“The excision of a transposon from a native location followed by its integration at another location in a host genome is referred to as a non-replicative mechanism of transposition.”
The non-replicative mechanism of transposition, a kind of ‘cut and paste mechanism’, is also known as the conservative model of the transposition that helps transposons to jump from one location to another without leaving a copy behind.
Henceforth, it is also known as the cut and paste mechanism of transposition. But you may wonder, Why is it known as conservative transposition? During the transportation, on the non-homologous locus, the sequence of transposon remains conserved.
A gene can be introduced into a genome by using transposons therefore it is practically utilized in gene therapy experiments. A gene of interest is excised, transferred into the vector and introduced into the host genome by transposase-mediated transposition.
Conservative transposition is applicable usually for gene therapy and gene transfer experiments in vertebrates and invertebrates. Sleeping beauty transposon system is the best example of non-replicative transposon-mediated gene therapy.
We covered almost all the information about the transposons, in the present series of articles. Some of the previous articles are:
- Transposons: A Jumping Entity and a Foe with Benefits
- Transposase, Transposons and Antibiotic Resistance in Bacteria
- Transposons in eukaryotes
- Role of transposons in evolution
Let me brief you on the topic.
The transposons are the mobile genetic elements present in prokaryotes as well as eukaryotes that can move from one location to another location. Structurally, it contains a gene body that can make a couple of genes, terminal repeats either inverted or direct and a piece of the previous host DNA.
See the structure below,
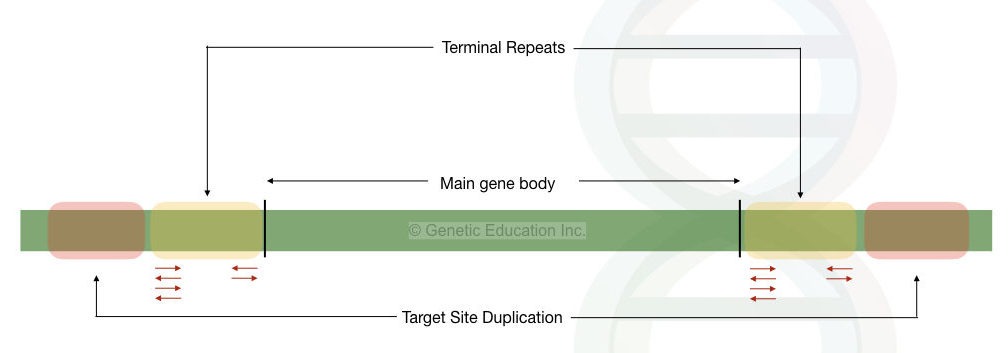
They are categorized into retrotransposons that can move via the RNA intermediate and DNA transposons that can transfer via the DNA intermediates. The transposition mechanism varies among different transposons.
Some of the transposons follow the replicative mechanism of transposition in which After excision from the native place a copy of it remains there. We had covered an entire article on replicative transposition. Please read it first: Replicative transposons.
While in the non-replicative transposition, the TE moves unidirectional, leaving a gap behind. In the present article, we will discuss the mechanism of the non-replicative transposition.
Key Topics:
What is non-replicative transposition?
Without leaving a copy behind, the excision of a transposon from one location followed by its integration to another location in the genome is known as non-replicative transposition. It cuts the transposon from one location and pastes it at another location hence it is also named the cut and paste mechanism of transposition.
Mechanism of transposition:
The non-replicative transposition begins with the activity of a transposase protein. The transposase protein encoded from the transposon, helps it to migrate within a genome.
“It is believed that transposons can move between different organisms too.”
The transposase creates a synaptic complex, initially. The binding of the transposase proteins to their terminal repeats makes a synaptic complex. Let me tell you that the TRS- terminal repeat sequences are present on both ends of transposons and help them to move.
Notedly, the transposase protein (or group of proteins) recognizes those sequences first to initiate the activity. The structure of the synaptic complex is given below,
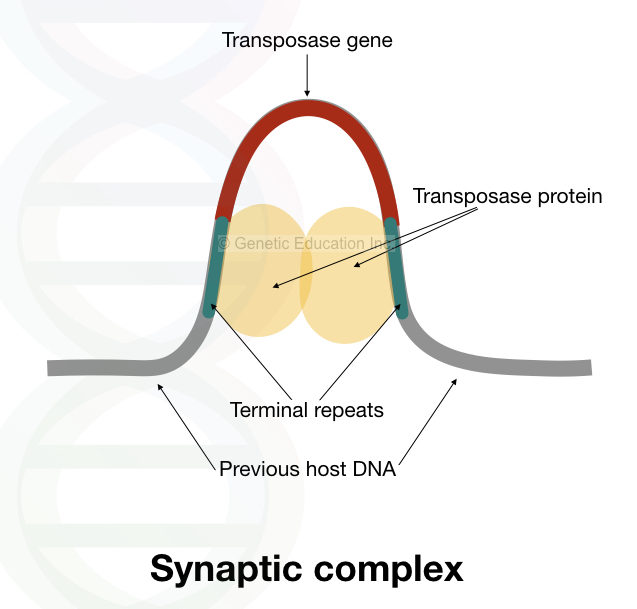
Once it binds to the terminal repeats, it brings the two ends together and creates a stable DNA-protein complex known as a synaptic complex.
