“DNA fragmentation is a technique of artificial breaking of DNA in order to use it in various downstream genetics and well as genomics applications.”
The first DNA fragmentation was reported by Williamson in 1970 in a cell in vivo. Albeit, If DNA fragmentation occurs in a cell, it can cause serious health-related problems, majorly, cancer.
Some intrinsic, as well as extrinsic factors, do DNA fragmentation. The technique known as the COMET assay is widely used to study and investigate the amount of DNA fragmentation in a cell.
The present technique is also used to encounter male infertility by studying the amount of fragmented DNA from a sperm cell.
But here, our purpose of making DNA fragments is different!
We can use DNA fragmentation to use gDNA, cDNA and various other DNAs in different experiments.
In the present article, we are going to discuss the artificial method of DNA fragmentation and why we practice it.
Key Topics:
Importance of DNA fragmentation:
The genome of an organism is a complex network of DNA and protein and it’s huge. For instance, the human genome contains around 3 billion DNA base pairs.
On the other side, a gene is a tiny unit of the entire genome that makes a particular protein.
Approximately, 23,000 genes are now known to us. Processing the entire genome in order to study a single gene or some genes is worthless!
Also, we can’t process the whole genome in a single experiment and we can’t interpret its results as well, perhaps. The reason is, it’s huge!
None of the machines can sequence or amplify the entire genome in a single run. So what can we do to study the whole genome?
The concept of DNA fragmentation facilitates to overcome this problem. If we somehow fragment the whole genome into smaller fragments that can be processed, then we can study it.
There are some reasons we can’t use the whole genome in sequencing, microarray and PCR.
- First, the genome is huge.
- Second, it’s very complex
- Third, there are so many repetitive sequences in it that are similar to some genes.
- Fourth, the major portion of the genome is non-coding and made up of high GC rich nucleotides. GC rich regions are hard to amplify.
Other reasons why we can’t use the whole-genome are the limitations of the machine.
- A PCR can efficiently amplify up to 5000bp DNA fragments, not the whole genome.
- The DNA sequencer machine can sequence up to a few thousand base pairs.
- Due to repetitive DNA’s non-complementary base pairing in PCR, microarray and DNA sequencing occurs if we use the whole genome.
Henceforth, if we want to study a gene or even the entire genome, we have to fragment it.
We need to make processable fragments of DNA that can be amplified, sequenced and hybridized efficiently in PCR, DNA sequencing and DNA microarray, respectively.
Restriction digestion, PCR amplification, sonication, heating and chemical methods are some techniques utilized to cleave DNAs. Based on that the DNA fragmentation methods are divided into three categories.
- Enzymatic methods
- Physical methods
- Chemical methods
Techniques of DNA fragmentation:
Enzyme based DNA fragmentation methods:
Some of the enzymes have the power to cleave DNA and are utilized to fragment DNA.
Restriction digestion:
The restriction digestion method is also known as enzymatic lysis which can cleave or fragment DNA at a specific location or known location.
The restriction enzyme brakes either single-stranded or double-stranded DNA precisely and at a known location. We can also create a restriction digestion map to use it after processing, to arrange the sequences of fragments.
The restriction digestion cuts the DNA and creates either blunt-end DNA or sticky end DNA.
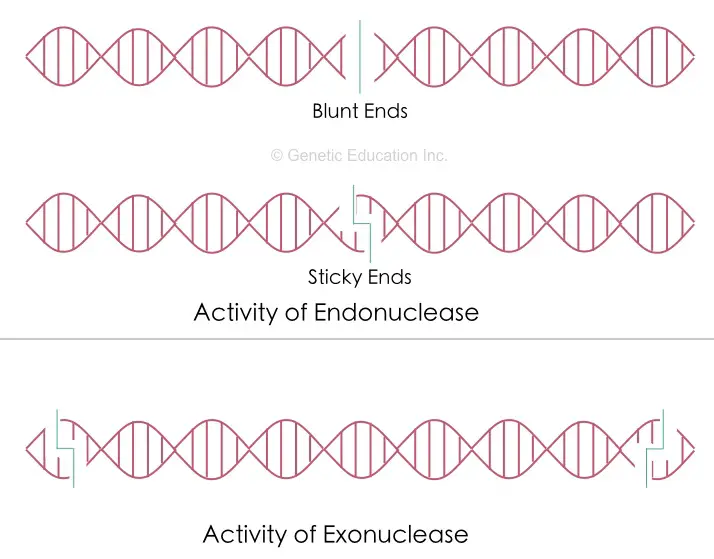
Non-specific nucleases:
The nuclease is a class of enzymes that can cleave DNA and can generate DNA fragments, the well-known example is the endonuclease. However, some other non-specific nucleases can also cleave DNA but at a random site and generate DNA fragments of random sizes.
