The “kit of housekeeping genes” are required for basic cell function, in all cells in all organisms.
Genes are one of the important elements of our genome, in fact, a must required component which encodes proteins. And that is the main function of the genes.
A genome consist of coding and non-coding DNAs helps in encoding proteins and maintains gene regulation, respectively (Genome).
However, the expression of a gene or set of genes is tissue-specific- some cells required a higher amount of some genes while some not.
For example, a high amount of keratin protein-encoding gene required in hair cells but not in muscle cells.
A hemoglobin encoding gene is required in blood cells, but not in hair cells that is how the entire mechanism is working, coordinately.
Notably, some genes called “housekeeping genes” are required in all cells for the basal functioning of a cell. In the present article, we will give you a complete guide on how housekeeping genes are working and what is their significance for us.
Related reads:
Key Topics:
What are housekeeping genes?
So let’s start with some basics,
Replication, transcription and translation is a machinery used in the process called “cell central dogma” from which a DNA copied, mRNA is formed and a protein is translated, respectively.
DNA polymerase is the main ingredient for the replication which synthesizing the DNA by taking the help of helicase, topoisomerase and primase.
The RNA polymerase is the main enzyme used in the transcription which also gets the help of other transcriptional proteins and factors as well for constructing the mRNA.
tRNA and rRNA are the major players of translation helps to encode a protein.
All the elements are must require in each cell for creating a protein thus we can say all these elements tRNA, rRNA, transcriptional factors and other enzymes must be produced in every cell for performing basic functions.
Genes which encodes for these elements or factor are our housekeeping genes and those required in every cell- for keeping our house (a cell) safe. ?
Now take a look at this example:
The ATF1 gene is a known transcriptional factor required to regulate the transcription and encodes a protein Cyclic AMP-dependent transcription factor. This gene belongs to the ATF family which regulates the expression of downstream genes.
The gene for ATF1 protein is highly conserved since evolution, present in all eukaryotes and must require during transcription.
For constructing mRNA transcript of any gene of any tissue, ATF1 gene is an important element of the “transcription kit”
Therefore in each cell, the ATF1 gene must be produced in a constant and same amount because it has to perform the basic function- the transcription.
And that is why the ATF1 gene and the family of ATF genes are categorized into the housekeeping genes.
Now you better understand what the housekeeping genes are!
16s rRNA is another important component of prokaryotic 30s ribosome sub-unit which helps in immobilizing and interacting of ribosomal protein for performing the translation. 16s rRNA is also a class of housekeeping genes.
Definition:
“Highly conserved regions of a genome possessed tissue-specific genes which required for the basal function of cell machinery is called housekeeping genes.”
In other words, we can say,
“Genes required for cell maintenance under normal as well as pathophysiological conditions are called housekeeping genes.”
The housekeeping genes have some diverge characteristics which make it different from other genes, some of them are enlisted here:
The housekeeping genes are simple in structure, made up of shorter exon and introns. Exon and introns are sequences of a gene, during the splicing the non-coding introns are removed and the final transcript is formed.
Due to the shorter sequences, processing and splicing performed easily and preciously.
Highly conserved:
The housekeeping genes evolved slowly because it has to perform the same function in all cell types in all organism thus no need to evolve or change. It is highly conserved in almost all organisms and therefore is less polymorphic in nature.
The repeated DNA sequences are very less and shorter in housekeeping genes. The major repeat DNA in housekeeping genes is short interspersed repeats and lacks longer repeats.
The 5’ untranslated is yet very simple made up of simple sequence repeat.
Because of this reason, the nucleosome formation capacity (during DNA packaging) is very low, due to the simple sequences on 5’ untranslated region.
It is constitutive in nature- thus transcribed continuously in all cells and in all organisms. As opposite, facultative genes which transcribed when needed, the constitutive nature of housekeeping genes makes it available during transcription and translation all the time when it needed.
Housekeeping genes are less complex and do not contain larger and longer exons as well as long and complex repeat sequences.
The promoters of housekeeping genes do not contain TATA-box.
In addition to this to make it conserved and unchanged, the genes are enriched with GC content and thus have high CpG islands.
Unlike the promoter of the normal genes, the GC boxes are present instead of TATA and CAAT boxes in housekeeping genes.
Housekeeping genes are categorised in “all-time active genes” – vital elements needed to survive.
List of some common type of housekeeping genes used in the realtime PCR are given below,
| Gene | Gene name | Level of expression |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | Hight |
| RER1 | Retention in endoplasmic reticulum 1 protein | low |
| PPIA | Peptidyl prolyl isomerase A | Medium |
| RPL41 | Ribosomal protein L41 | Medium |
| RN18S | Ribosomal 18s rRNA | High |
| HPRT1 | Hypoxanthine-guanine phosphoribosyltransferase | Low |
| ACTB | Beta-actin | Medium |
| ACTA | Alpha actin | Medium |
| ALDOA | Aldolase A, fructose- bisphosphate | high |
| RPL7L1 | Ribosomal protein L7- like 1 | low |
These are some of the common housekeeping genes used in the real-time PCR quantification, besides these other common housekeeping genes are enlisted at the end of this article.
