“Long-range PCR is a method to amplify the DNA fragment of more than 5Kb using a special type of high-fidelity DNA polymerase.”
Using the PCR ingredients like dNTPs, PCR buffer, DNA primers and template DNA, copies of DNA can be generated in a PCR. It is a cyclic biochemical reaction based on the mechanism of temperature-based amplification.
A conventional 25μL or 50μL reaction can amplify only several thousand basepair DNA fragments. Efficiently up to 1500 to 1800bp fragments can be amplified into the conventional normal PCR reaction. However, to amplify the fragments of 5kb, 10kb or 20kb is quite impossible using the normal PCR setup.
We are willing to amplify a long fragment, right! so it is obvious that we need more reagents, PCR cycles and primers. Also, to achieve amplification for longer PCR, one should have expertise and experience in using different combinations of chemicals. Great precision and high-end experiment setup are must require for long-range PCR.
In the present article, we will explain to you what a long-range PCR is and how to perform it. We will also give you a complete protocol and set up for you.
Key Topics:
A conventional PCR:
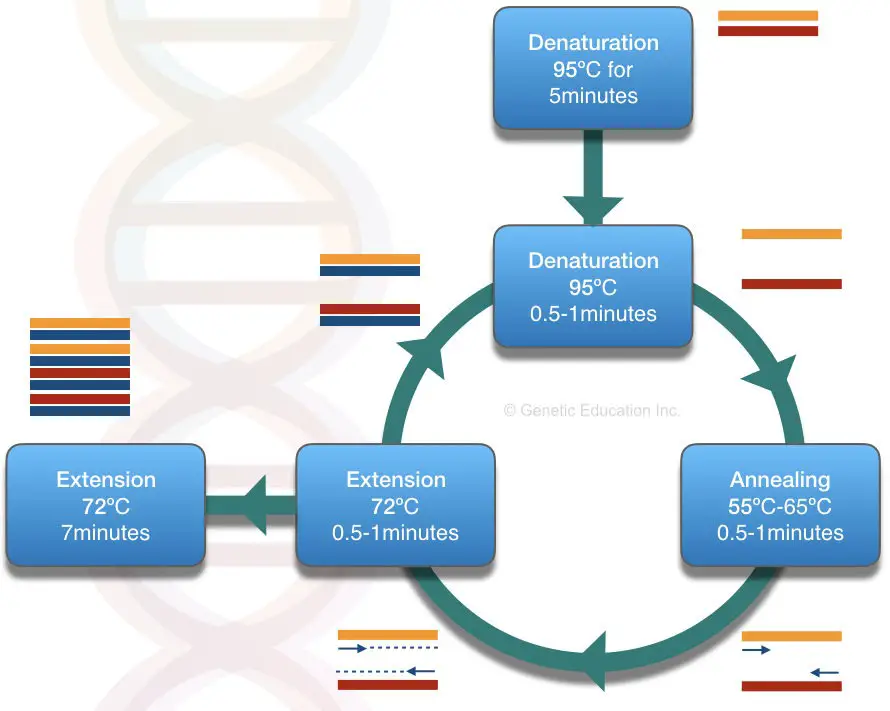
A PCR reaction is a cyclic condition in which our DNA of interest is amplified. It is a temperature-dependent in vitro process of replication completes in three steps
- Denaturation
- Annealing
- Extension
In the denaturation step, at the 94 ̊C, the DNA template of our interest denatured or break opens into two single-stranded DNA.
In the next step- the annealing period, the sequence-specific primers bind to the DNA between the temperature of the 55 ̊C to 65 ̊C (at its annealing temperature).
Finally, in the extension step, the DNA is synthesized with the help of the Taq DNA polymerase. The process of PCR is explained in the figure below,
Read more,
The PCR products are run on the agarose gel electrophoresis.
Depending upon the requirement of the researcher different modifications to the native PCR reaction can be done. For example, to achieve higher specificity into the reaction Nested PCR or Hot start PCR can be performed. To amplify a specific allele, allele-specific (ARMS) PCR can be performed.
The long-range PCR is also a kind of modification of native PCR, designed specially to run for long DNA.
What is a long-range PCR?
The extensive protocol setup, the makeup PCR reaction buffer and optimized Taq DNA polymerase are three key factors to achieve success in long-range PCR. But the question strikes in your mind that what is the need to perform a long-PCR?
We can identify the mutation and single-gene disorders by conventional PCR, then why do long-range PCR? What is the need?
Well, the size of the fragment is also as important as the content of the DNA fragment, for the detection of abnormalities in some diseases the size of the DNA fragment matters! for example, the fragile- X syndrome.
Fragile-X syndrome occurs by the abnormal expansion of the high GC-rich region known as triplet CGG present on the X chromosome. The region is too long but if we amplify that region and compare it with a normal one, we can interpret the results.
Based on the size of each fragment normal, carrier and disease alleles can be detected. Moreover, the long-range PCR method is also used in other cases such as,
- Genotyping,
- DNA fingerprinting
- The DNA sequencing of long fragments
- Copy number variation studies
Now before discussing the protocol, let us see some of the optimization needed for the long-range PCR.
