a gel.
So in the very first step, the genome is broken into smaller fragments or a selected larger gene or DNA fragment is fragmented and inserted into bacteria plasmid. Clones of DNA fragments are generated in bacterial vectors.
In the next step,
These DNA fragments are digested with specific RE and run on the agarose gel. In the agarose gel, it is separated based on its size.
The resulting fragment size is analyzed and a genetic map of the restriction site can be constructed based on the similarity and differences between the band patterns.
See the image below.
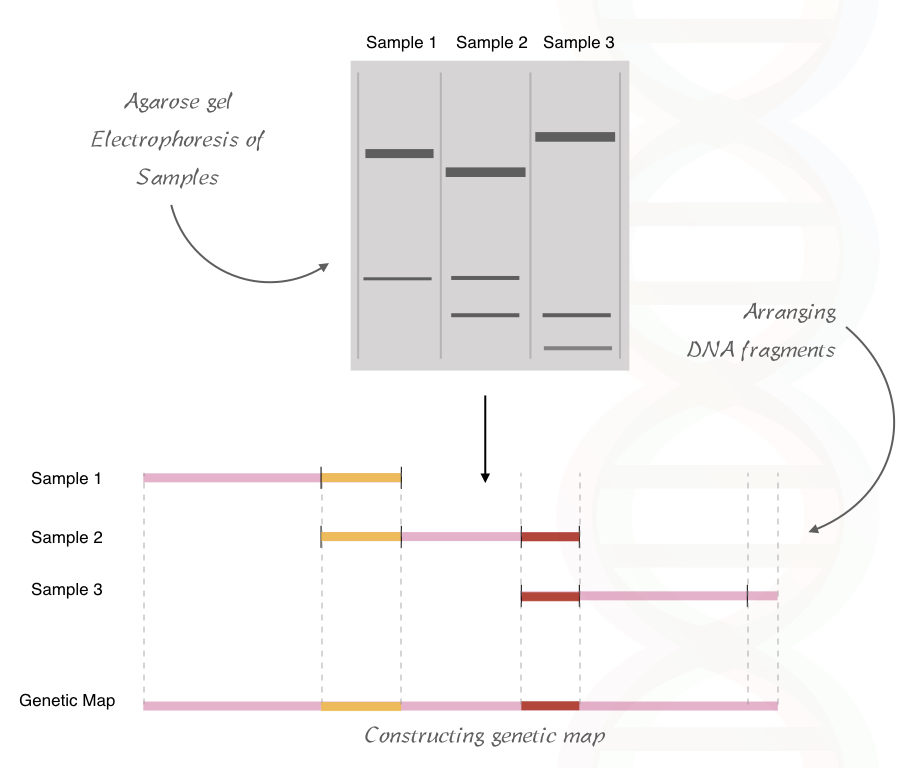
Optical gene mapping:
The optical map is constructed using the fluorescent intensity emitted by the gaps generated during the digestion.
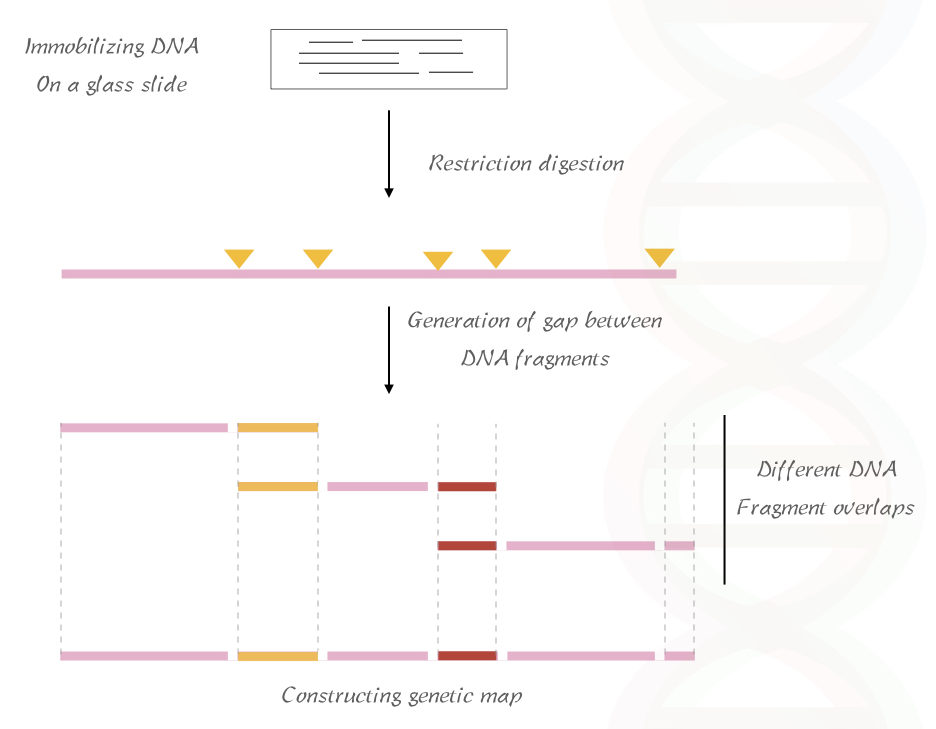
In the first step, the DNA fragment is immobilized on a glass slide and subsequently digested with the RE which generates a gap between the digested fragments.
