“Numerous genetic testing techniques and assays are now available, however, selecting one for your experiment is a crucial decision. This article provides a comprehensive guide for various genetic methods and how to choose one, depending on your requirement.”
Since the DNA was chemically explained by Watson and Crick (1953) many basic and sophisticated genetic techniques evolved timely. Some were considered excellent in the past but are least useful now while some are still important in daily research.
In genetic research and diagnosis, assay selection is a crucial decision for getting accurate, trustable and meaningful results. Having a few traditional, popular and important techniques on the board, a plethora of genetic assays are now available for various purposes.
Now, we have dedicated technology, techniques and assays for chromosomal studies, genetic studies, disease studies, phylogenetic studies, gene expression studies, population studies and even microbial genetic studies.
Genetic assays and techniques are now so useful that scientists can use them for disease diagnosis, treating infection and controlling outbreaks. However, each technique would come with its own advantages and limitations. And therefore, selection becomes so crucial.
The selection of genetic assay depends on the molecule you are working on, the expertise and experience you have and requirements of the assay, etc. Today, we will explore different types of genetic assays or techniques, their importance and their applications. Most importantly, I will let you know how to select one for your experiment.
This article will be a ‘sure-sure’ guide for researchers, although, if you are a student or newbie scientist, please read it, it will help you to design your experiment or at least to learn about different techniques.
Stay tuned.
Key Topics:
Importance of genetic assays:
Popular genetic techniques like PCR, restriction digestion, DNA sequencing or karyotyping are routinely used in genetic research for years. However, it has become increasingly important in medical science and healthcare.
Plenty of techniques are now used for the screening of genetic disease at the prenatal, postnatal and pre-implantation levels and are successful. Hence, starting from karyotyping to new-age Next-generation DNA sequencing, every technique now has been used extensively in the medical field.
Take a close look at various applications.
Extensive research: The foremost and priority application of genetic techniques is in research. It is used in plant, animal and microbial research to find new genes, and species, prepare taxonomical relations and understand various mechanisms and processes related to organisms.
They are, furthermore, used in genetic engineering, gene editing and recombinant DNA research and finding novel ways to treat ‘ailments.’
Diagnosis of genetic disease: Inherited as well as non-inherited single gene and polygenic diseases can now be diagnosed using genetic techniques. A single NGS or microarray assay is capable enough to find and screen millions of alterations associated with the disease.
This can also be done at the prenatal and preimplantation level so that the condition can be managed, ruled out or treated early.
Microbial genetic studies: Genetic techniques have increasingly become popular in microbiology and microbial research. Due to high-end safety, and the capacity to assess thousands of microbes in an assay, genetic assays like metagenomics analysis are extensively penetrated in microbial studies.
Learn more: What is Metagenomics?- Definition, Steps, Process and Applications
Prenatal testing: As aforementioned, in recent trends, genetic techniques become increasingly important in prenatal testing. A fetus’s genetic condition can be ruled out as early as possible, allowing us early intervention, treatment or management for the condition.
Population screening: ultra-modern genetic assays like NGS or microarrays are capable enough to perform accurate screening at the population level, for a particular or group of alleles. Thus, it gives immense strength to study a disease or alteration at the population level.
Predictive genetics: Advanced genetic technologies also have the capacity to predict any disease condition and associated genetic alterations. Millions of alterations associated with cancer and other diseases can be screened or predicted in a single assay.
Infection studies: Various infectious and related pathogens can be identified and quantified using techniques like real-time or qPCR.
Besides, other potential applications of modern-day genetic assays are disease risk assessment, outbreak screening and monitoring, pharmacogenetics, gene therapy research, rDNA technology and genetic engineering.
Types of genetic testing techniques or assay:
We already have so many articles explaining various genetic techniques, comprehensively. So in this section, I will only give a brief overview of each technique and also provide a link to the complete article, when required.
Karyotyping:
It is a conventional approach to studying chromosomes and is still widely used in the screening of chromosomal alterations. It studies chromosome number, structure and related alterations. Karyotyping is used in the diagnosis of chromosomal diseases like Down syndrome. Also, it is utilized in cancer cytogenetic studies as well.
