MLP- Multiplex ligation-dependent Probe AMplification is a nucleic acid quantification technique used in the detection of CNV, SNPs, DNA methylation and mRNA quantification. It is an amplification-based molecular genetic technique.
Molecular genetic techniques are commonly employed in the study of gene expression, amplicon quantification, gene quantification, mRNA expression studies. Some of the techniques like RT-PCR, DNA microarray and MLPA are utilized for this.
Nucleic acid quantification is important in studying gene expression- how much amount a gene expresses in a tissue or cell is a gene expression. Abnormal gene expression causes abnormal protein translation. Quantification techniques quantify either type of nucleic acid present in a sample, thereby helping in studying gene expression or mRNA expression studies.
MLPA is a technique scientists commonly use to study copy number variations like deletion or duplication. Simply put, the technique is based on the principle of hybridization, ligation and amplification.
It’s a multiplex amplification reaction, allowing the amplification of multiple variable-sized templates in a single reaction using a single set of universal primers.
However, analyzing and interpreting data of MPLA requires a high-end, sophisticated and state-of-the-art instrumentation setup like capillary gel electrophoresis.
The present technique is the first choice for DMD gene mutation studies and other copy number variation studies. This article focuses on the technique MLPA (Multiplex Ligation-dependent Probe Amplification ), its principle, technique, process and its applications.
Related article: Polymerase Chain Reaction (PCR)- Definition, Principle, Steps, Procedure, Protocol, Applications and Types.
Key Topics:
MLPA:
The MLPA technique is based on the principle of DNA hybridization, ligation and amplification. The original technique was developed by MRC Holland to quantify copy number variations efficiently. Henceforth it is also known as a semi-quantitative polymerase chain reaction. The technique is completed in the following steps:
- Denaturation: the target template nucleic acid denatures
- Hybridization: probes are hybridized to the target nucleic acid
- Ligation: two probe sequences ligated using a ligase
- Amplification: ligated products amplified in the PCR reaction
- Amplicon separation and data analysis
Principle of MLPA:
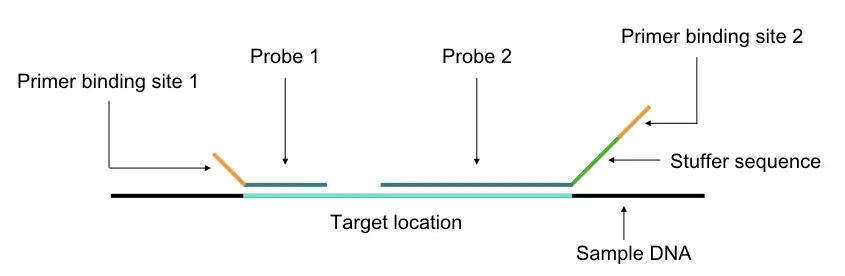
When the dsDNA or nucleic acid denatures probes are hybridized on different complementary locations on the target location. Two different hybridization sequences are ligated using a ligase enzyme. In the next step, A PCR-primers bind to the complementary sequence provided by probe hybridization and allow amplification of various-sized fragments.
The fragments are separated and analyzed for the quantification of nucleic acid.
Process of MLPA:
DNAs are double-stranded, we need to separate them to study them. At 95°C to 98°C temperature, denaturation occurs which separates two DNA strands.
Two different single-stranded DNA for hybridization are obtained.
Followed by the denaturation, hybridization occurs at 60°C.
During the hybridization, varied length probes bind to the target DNA sequence. As it is a multiplex, many different probes are hybridized to the DNA. The probes are 20 to 30 nucleotides long fluorescently labeled nucleotide sequences that bind to the target gene.
Two probes bind adjacent to each other on the DNA having a PCR primer binding site on it which allows amplification in the later stage. Probes have sequences complementary to the DNA.
Note that one of the probes also has a unique stuffer sequence allowing different-sized fragment amplification. It makes each amplification product unique in length. The length of the stuffer sequence varies among different template DNA.
Technically, there are two different types of probes that can be used in the MLPA assays. One is the probe set without the stuffer sequences that can produce products between 94 to 130 nucleotides. This probe usually comes in pairs.
Another probe uses stuffer sequences having one synthetic and other derived probe, and can produce products between 130 to 481.
Which probe set to use depends on assay or mutations to study.
In the next step, the gap between the two probes is filled by the process known as ligation. The ligation process completes in three different temperature-dependent steps at, 54°C, 98°C and 20°C for 15 minutes, 5 minutes and pause, respectively.