Use of housekeeping genes in RT-PCR:
Because we have smarter minds, we are scientists and us obviously different. We know how to use positive characteristics of something for betterment.
Scientists are using housekeeping genes as a reference or marker in RT-PCR experiments for quantification, why? Because as we said their expression is stable and relatively unchanged in all cells and tissues, across all organisms.
Due to this reason, it is used in mRNA relative quantification for normalizing data.
18s rRNA, ACTN, tubulin, GAPDH and PGK are some of the common housekeeping genes used in the realtime PCR quantification.
Quicklook on RT-PCR:
For mRNA quantification, the cDNA is constructed from the sample and used for reverse transcription.
Using the reverse transcriptase enzyme, the mRNA is synthesized from the cDNA and used for quantification.
Using the fluorescent dye or fluorochromes, the relative and absolute quantification of mRNA can be done in realtime PCR.
A standard curve is generated for absolute quantification while the housekeeping gene or reference sample is used in relative quantification.
Ultimately, the amount of fluorescence emitted by hybridization is measured for the quantification of the template mRNA.
Using the housekeeping genes in our RT- PCR assay provides a reference value to estimate the concentration of the mRNA present in a sample because of the stable expression of it in all cells.
Though it is used for quantification of different mRNA, it is not suitable for all type of assays because not all the housekeeping genes have stable expression in all cell types, yes, it is fact (scientific data suggest).
The minor deviation is reported all the time.
And therefore, we need to perform normalization all the time for calculating different templates.
Read more: Reverse transcription PCR: Principle, Procedure, Applications, Advantages and Disadvantages.
Take a look at the example here:
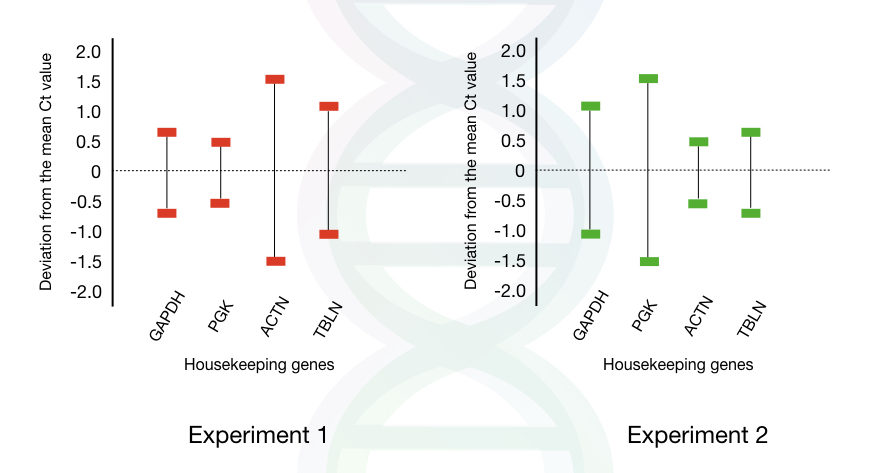
From the experiment, you can see that the deviation in the standard value of GAPDH and PGK is very low in experiment 1 thus both housekeeping genes are the best choice for experiment 1.
While the ACTN and TBLN are the best choices for experiment 2 due to the lower deviation rate.
This is the reason normalization must be performed before doing RT- PCR for different cell types.
Conclusively, we can say due to the nearly unchanged expression of housekeeping genes, it is used as a reference for mRNA quantification in real-time PCR.
List of other common housekeeping genes:
Transcriptional factors:
- ATF family genes – Activating transcriptional factors
- HMGB1- high mobility group box binding DNA
- ILF2- (H. Sapiens) interleukin enhancer-binding factor 2
Translational factors:
- EIF family genes- eukaryotic translational initiation factor
- TUFM- Tu translational elongation factor mitochondrial
Repression:
- PUF60- Poly (U) binding splicing factor.
RNA binding protein:
- ELAVL- ELAV like protein 1
RNA splicing:
- PABPN1- poly (A) binding protein, nuclear 1
- HNRPK- H. Sapiens heterogeneous nuclear ribonucleoprotein K, transcript
tRNA synthesis:
- AARS- aminoacyl tRNA synthetase
- DARS- Aspartate tRNA synthetase
- FARS- phenylalanine tRNA synthetase
- IARS- isoleucyl-tRNA synthetase
- TARS- threonyl-tRNA synthetase
Ribosomal proteins:
- RPL family genes- 60s ribosomal protein (RPL5, 8, 9, 10A, 11, 14, 25, 27, 30, 32, 34, 35, 36)
- RPS- 40s ribosomal protein (RPS20, 23, 24, 27).
Heat shock proteins
Histone
RNA polymerase
Cell cycle protein
Conclusion:
The set of housekeeping genes are practiced as an internal control to improve the reliability of the quantification reaction. Nearly accurate quantification of mRNA can be done, however, normalization of housekeeping genes must be required before it is used in the realtime PCR to improve the efficiency of the reaction.
Sources:
Kozera B, Rapacz M. Reference genes in real-time PCR. J Appl Genet. 2013;54(4):391–406. doi:10.1007/s13353-013-0173-x.