FISH:
FISH is a Fluorescence In Situ hybridization which is a hybridization-based assay used to study chromosomal alterations which can’t be studied using karyotyping. It’s a type of high-resolution karyotyping where an individual target-specific probe binds on chromosomes and provides us with an idea about the alteration.
Chromosomal microarray analysis:
CMA (chromosomal microarray) has a higher resolution than conventional karyotyping and FISH and can screen thousands of indels (insertions and deletions) from all the chromosomes. The technique is more sensitive and powerful enough to find alterations associated with any genetic disease or cancer. It has been widely used in cancer genetic studies.
PCR:
PCR is a polymerase chain reaction, a technique scientists used to generate copies of DNA. PCR is extensively used in the study of mutations, single-gene disorders, DNA sequencing, gene analysis, marker-assisted selection, DNA testing and DNA fingerprinting.
qPCR:
qPCR is a quantitative PCR having crucial applications in gene expression studies, infection studies, microbial analysis and virology and bacteriological studies.
RT-PCR:
Reverse transcription PCR can reverse transcribe the mRNA into cDNA and allow us to study retroviruses, gene expression, infection and other transcriptome-based analyses.
Sanger sequencing:
Sanger sequencing is an important technique to study new genetic variations, identifying alterations associated with the disease and studying genes and sequences at a sequence level.
NGS:
NGS is next-generation sequencing, a powerful sequencing tool that can even sequence the entire genome of an organism. It can generate a huge amount of data and can screen millions of genomic alterations.
Thus, it is extensively utilized for disease studies, gene studies, genome level analysis, transcriptomics studies, genomic studies and pharmacogenomic analysis.
Related article: DNA Sequencing: History, Steps, Methods, Applications and Limitations.
Methylation assays:
Methylation assays like methylation PCR and methylation sequencing are popular ways to study gene expression and perform epigenetic analysis.
Restriction digestion:
Restriction digestion is a technique that enables us to study genetic alterations using restriction enzymes. It is also used to study genotypes and single-gene disorders.
Chromatin immunoprecipitation assay:
The chiP assay can study chromatin and helps in epigenetic analysis.
Single-cell RNA sequencing:
Single-cell RNA sequencing is a powerful sequencing platform that enables us to study the RNA, particularly the mRNA from even a single cell. It is widely used in gene expression studies.
How to select a genetic assay or technique?
Don’t worry if you are a beginner and don’t have any idea about where to start. I will stepwise explain the process for selection.
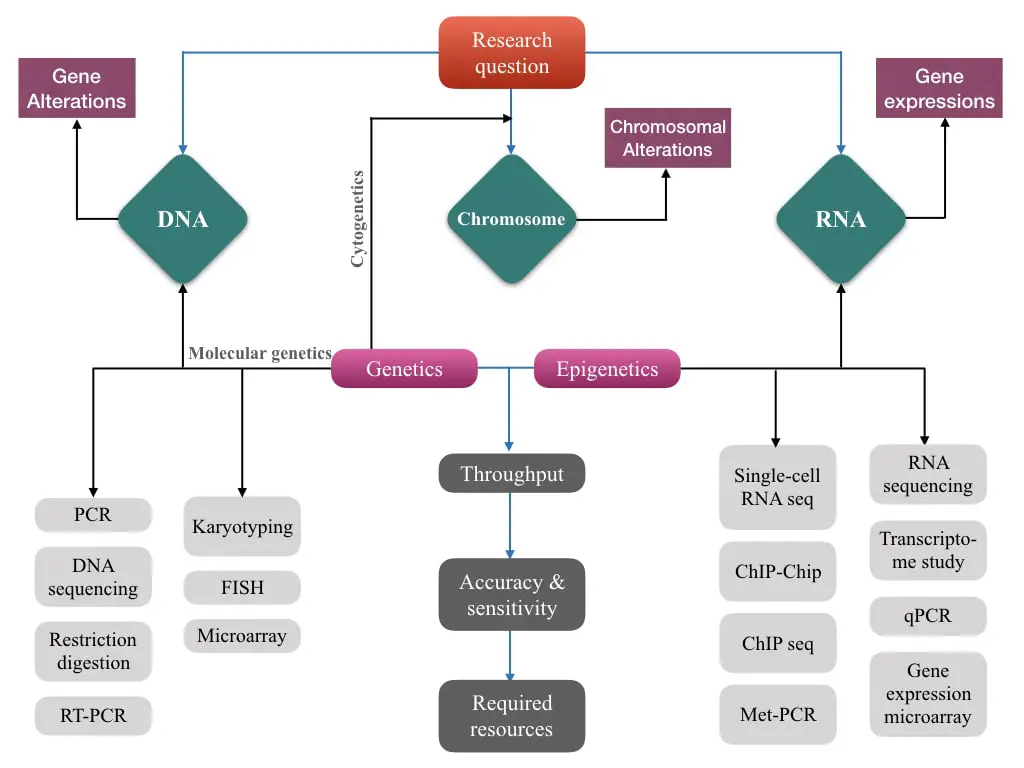
Step 1: Understand the Research question:
The very first step is, you have to select and understand your research question whether you are working on a disease, gene, sequence, mutation, condition, population, microbe or microbial community. Be clear on what you are working on and then go ahead. To understand it, carefully analyze your research question.
