“The in situ PCR or slide PCR is a method used to amplify the DNA present in the paraffin-embedded tissue or formalin-fixed tissue on the glass slide.”
In the traditional PCR or in the conventional PCR method, the DNA is amplified once it is extracted from the sample.
For extracting DNA, the sample of the tissue is proceeding from one of the methods of DNA extraction.
Because the molecules present into the cell wall or cell membrane as well as into the cytoplasm interfere in the amplification process and the DNA cannot be amplified.
So it is very important and crucial to extract pure DNA from the cell. For performing PCR, the purity of the DNA must be nearby ~1.80 and the quantity must be above 50ng.
The amplification of the DNA is taken place in a temperature-depended cyclic manner. The denaturation occurs at 94°C, annealing at 55 °C 65°C and extraction of the newly grown DNA occur at 72°C.
See the PCR cycles in the figure below,
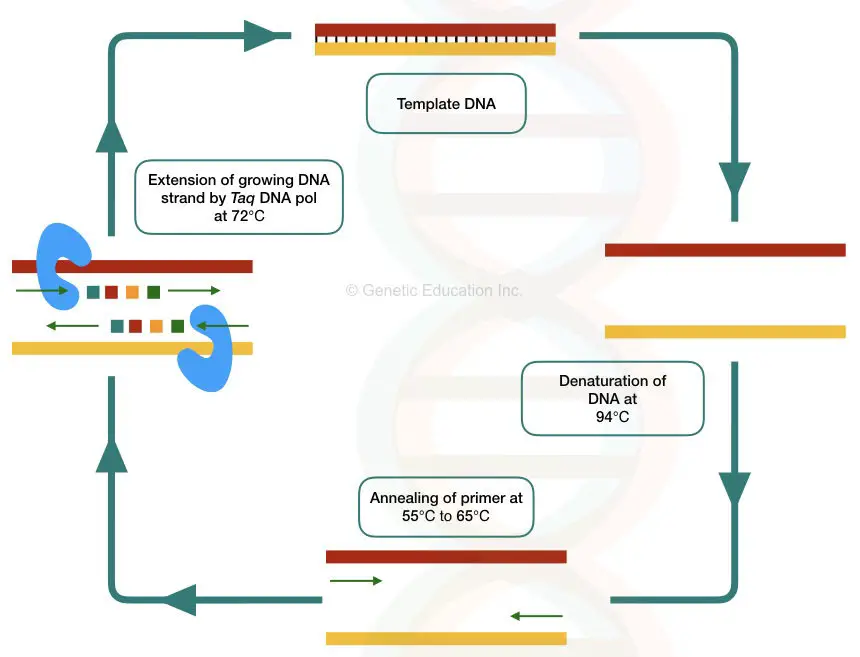
In the in situ PCR, the PCR reaction is directly taken place on the slide containing the tissue.
The tissues may be frozen, older, ancient, paraffin-embedded or cultured cells. The master mix is directly applied on the surface of the slide, and the entire amplification process occurs on this slide. However, the master mix used into the in situ PCR is specially prepared.
In this article, we will discuss the general process of the in situ PCR and also its benefits and limitations. The content of the article is,
Key Topics:
What is in situ?
The word in situ PCR is a combination of two words: in situ + PCR.
The Wikipedia defines the word “ in situ” as “on position”, “on the site”, “on the original place.” The word is Latin.
Clearly, it means that any reaction taken place in situ means it occurs on its original place or position.
Our sample is paraffin-embedded tissue thus the reaction occurs directly on the paraffin-embedded tissue, after several purification processes.
If our sample is a layer of cells, the biological reaction taken place on the surface of the cell layer, is in situ.
The in situ reaction occurs in an appropriate position while the ex-situ reaction occurs other than its original position.
The general in situ reaction is the hybridization used in the FISH and DNA microarray.
What is in situ hybridization?
The hybridization process happens directly at the exact position on the tissue, on the surface of the slide with the help of the DNA or RNA is called as in situ hybridization.
The excellent example of in situ hybridization is the use of the fluorescent labelled probes in the FISH (fluorescent in situ hybridization).