Enzymes ligase-65 or NAD- ligase catalyzes the ligation reaction to seal the gap.
Note that the ligase used in the present assay should be thermostable.
Steps and process:
Here we have explained the whole procedure of the MLPA in the present figure.
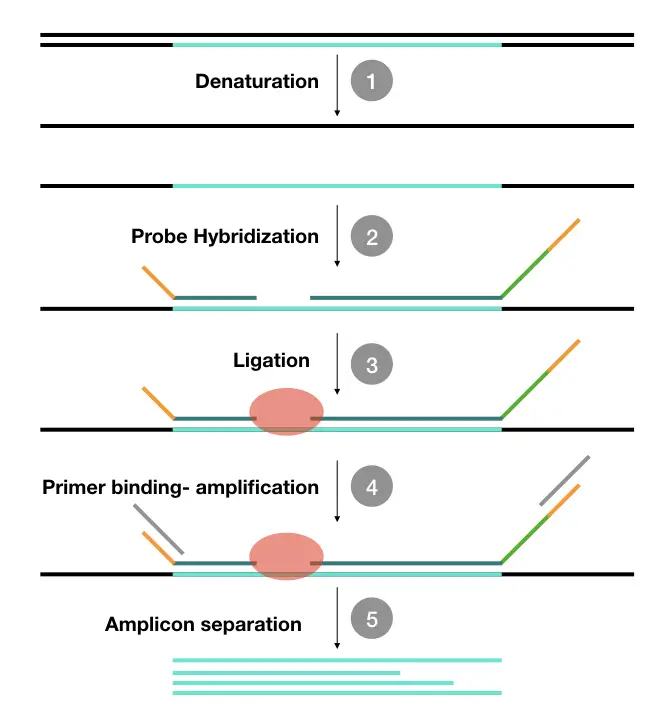
Why is ligation required? If ligation can’t be performed, various-sized fragments can’t be achieved during the amplification.
PCR amplification follows the ligation process. The thermocycler now amplifies the set of reactions, using a single set of universal forward and reverse primers (Every probe set has the same primer binding sites).
Enzyme DNA polymerase catalyzes the amplification reaction and synthesis of new DNA strands using the dNTPs, set of primers, PCR buffer and other ingredients.
The reaction completes 5 steps, 95°C, 60°C, 72°C, 72°C and 15°C for 30 sec, 30 sec, 60sec, 20 minutes and hold.
Note that a separate pair of PCR primers are used which are complementary to the universal primes, one is labeled and another is normal.
In the next step, the fragments are separated on the special type of gel electrophoresis set up known as capillary gel electrophoresis- the technique that has the power to even separate DNA fragments of smaller size.
Here the process of getting MLPA results completes in two steps, separating amplicons on capillary gel electrophoresis and analyzing the fragment using the hybridization data.
Different lengthened fragments run on small capillaries and are monitored. Fragments run as per their length and appear as a peak which is known as Electropherogram.
Different sized fragments have different lengthened peaks due to the use of stuffer sequences used in the amplification. At the end of separation, every amplicon is quantified as well.
Data obtained from the separation are now analyzed in the computational software for further studies. Samples are compared with the reference values in order to study homozygous and heterozygous values.
Read more: What is a multiplex PCR?
Interpreting the results of MLPA:
We have so far quantified the templates right? Now that can also be done in the qPCR too, so you may wonder how it is used in the study of copy number variation.
The copy number variation is when the copies of a gene change in the genome because of deletion or duplication, now let’s how it can be detected using the present assay.
The reference sample probe ratio for reference is always 1.0, having a set of genes present on homologous chromosomes. Two same genes are present on two same chromosomes in us because we are diploid, and that is why both genes should be present in normal conditions.
The score or ratio of 1.0 means both genes are present. A score of 0.5 indicates the deletion of one gene or allele. Which are the heterozygous conditions. Similarly, ratio 1.5 indicates the presence of one extra copy, which means duplication of a gene.
This technique to detect deletions and duplication is routinely employed in Duchenne muscular dystrophy copy number variation studies.
Advantages of MLPA:
The present technique is a version of multiplex PCR that can amplify many templates in a single reaction. Therefore many target genes or gene alteration or copy number variations can be studied in a single reaction.
The present technique is efficient and accurate as well. The chances of error are very less.
It’s rapid and can give results within 24 hours.
It allows the separation of small-sized fragments as well. Even a single nucleotide polymorphism can also be accurately encountered.
It is a high throughput and low input technique.
Related article: Agarose gel electrophoresis.
Limitations:
MLPA technique is an extremely sensitive technique.
State of the art and high-end instrumentation set up is required to conduct the study.
It needs expert hands to operate, perform and evaluate the results of MLPA.
Impurities in the present assay cause failure in the reaction which increases the cost of the assay.
Applications of MLPA:
MLPA is widely used in many different clinical studies and mutation detection systems. One of the most common applications of it is in the detection of copy number variations.
Copy numbers are the number of copies of a particular gene present in a person. Deletion or duplication of chromosome fragments changes the number of copies.
MLPA assay is an efficient method to detect various types of CNVs and henceforth it is used in the exon deletion in case of DMD. deletions or duplications of more than 40 to 60 different DNA samples can be performed using the present assay.
It identifies copies of a gene as well as quantifies the genomic template DNA too.
Detection of single nucleotide polymorphisms:
Another important application of the present technique is in the detection of SNPs, a single base mutation that can’t be identified using conventional PCR.
The MLPA is powerful enough to detect two different templates having a single base change by utilizing two different probe sets and a universal primer set.
Capillary electrophoresis separates various sized SNPs in the run and the hybridization helps in the quantification of templates.
mRNA quantification and profiling:
The ligation based amplification technique- MPLA can quantify not only the DNA but also the mRNA too. Probes specific to mRNA hybridize the template, the machine detects the amount of mRNA template present in a sample.
Smaller gene alterations:
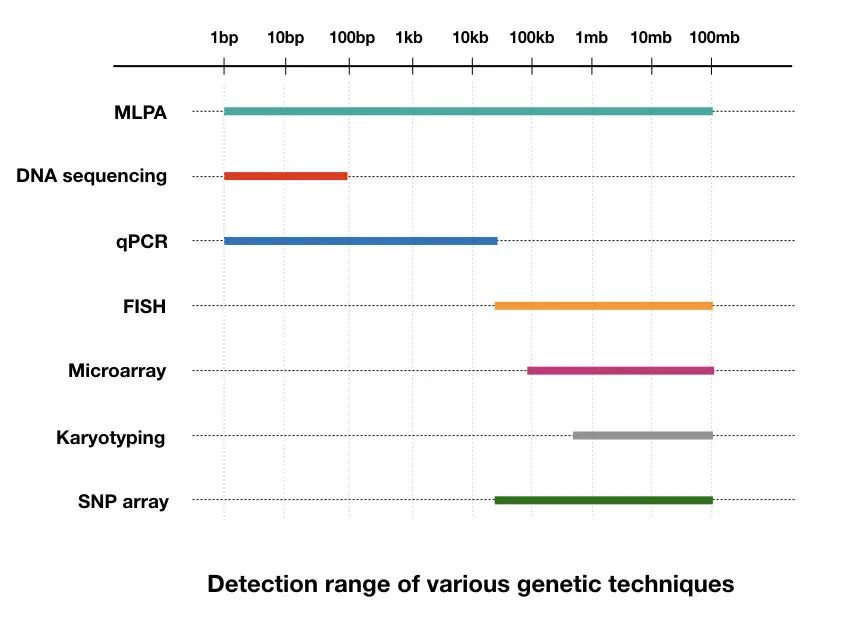
Competitively MLPA is superior to FISH. Though FISH relies on fluorescence chemistry too, it can’t identify smaller gene alterations like small deletion/duplications, additions, translocations or insertions. It finds alterations at the chromosome level. Contrary, the MLPA can even find smaller variations as well, unlike the FISH.
Studying DNA Methylation:
One of the most useful alterations of MLPA known as MS-MLPA or methylation-specific multiplex ligation-dependent probe amplification can find methylation pattern or amount of methylation of template DNA or sample.
An extra restriction digestion step makes it efficient to study methylation related to DNA. methylation profile or methylation pattern is very important in studying transcriptional activity or gene expression studies.
Read more: What Is Genetic Engineering?- Definition, Types, Process And Application.
Conclusion:
The present technique, MLPA is one of the best amplification-based nucleic acid detection techniques that helps in precise quantification of templates. The technique is rapid, accurate and can detect copy number variation effectively.
It is commonly practiced for DMD mutation- exon deletions studies.
Sources:
Stuppia L, Antonucci I, Palka G, Gatta V. Use of the MLPA assay in the molecular diagnosis of gene copy number alterations in human genetic diseases. Int J Mol Sci. 2012;13(3):3245-3276. doi:10.3390/ijms13033245


