“The mRNA possesses the sequences to encode protein only thus to construct the DNA library or the cDNA library, the mRNA is reverse transcribed into the cDNA and processed.”
The mRNA is the messenger RNA transcribed from the DNA or a gene. It’s a part of the central dogma system of a cell.
Actually, the mRNA is not directly used to construct the DNA library (otherwise it is not called a DNA library!). First, it is converted into the DNA which are lesser sequences than the whole genomic DNA.
To understand the present topics, we will first understand the structure of the genome first and soon after we will learn why only the mRNA?
Key Topics:
Why is mRNA chosen for making DNA library instead of genomic DNA?
The genome of us is a huge collection of coding and non-coding DNA. The coding sequences make protein and that is why they are more important than other sequences.
However, the non-coding sequences are important as well.
The coding sequences of a genome are our genes which is only 2% of the total genomic content. It is believed that the coding portion of the genome is associated with vast majority disorders in which either protein function or its structure altered.
This is the reason it is a wise decision to study mRNA instead of the whole genome, right! Why waste money, time, and manpower on 98% of the genome which is not useful to us.
Furthermore, epigenetically, the study of mRNA helps to read gene expression. The gene expression is the amount of a particular gene expressed in a cell.
So by using the mRNA in the experiment, we can get the protein-related information and its expression as well.
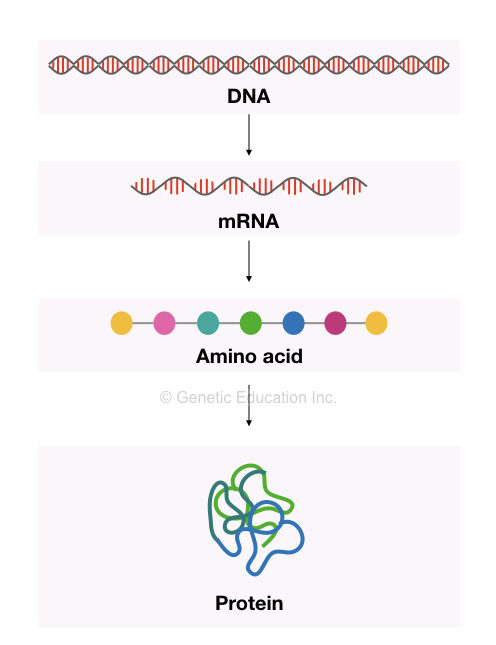
The mRNA is a messenger RNA and the type of ribonucleic acid that has the massage to form a protein.
Once the mRNA is formed, it leaves the nucleus and moves to the cytoplasm to make a protein. Here the mRNA forms the chain of amino acid by taking the help of tRNA and rRNA.
So if we study the mRNA related to genes or the whole coding part of the genome, we can study or know-how actually protein forms.
Although, commonly the mRNA directly is not used (there are so many reasons behind that). We must have to convert it into DNA.
To construct the DNA library or the cDNA library, first, the mRNA is isolated from a cell.
After that, the complementary DNA or cDNA is synthesized from the mRNA. The mRNA dependent DNA polymerase performs this amazing function to convert the mRNA into the cDNA, thanks to the retrovirus!
So technically, once we have synthesized the cDNA, we have all the coding region information encoded back from mRNA to DNA.
The entire set of mRNA of a cell is known as transcriptomics and the technique is known as transcriptomics analysis.
There are several advantages of using the mRNA to construct the cDNA.
The lack of intervening sequences:
The non-coding regions are intervening or introns which are not useful actually. The mRNA lacks intervening sequences that actually we don’t need.
Second, the mRNA copies in cells are high. The mRNA is the expressed portion of a gene thus there are multiple copies of the mRNA present in a cell. We get so many mRNAs at once.
Less background noise:
As the mRNA is the only 2% portion, we no need to process other DNAs that we don’t want to study so there is less background noise or data obtained during the analysis.
Another factor that justifies why we are using mRNA to construct cDNA is the information we get!
The coding sequences have huge information regarding the protein. As it is the collection of the coding nucleotides only. It can provide information on how protein format, its expression, alteration, its structure and its influence on phenotype.