In the FISH, the tissue sample is fixed on the slide and permeabilized with the help of the chemical methods.
A fluorescently labelled short DNA or RNA probes are applied to the site of hybridization and with the repeated process of chemical treatment and washing, the micromolecules hybridized.
The in situ hybridization technique is very useful in the identification of minor chromosomal deletions, duplications and additions. Also, other cytological and molecular aberrations can be detected using the in situ hybridization.
Read the article on different types of mutation: Different types of Genetic mutations
What is in situ PCR?
The idea behind the in situ hybridization may have been cleared, I hope so,
Right?
So the question is how PCR is possible on the slide? Because we have to extract DNA to do amplification.
“The PCR reaction occurs on the tissue, located on the slide, followed by proteolytic treatment and washing is refers to as in situ amplification or in situ PCR.”
The principle of in situ PCR:
The in situ PCR is completed into the three steps.
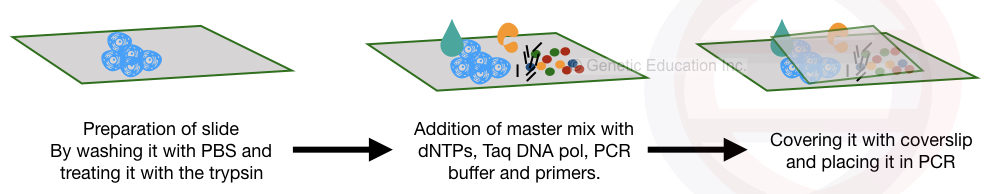
Preparation of the tissue for the amplification, proteolytic digestion and DNA amplification.
The tissue is fixed on the silane-coated slide. The silane is used to fix the tissue or the cell on the slide which means it helps in cell adhesion.
After fixing and cleaning the tissue, the proteolytic enzymes digest the protein portion of the cell. Pepsin, trypsin and proteinase K are generally used in the digestion reaction.
Pepsin, trypsin and chymotrypsin are always preferred in the in situ PCR digestion because of its shorter life span.
Once the digestion is over, the specialized mastermix is applied to the surface of the cell along with the primers and other PCR additives.
After covering it, the slide is directly placed in the thermocycler for the amplification.
The in situ PCR technique is quite tedious and sensitive. The yield of the reaction depends on how we prepare the sample for the amplification.
Sample preparation for in situ PCR:
The technique is specialized and sensitive, starts with the sample preparation.
The starting material or tissue for the in situ PCR are paraffin-embedded tissues, cell culture, a cell suspension or a thin layer of cells.
Now add formalin to the sample and place it for 10 to 12 hour or overnight. Use 10% formalin.
Wash the sample or tissue with phosphate buffer saline twice or thrice to remove other impurities.
If the sample is cell suspension, wash with PBS followed by the two round of centrifugation of 25,00 rpm 10 minutes.
Washing is very important to remove other impurities from the surface of the tissue.
Note
Use DEPC-treated water to wash all the utilities while doing RNA extraction and reverse transcription PCR. To know more, you can read this article: 11 Effective Ways to Use DEPC-Treated Water in RNA Isolation.
Now prepare the slide by coating it with the silane. Silane facilitates cell fixing.
Place our tissue or the thin film of the paraffin-embedded tissue block to the slide.
Paraffin can block the amplification so it is necessary to treat the tissue (which is embedded in the paraffin) with the xylene-ethanol treatment.
Place the slide in the fresh xylene solution for 8 minutes and then place it under the pure ethanol (100%) for 10 minutes.
Air-dry the slide to remove excess ethanol.
Now our tissue is fixed on the slide and ready for the PCR.
Read more:
The procedure of in situ PCR:
The process of in situ PCR start with fixing the tissue which we had already discussed.
The fixing helps to maintain the structural hierarchy of the tissue and also prevents the morphology of the tissue.
This will preserve the integrity of the DNA too.
In the next step, the sample or the slide is processing for the proteolytic digestion.
2mg/ml of one of the proteolytic enzyme can be used for the digestion followed by the washing with the alcohol.