Manufacturing synaptic complex begins the process of non-replicative transposition, we can say it’s the very first step in the process. “Transposome” is another name used for the synaptic complex.
The synaptic complex truly decides the fate of the transposon, it ensures DNA cleaving and joining which a transposon needs to move. The next step is excision.
In this step, the DNA-protein complex, a ‘transpososome’ consecutively cuts the transposon. First, it cleaves one DNA strand on each side exactly at the junction between the TE and the flanking (previous) host DNA.
(Remember! the transposon contains some of the DNA sequences at their ends from their previous host DNA).
Here, it aims to generate a free 3’ OH end needed to fill the gap. In the second part of this reaction, the synaptic complex cleaves another strand at both ends. Note that the same transposon isn’t required to catalyze both reactions.
Once the transposon is completely excised from the original position, it’s now ready to move. In the very next step, The transposase creates two single-stranded cuts on the host DNA. Immediately after that, the free 3’ OH end of the transposon DNA attacks the 5’ and the host DNA.
At a molecular level, the reaction occurred as mentioned below,
- The 3′ end attaches to the 5′ end of the host DNA.
- It cleaves the bonds and releases two diphosphates by joining with the 5’ end of the target DNA.
- The reaction occurred on both ends at the same time.
Transposons fill the gap between the junction of the 3’ end (of transposon) and the 5’ end (of the target DNA). An important event that occurs at this point is the gap-filling.
Once the gaps are filled, the transposases are separated from both ends, but another end of the target DNA remains unfilled. The entire process of non-replicative transposition is shown in the figure below,
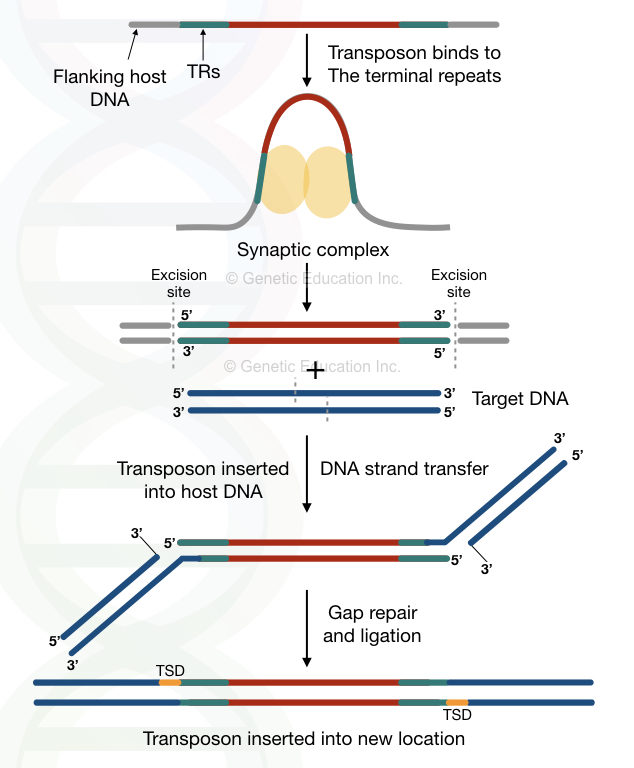
The transposase cannot fill the gap between the 5’ and 3’ ends of the transposon and target DNA, respectively. The 5 to 9 nucleotide gap remains unfilled on both ends.
A cell’s DNA repair mechanism does do this. The 3’ end of the target DNA is free, this site is utilized as a primer by the DNA repair polymerase.
The polymerase fills the gaps by adding nucleotides between the gaps. The same gap is filled by the polymerase on the other side which generates two duplicate sites on both the flanking regions of a transposon.
The entire process is governed by the cellular repair protein and is known as target site duplication. We have covered a whole article on it, you can read it by clicking the link.