Then the slide containing the DNA fragment is stained using the fluorescent dye and observed under the microscope (fluorescent microscope).
A brief overview of the method is illustrated in the figure below,
Based on the fragment size and gaps the map of the DNA is being constructed.
Related article:
2. FISH:
The fluorescent in situ hybridization method is used to detect a DNA sequence or the disease gene within a cell using the fluorescent probe.
The physical gene mapping methods rely on sequence information.
In FISH, a short sequence of DNA complementary to our DNA sequence or gene is artificially synthesized and labeled with the fluorescent dye.
The resulting fluoro-labeled, short oligonucleotide sequence called the probe is used in the hybridization.
If the complementary sequence of the gene of our interest is present on a chromosome, the probe will hybridize with it and give the fluorescent signal.
The resulting hybridization is visible under the fluorescent microscope because the probe directly hybridizes the DNA sequence on a chromosome.
Multiple DNA sequences or genes can be hybridized using multiple probes, the method is called mFISH.
The fluorescent hybridization signal allows us to determine the location of a DNA sequence or a gene on a chromosome.
See the image below,
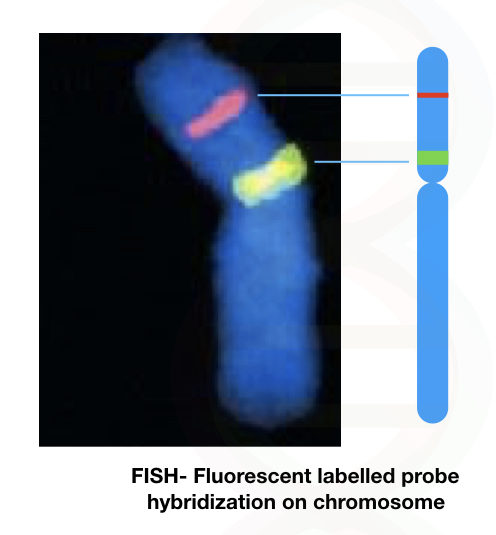
Many disease-causing loci are identified using the FISH and probes related to this are now commercially available.
Read our article on FISH: A Brief Introduction To Fluorescence In Situ Hybridization (FISH).
3. STS mapping
A Sequence-tagged site is a short stretch of the repeated DNA sequences of 100 to 500bp repeatedly present within a genome.
The STS are highly polymorphic regions thus they can easily be distinguishable.
The STS-based gene mapping technique relies on PCR and agarose gel electrophoresis.
The DNA sample is fragmented using the RE and inserted into the bacterial plasmid.
Once enough numbers of DNA copies are obtained, the fragments are collected and processed for PCR.
The DNA sample was amplified using the known primers specific to the STS in the polymerase chain reaction.
The PCR amplicons are run on 2% agarose gel electrophoresis.
From the results,
If two different STS markers are present on one DNA fragment, both must be nearer to one another in the genome.
Similarly, if two different DNA fragments contain the same STS, those two DNA fragments must represent the overlapping part of the genome.
Related article: Polymerase Chain Reaction.
Genetic mapping:
The gene genetic mapping technique is entirely different than the physical mapping method. The genetic mapping method is based on the number of markers used and the size of the study population.
The genetic mapping method is dependent on the principle of showing the positions of related genes or DNA sequences related to the phenotype or disease on a chromosome using the techniques such as pedigree analysis or cross-hybridization.
(Instead, in the physical mapping, the DNA sequence related to the disease or phenotype is directly examined using molecular genetic techniques).
Thus the genetic map constructed using the genetic mapping shows the relative position of a particular phenotype on a chromosome or genome.
For constructing a genetic map, scientists collect the blood samples from the family members having a disease trait and other family members without the disease.
After that, DNA extraction is performed from the collected samples, scientists then analyze the DNA of the family members having a disease trait as well as members who do not contain the disease trait.
The results are developed as a marker for that particular trait or disease.
The genetic map indicates the location of a particular phenotype-related genotype on a chromosome.
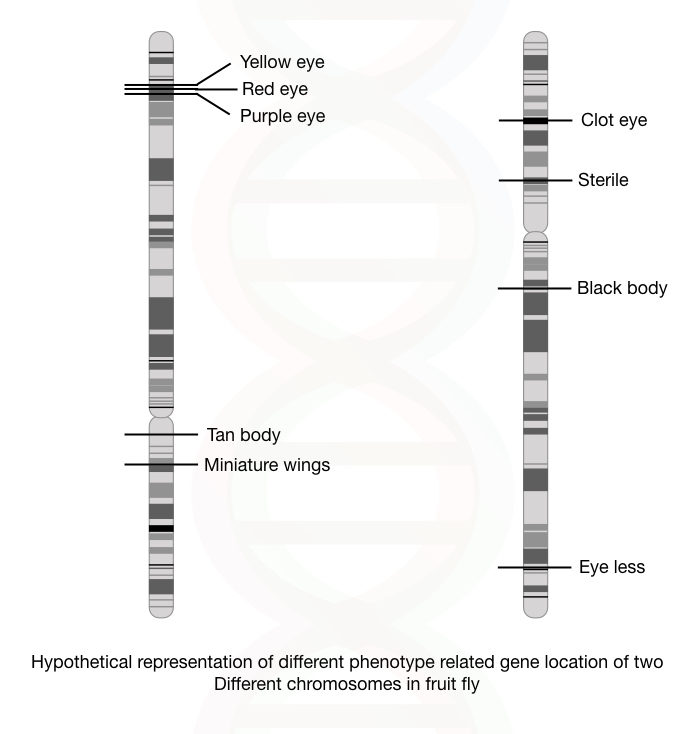
In the early days, the genetic map is used only for the visible or distinguishable characters like eye color, body color, height and wing shape, etc in fruit-fly.
Now, genetic mapping is used for visible as well as biochemical characteristics and the disease-causing gene can be mapped on a chromosome.
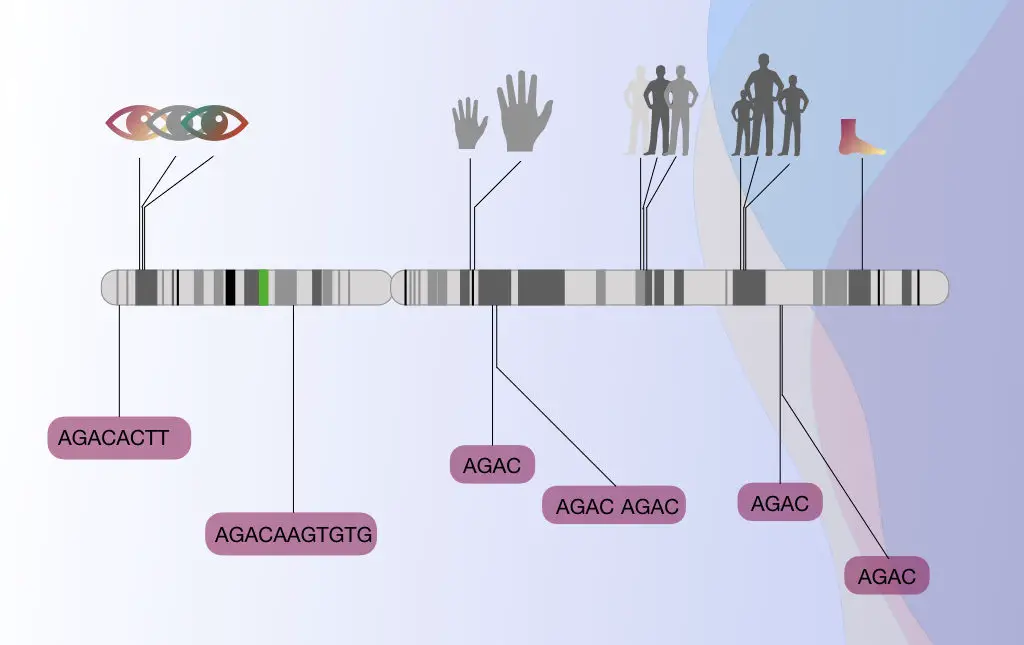
STR, short tandem repeats is one of the useful microsatellite markers used in the genetic marker technique.
Besides this, several other markers such as SSCP, VNTR, SNP, RFLP and AFLP are also used in genetic or gene mapping.
The STR (microsatellites) are frequently used in genetic mapping because the minisatellites are more prevalently present on the telomeric regions of chromosomes thus an accurate mapping can not be done.
Related article: Genetics Basics: A Beginners Guide To Learn Genetics.
Applications of gene mapping:
One of the important applications of gene mapping/genome mapping or genetic mapping is of identification of genes responsible for the traits.
(Trait is a distinguishable or observable phenotype of one).
For example the mapping of the disease-resistant genes in a plant genome.
It is also used in the identification of quantitative trait loci which are economically important.
Mapping of a milk-producing gene in animals can also be possible using gene mapping.
In modern genetics, identification of the disease-causing gene in the human genome is one of the important applications of gene mapping.
The heritable, as well as non-heritable, cancer-like disease-causing genes, can be mapped too.
Conclusion:
Advancements in DNA sequencing led to discoveries of more than 20,000 genes until now. The human genome project was completed in the year 2003 which sequenced the entire genome of us.
Gene mapping becoming a very crucial process for the identification of disease causes genes. The location of it, inheritance pattern and penetrance of the allele can be determined by using the data of gene mapping.
Sources:
- Brown TA. Genomes. 2nd edition. Oxford: Wiley-Liss; 2002. Chapter 5, Mapping Genomes. Available from: https://www.ncbi.nlm.nih.gov/books/NBK21116/.
- Restriction Mapping. Nature Masterclasses.
- Genetic mapping factsheet by NIH.


It’s a niece piece especially for beginners in genomics