Physical shearing:
The physical methods of DNA fragmentation are commonly used in next-generation sequencing and other sequencing techniques. Different physical shearing methods are capable of making different sized DNA fragments of the genome.
For instance, the nebulizer can make DNA fragments of 100 to 3Kb using the atomized liquid and compressed air.
The acoustic shearing method is capable of making DNA fragments of 100 to 5Kb while the sonication technique can generate DNA fragments of 100 to 1Kb.
Heat denaturation:
Yet another physical method to shear DNA is the extreme heat! Under high temperature, the DNA strands break into fragments. Nonetheless, the heat shearing method is less effective and hence not used in library preparation of next-generation sequencing.
PCR-based amplification:
The PCR based amplification of a gene of our interest or the DNA fragment we wish to study can be separate or we can say fragmented.
Based on the location of a gene in the genome the primer set is designed, amplified and separated on the agarose gel electrophoresis.
The separated fragment of DNA is then isolated and sent for either DNA sequencing or restriction digestion.
The PCR based amplification method is one of the most trusted DNA fragmentation methods so far.
Also, the random amplification in order to get different fragments of the whole-genome is performed to create the entire librate of the DNA fragments for DNA sequencing and DNA microarray.
Chemical methods:
Chemicals like the alkaline agents can break DNA in pieces. And hence recommended by some of the manufacturers in DNA fragmentation. Although chemicals are less effective than the restriction digestion method.
Related article: What is DNA sequencing and how to do it?
Besides several other fragmentation techniques are discussed here.
Centrifugal shearing in which the sample DNA passed from the specific size pores under the high centrifugal forces. Larger size DNA can’t pass from and break under the influence of centrifugal forces.
Needle-shearing is yet another technique used to do so in which the DNA is forced to pass through the needle which generates the fragment of DNA up to a few Kb in size.
Applications of DNA fragmentation:
To perform DNA sequencing:
The main application of DNA fragmentation is to facilitate smaller DNA fragments to sequence. Larger DNA makes it difficult for a machine to sequence and amplify. Thus we need to fragment DNA in sequencing.
A library of different DNA or RNA fragments is sequenced one after another in high-throughput DNA sequencing.
To perform it, the genome is first fragmented using the technique enlisted above. Then in the next step, it is amplified in the PCR. The PCR amplification generated copies of fragments.
After that, each fragment are ligated with the known adapter sequences to generate the library of DNA fragments.
The adapter sequencer is known to us therefore it’s a kind of tag used to evaluate the results. After the completion of DNA sequencing, we will locate each adapter sequence, remove it and join each fragment computationally.
To perform DNA microarray:
Again much like DNA sequencing, DNA fragmentation and library preparation are the common steps in DNA Microarray as well.
The fragments are allowed to hybridize on a glass slide where millions of sequence-specific probes are located.
The probes are known as to us and by their hybridization, we get information about the sequence.
SNP microarray, whole-genome microarray and whole-chromosome microarray are popular nowadays to hybridize whole genomes.
Related article: DNA microarray.
Conclusion:
The objective of DNA sequencing or DNA microarray is to screen various mutations associated with DNA.
However, the microarray can’t find new mutations or copy numbers variations but a sequencer can! The process of DNA fragmentation and library preparation facilitate ease in the experiment and reduce complexity.
Sources:
Keisuke Tanaka, Rumi Ohtake, Saki Yoshida, and Takashi Shinohara. Effective DNA fragmentation technique for simple sequence repeat detection with a microsatellite-enriched library and high-throughput sequencing. BioTechniques 2017 62:4, 180-182.