Optimization of long-range PCR:
Several optimizations are required to achieve amplification in the longer template DNA. We need a different type of Taq DNA polymerase, different types of primer sets and PCR reaction buffer.
Taq DNA polymerase:
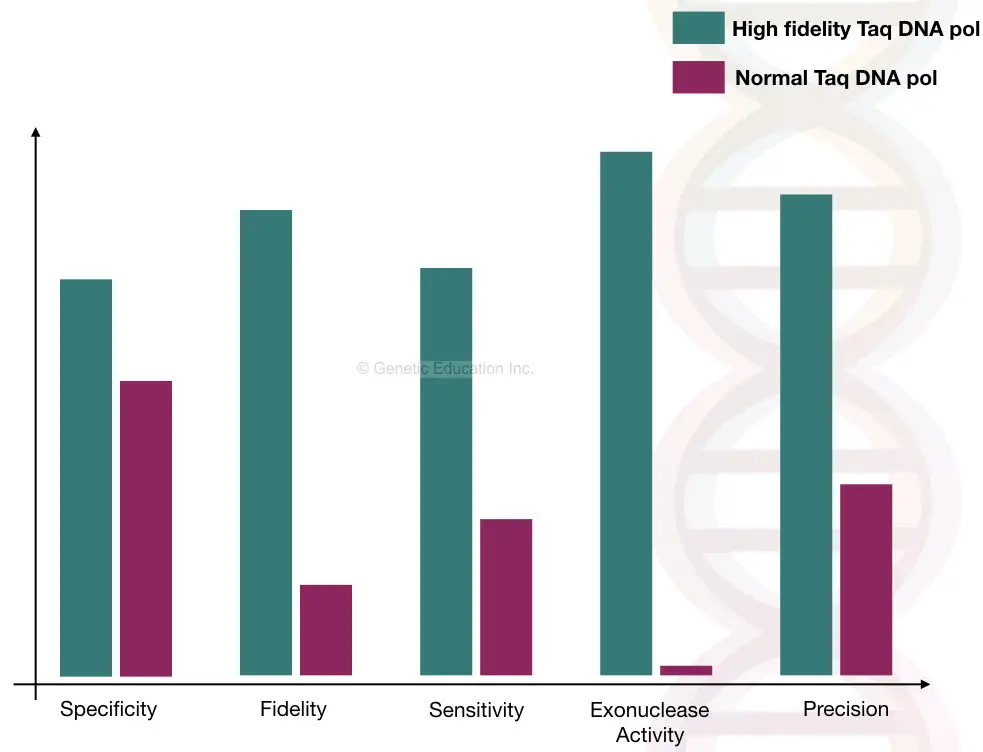
Now one thing is clear, the normal Taq DNA polymerase is not working here. The normal Taq has several limitations,
It has low fidelity, can add up to several nucleotides and does not have the proofreading activity. The routine Taq DNA polymerase can add dNTPs up to 1kb to 2kb, after that the chance of the incorporation of the wrong dNTPs is increased because the specificity of the Taq will gradually decrease.
That is why the normal Taq DNA polymerase cannot be used in the long-range PCR reaction. Nonetheless, the Taq DNA polymerase is the best choice for the conventional short PCR cycles. We have explained the importance of Taq DNA polymerase in one of our articles, read it here: Function of Taq DNA Polymerase in PCR.
The Taq DNA polymerase that we want to use in the present technique must have a high amplification rate, low mismatch rate, high fidelity, efficiency, and high specificity. Importantly, the Taq used in long-range PCR must have the 3′ to 5′ exonuclease activity.
The best alternative we can use instead of Taq DNA polymerase is the ‘Vent‘.
The Vent has all these characteristics. The Vent has 3′ to 5′ exonuclease activity, high-fidelity and a high amplification rate. A higher concentration of enzyme is required for a long-range PCR, though.
In another method,
Use a combination of two different polymerases viz one with high concentration, normally the Taq DNA polymerase and another one with low concentration, a high fidelity DNA polymerase. Use 1 to 1.5 unit normal Taq and 0.5 to 1 unit of high-fidelity DNA polymerase.
If you want to use a single high-fidelity Taq DNA polymerase use 2.5 to 3 units into a single reaction.
PCR primers:
The primer set of normal PCR can’t work here. The primer set employed in the long-range PCR should have some unique and different properties.
The primers used in the long PCR are 22 to 30 nucleotides long and 50% of less GC content in it. Furthermore, it should have a low capacity of secondary structure formation and annealing temperature between 58 ̊C to 68 ̊C.
As per the requirement use 0.1 to 1.0 mM of primers. Ideally for the fragments up to 20kb use 100pM to 200pM of primers. Nonetheless, optimize the reaction by using different combinations of primer concentration if needed.
Readout our guide to designing PCR primers: PCR primer design guidelines.
Important
For the complex PCR template (DNA) use a low concentration of primers; For simple or lower complex template (DNA) use a high concentration of primers.