Step 2: Select the study molecule- DNA, RNA or chromosome:
Now closely take a call on whether you would like to work on DNA, RNA or chromosomes. You have to make this decision based on your interest, understanding and expertise. For example, if you are working on some cancer, you can work on chromosome, DNA and even RNA levels.
Step 3: Epigenetics vs genetics:
You also have to make a decision on what type of study you would like to conduct. You can perform either gene expression or gene alteration analysis and depending on that techniques may vary.
For example, PCR, restriction digestion and DNA sequencing are the techniques commonly employed for genetic studies while qPCR, RT-PCR, microarray and RNA sequencing are commonly employed for gene expression studies.
| Field | Techniques |
| Cytogenetics | Karyotyping, FISH and chromosome microarray |
| Molecular genetics | PCR, restriction digestion, southern blotting, DNA/RNA sequencing etc. |
| Epigenetics | RT-PCR, ChiP-seq, RNA sequencing, transcriptome sequencing, methylation assays, chromatin immunoprecipitation assays, DNA-protein interaction studies. |
Step 4: Consider the throughput:
Throughput is a crucial factor in choosing the right genetic assay. For example, PCR and DNA sequencing can perform mutational and gene analysis, but PCR may be well-suited for a small number of ‘well-studied’ samples and/or mutations.
Contrarily, sequencing techniques can study many sequence variations, entire genes or even genomes and give us ideas about novel variations. Thus NGS and Sanger-like sequencing platforms can analyze a large number of samples, mutations or genes in parallel.
Keep in a note, throughput is also important in cytogenetic and epigenetic studies too. Karyotyping can have a limited resolution and can study numerical and some larger chromosomal alterations, accurately. While FISH has higher resolution and can even study smaller alterations which karyotyping can’t.
| Technique | Low throughput | Medium throughput | High throughput |
| Cytogenetics | Karyotyping | FISH | Chromosomal microarrays |
| Molecular genetics | Restriction digestion, southern blotting and PCR. | Sanger sequencing and other old-gen sequencers. | Next-generation sequencing. |
| Epigenetics | Methylation-specific PCR, RT-PCR | Chip-chiP assay | Chip-seq, single-cell RNA sequencing and transcriptome analysis. |
Step 5: evaluate accuracy and sensitivity:
Accuracy and sensitivity are two crucial factors one must have to keep in mind while choosing any technology or technique. For example, longer-read sequencing like Sanger sequencing is still more accurate than NGS-like short-read sequencing platforms.
However, NGS comes with high-end sensitivity and thus offers excellent results, guaranteed. Likewise, conventional PCR is an accurate method but real-time PCR is more sensitive and gives us real-time information regarding the copies of the gene present in the sample.
Step 6: evaluate required resources:
Now in the last step, check if you have all the resources, utilities as well as required expertise, experience, or manpower to perform the assay, and go ahead.
Note: you can also consider the cost and time of the assay if the assay is crucial and involves a huge investment.
Wrapping up:
I hope now you get clarity on how you can choose a significant genetic assay for your test, experiment or lab. By following these steps you can select the most appropriate and matching genetic assay for your specific need.
Keep in mind that each assay has its own advantages, limitations as well as applications. Take an expert’s advice or read about the assay before planning to acquire it or select it.