The image below clarify things more effectively,
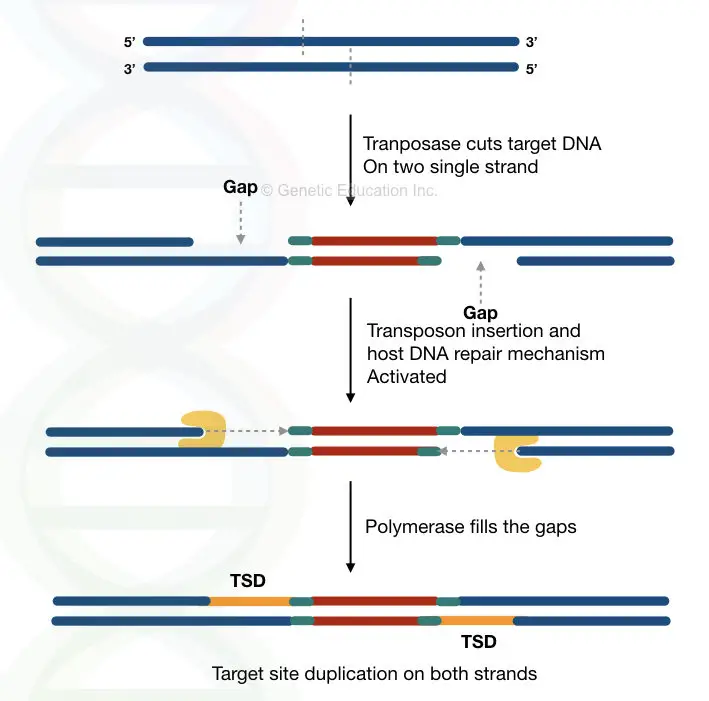
Target site duplication is another important characteristic of a transposon.
“The target site duplication length divulges a distance between the transposase-action site on the host DNA.”
In the final step, the DNA ligase seals the gaps between the newly synthesized DNA. Apart from their role in the transposition, the transposase also performs one important function needed to complete the reaction.
It protects the transposon DNA ends attacked by cellular enzymes during the process of strand transfer.
The gap…..
Now, what about the gap that remains behind!
After completion of the entire process of conservative transposition, a gap, more precisely we can say a double-stranded break remains as such, at the native or old place.
Later on, that gap is filled by the repair mechanism of a cell. It might be joined directly or filled by homologous recombination.
See the figure below,
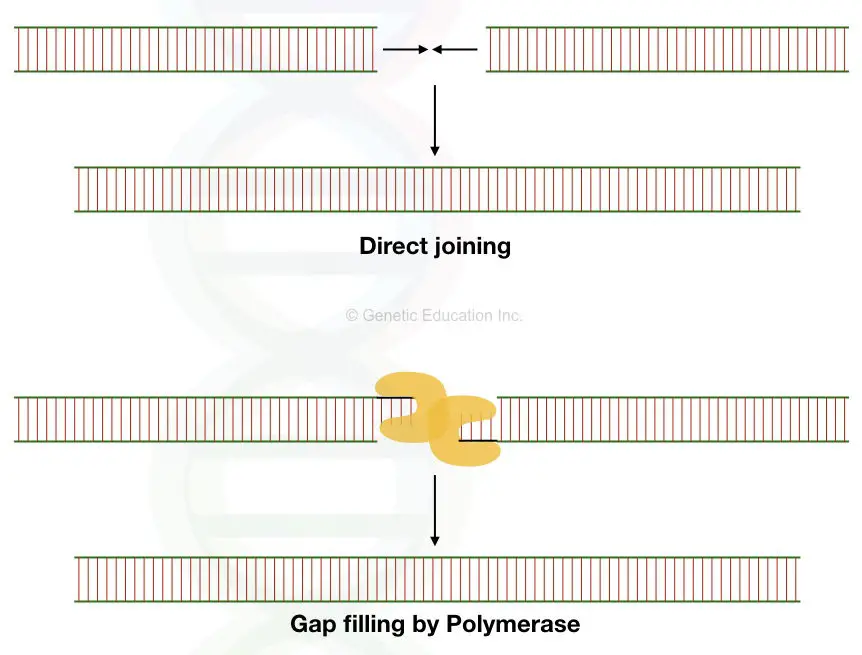
The bacterial Tn10 transposon follows the non-replicative modes of transposition. Our sleeping beauty transposons also do the same.
Conclusion:
The cut and paste/non-replicative/ conservative transposition have significant importance in gene therapies and gene transfer. Researchers are trying to understand the genome complexity using it.
Naturally, the phenomenon helps to produce new variation as the gaps are filled with new nucleotides, which isn’t a part of the original DNA sequence. Hence, the non-replicative mechanism of transposition has a crucial function to perform in a genome- to produce new alleles.
Also, at the new insertion site, the transposon can ON/OFF gene expression, alter gene function or create new allelic variation.
Therefore it is believed that transposons played a significant role in the process of evolution by creating new allelic variations in the population.
Sources:


