Genotyping is a process of investigating a variant(s) present in an individual while sequencing is a process of investigating the whole sequence, gene or genome.
Genotyping and sequencing are separate techniques having a different work platform, the principle of working and applications. This article gonna be an elaborative explanation with technical clarity.
So if you are a beginner and wish to learn more regarding each separate term, please read these articles first.
Without wasting much time, let’s quickly discuss some of the common differences, technical foundations and applications of genotyping and DNA sequencing.
Key Topics:
What is genotyping?
A technique that enables us to detect various genotypes present in a gene, DNA sequence, genome, or in a person or in an organism is known as genotyping. For example, FISH and DNA microarray.
A genotype is a genetic constituent that produces a specific phenotype through mutation or CNV for a gene. Commonly, the genotyping technique relies on the location of a genotype and genotype (mutation) itself. For example, IVS1-5 (G/T).
Meaning, it locates and identifies known variants for a gene or a person. These two parameters are usually known.
FISH (Fluorescence in situ hybridization) is a well-established genotyping technique based on the principle of DNA hybridization. A probe specific to known (or previously identified) genotype, is designed and allowed to hybridize at a target sequence.
As the probes are highly sequence-specific, it only binds to complementary region, viz at a genotype. Through fluorescence analysis, the region is located and studied.
Another technique, DNA microarray also works on the same platform but is powerful enough to investigate millions of genotypes in a single assay.
Techniques of genotyping are used to establish, investigate and study the genetic basis of polygenic disorders.
What is sequencing?
A technique that enables us to identify or study a whole sequence is known as sequencing. DNA sequencing, gene sequencing and genome sequencing are three broad categories that study a sequence of DNA, gene or entire genome of an organism.
Gene sequencing, exome sequencing and genome sequencing are some of the examples.
In a broad sense, in principle, a DNA primer binds and amplifies in a reaction and expands the sequence which is read. The ‘read’ identifies each base of a sequence.
Differences between genotyping vs sequencing:
The Helix blog amazingly defined genotyping and sequencing terms as,
“Genotyping is like reading some known words, randomly on a page or book while sequencing is like reading the whole sentence or book.”
Helix blog.
Genotyping is a process to investigate or determine variation in a gene or individual while sequencing is a process to study or investigate the whole sequence or gene.
Genotyping studies ‘change’ within the sequence which is known while sequencing studies ‘the whole sequence’ of interest.
Genotyping techniques need previous genotyping data or sequence information for the study, while sequencing doesn’t need any previous data for a sequence.
Henceforth, genotyping can’t find new variations, however, sequencing has the power to discover new variations or genotypes.
On a common note, genotyping assays are based on the principle of DNA hybridization, for instance, FISH and DNA microarray; whilst sequencing assays are based on the principle of DNA amplification, for instance, Sanger-sequencing or whole-genome sequencing.
Note that PCR amplification is also a type of genotyping that uses the principle of amplification.
Genotyping provides us less data that can be processed faster while sequencing provides us a huge data that needs further computational investigations to study a sequence and find a new variation.
Genotyping techniques are recommended for population-based studies, SNP studies and polygenic studies; A genetic basis behind complex disorders can be studied, like diabetes or Autism Spectrum Disorders.
On the other side, sequencing-based techniques are used in diagnostics, investigating sequence variations, finding new mutation or copy number variation for a disease or gene.
In addition, techniques like whole-genome sequencing are used in GWAS and Exome sequencing for whole-exome analysis.
Though both techniques are costly, sequencing platforms especially, whole-genome and exome sequencing are more costly than genotyping techniques.
Note
Sequencing platforms can determine known genotypes as well as find a new one.
Technologies in genotyping study sequence variation (obviously which is known), compare it with other person or population and find the prevalence of particular genotype in a population.
By doing so, the genetic susceptibility of a particular genotype can be determined, also it can determine the population risk for that particular genotype.
On contrary, technology in sequencing study or can discover new sequence variations and investigate their association with disease conditions.
By doing so, the genetic basis for why a genetic defect occurred, can be determined. However, more studies will be needed for validation and establishment.
| Techniques in genotyping | Techniques in sequencing |
| FISH | DNA sequencing |
| SNP genotyping | Gene sequencing |
| CNV genotyping | Exome sequencing |
| Whole-Chromosome microarray | Whole-genome sequencing |
| Single chromosome microarray | Next-generation sequencing |
| RFLP, RAPD, AFLP and PCR | Pyrosequencing |
Note that PCR-based amplification is also a type of genotyping technique, however, it has DNA amplification base.
Read more: What is Whole-Exome Sequencing?
| Genotyping | Sequencing | |
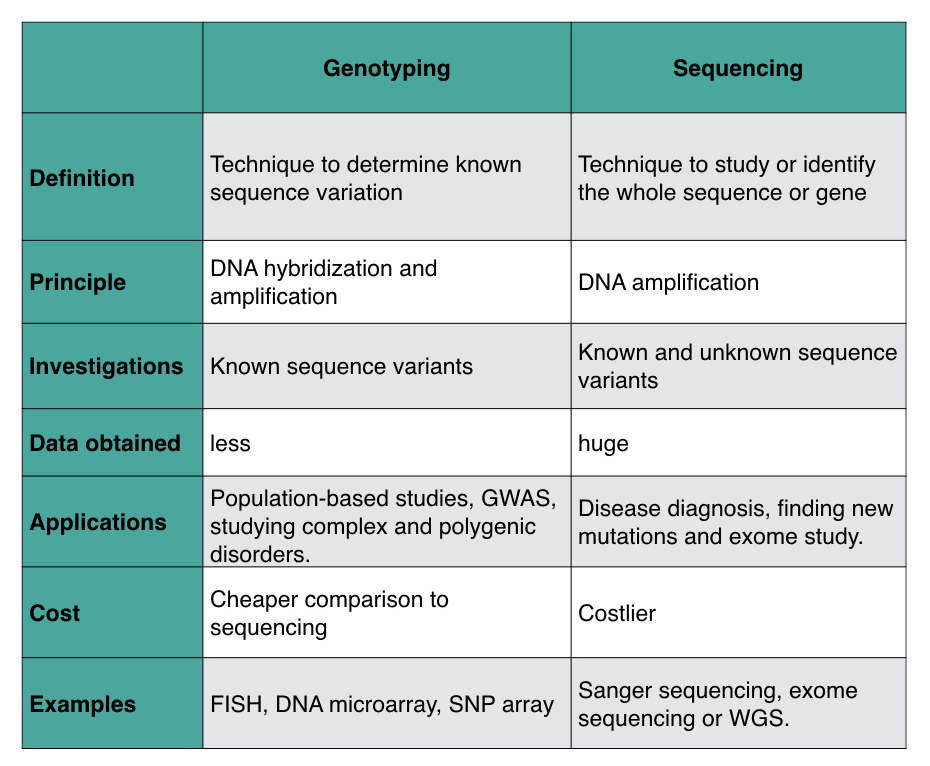
| Definition | Technique to determine known sequence variation | Technique to study or identify the whole sequence or gene |
| Principle | DNA hybridization and amplification | DNA amplification |
| Investigations | Known sequence variants | Known and unknown sequence variants |
| Data obtained | less | huge |
| Applications | Population-based studies, GWAS, studying complex and polygenic disorders. | Disease diagnostic, finding new mutation and exome study. |
| Cost | Cheaper comparison to sequencing | Costlier |
| Examples | FISH, DNA microarray, SNP array | Sanger sequencing, exome sequencing or WGS. |
Wrapping up:
Genotyping has been a valuable tool for population-based studies. It can find the prevalence of a specific genotype in a population, high-risk group and its dependency to cause disease.
However, on the other side, sequence though provides more powerful and vast data, it can’t directly be used for diagnostics.