The sample is ready for the amplification step.
We have to prepare the master mix for the in situ PCR. for that, one of the types of primers are selected from the list below,
- Fluorescently labelled primer (20 to 22 nucleotides long, for real-time PCR)
- Random hexa or octa nucleotide primers (simple PCR)
- Oligo (dT) primers (20 nucleotide long)
It is the best variation of real-time PCR. Hence the in situ PCR is usually applied to quantify the DNA present into the sample.
Use fluorescentl-labelled oligo (dT) primers for Real-time PCR.
Read more on realtime PCR: Real-time PCR: Principle, Procedure, Advantages, Limitations and Applications
Once the reaction is prepared, the slide is covered with the coverslip and placed into the thermocycler.
If you are doing reverse transcription PCR, additional DEPC wash and DNase digestion step are required before PCR reaction preparation.
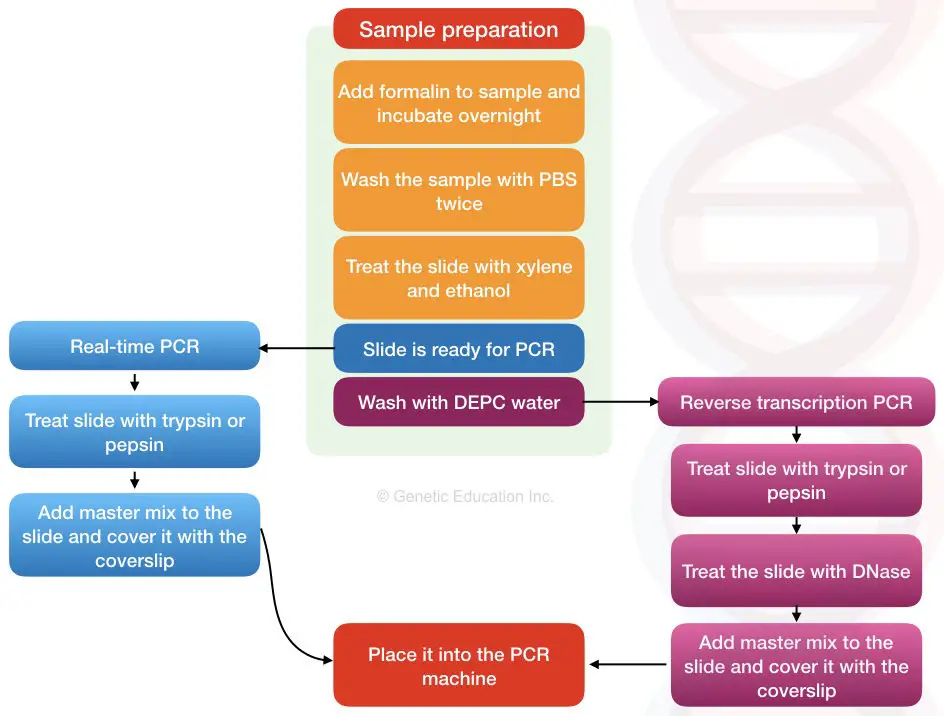
Application of in situ PCR:
By using the in situ PCR even a single copy of the DNA present into the sample can be measured or amplified.
Localizing and visualizing the amplicon within the cell is possible by the in situ PCR.
The low copy number of DNA can be detected with high sensitivity by this method.
It is widely used in the study of organogenesis and embryogenesis, also, it is used in infectious disease diagnosis such as HIV.
Quantification of DNA is also possible in real-time in situ PCR.
Gene expression can also be measured using the reverse transcription in situ PCR.
Limitation of in situ PCR:
The chance of non-specific binding is higher during in situ PCR.
The precision of the reaction is also low.
Multiplexing is not possible due to the fewer copy number.
Wrapping up:
The in silico PCR or the slide PCR technique is the best method for the detection of low copy number of nucleic acid. In the reverse transcription in situ PCR the mRNA is the target from which cDNA is generated and the gene expression level is directly measured on the surface of a slide.



Its good article, thanks.