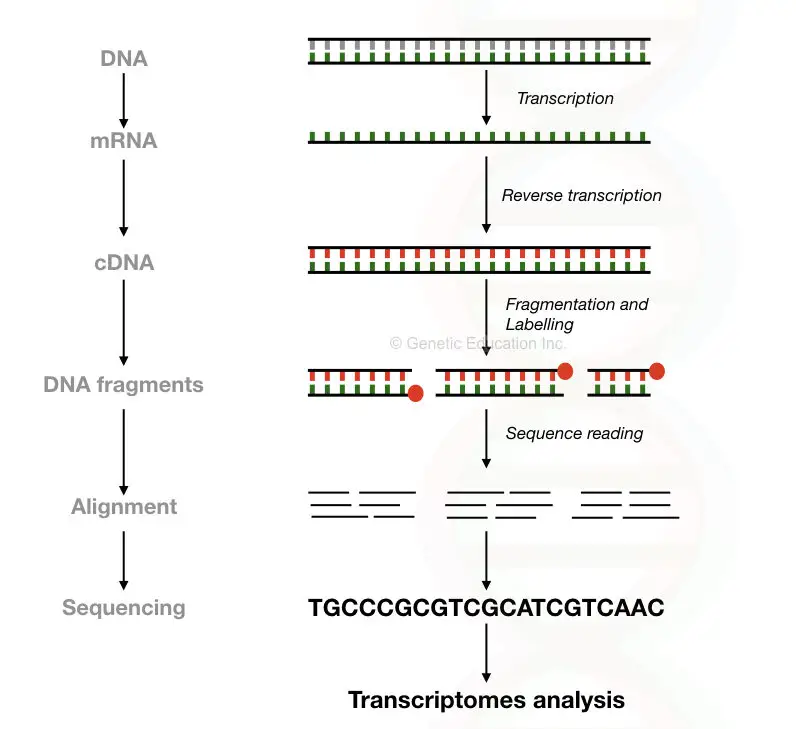
Using the technique known as the reverse transcription PCR, the cDNA is first constructed from the mRNA in the process of making the cDNA library.
The enzyme reverse transcriptase converts mRNA into DNA. After that, the cDNA is fragmented into pieces using various methods and saved into the cDNA library.
The library is the known location of various DNA fragments.
After that, the DNA library or the cDNA library is processed to sequencing where every nucleotide of the library is read, arranged and analyzed.
Furthermore, the cDNA library is also applicable in the microarray analysis to encounter the expression of various genes and alleles. Here I am not discussing the whole process because we already did it. You can read the related article here:
The cDNA library studies are used in reverse genetics in which the genes of eukaryotes are expressed in prokaryotes to study the effect of genes. But the prokaryotes don’t have a definite mechanism to remove the introns or intervening sequences unlike the eukaryotes and henceforth, the cDNA library preparation is required.
Let us check our several differences between cDNA and gDNA:
cDNA vs gDNA library:
The cDNA (complementary DNA) and gDNA (genomic DNA) library are different things!
The genomic DNA is the whole set of the genome or the genomic DNA of an organism while the cDNA is constructed from the mRNA only.
The cDNA is expressed-sequences derived from genes via the mRNA transcript while the gDNA contains coding and non-coding DNA sequences.
The cDNA gives information about gene expression only thus can’t define the organism fully while the gDNA is all DNA and therefore gives more information about the organism from which it originated.
Conclusion:
I think this information is enough to justify why we chose to use the mRNA for constructing the cDNA. Nowadays ready to use kits to make a cDNA is available, also different methods for DNA fragmentation and library preparation are available too.
It is widely used in genome-wide association studies and whole transcriptomics analysis. Read related article: What is transcriptomics?
Sources:
Berger J, Butler P, Troup C, et al. High-Throughput Strand-Specific mRNA Library Preparation for Illumina Sequencing from Total RNA Isolated from Normal and Cancerous Ovary Tissue. J Biomol Tech. 2013;24(Suppl):S42.