PCR reaction buffer for Long-range PCR:
The reaction buffer or the PCR reaction buffer used in the present type of PCR is also different. We can’t use a routine buffer for it. For example, to facilitate a great amplification rate it should have a high amount of MgCl2.
The PCR reaction buffer must contain 2mM to 5mM MgCl2. However, the concentration of magnesium ions depends on the purpose of our reaction because a higher concentration of reaction increases the non-specific bindings into the reaction.
Also, we can use KCl, Tricin, DMSO and other chemicals as per the requirement of the reaction. For the long-range PCR, we personally recommended using the ready-to-use PCR reaction buffer.
Template DNA:
we are amplifying longer DNA fragments, which means we need a larger amount of each chemical and PCR template DNA into the reaction. Use up 500 ng to 1000ng template-DNA into the long-range PCR.
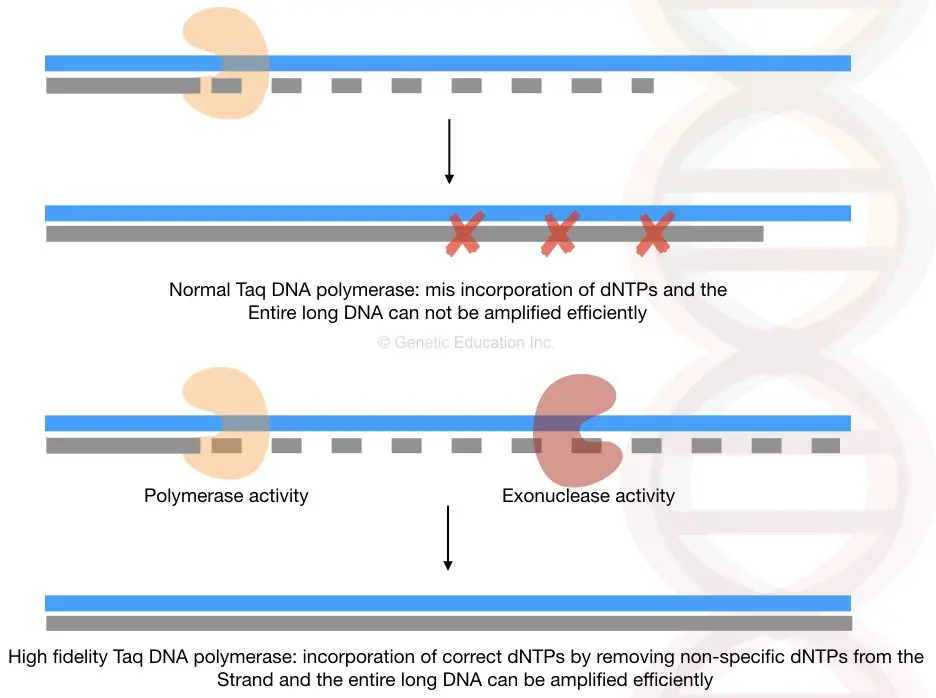
Also, the template DNA must be intact and highly pure otherwise DNA polymerase cannot work efficiently over a period of time. The exonuclease activity and polymerization activity of high-fidelity DNA polymerase is explained here,
Protocol for long-range PCR:
The concentration of each component used in the PCR reaction is already given in the above section. The PCR reaction preparation for long-range PCR is given below,
| Component | Stock | Concentration | Quantity |
| dNTPs | 200µM | 400µM (100µM each) | 8µL |
| PCR reaction buffer With MgCl2* | 5X | 1X | 5µL |
| High fidelity Taq | 5U | 2.5U | 0.5µL |
| Forward primer | 100pM | 200pM | 2µL |
| Reverse primer | 100pM | 200pM | 2µL |
| Template DNA | 250 ng | 2µL | |
| Water | 30.5µL | ||
| Total | ————————– | 50µL |
If needed add MgCl2, if not provided into the buffer,
Note: the PCR buffer must have MgCl2.
The PCR reaction conditions for the long-range PCR is divided into two steps and shown below,
The first set of a cycle:
| PCR Steps | Initial Denaturation | Denaturation | Annealing |
| Temperature | 94̊C | 98 ̊C | 68 ̊C |
| Time | 1min | 20 sec | 20 min |
| ——————– | 10-15 cycles | ——————- |
The second set of a cycle:
| PCR Steps | Denaturation | Annealing | Final extension |
| Temperature | 98 ̊C | 68 ̊C | 72 ̊C |
| Time | 20 sec | 20 min | 10 min |
| ——————– | Remaining 16 cycles |
After the completion of the PCR reaction, the PCR products are run on the 2% agarose gel electrophoresis.
Advantages of long-range PCR:
The specificity of the reaction increases by using the high fidelity Taq.
It facilitates the amplification of longer DNA fragments and also the high GC-rich regions.
Disadvantages of long-range PCR:
The process is time-consuming.
The success rate is also very low.
The chance of misincorporation of dNTPs is high.
The technique is not suitable for routine use.
Read more:
Conclusion:
Long-range PCR is the first choice for the diagnosis of long CpG repeat-containing templates. Also, it can be useful in downstream applications. However, highly purified template DNA and extensive experimental setups are required for the present technique.
Still, up to 20kb DNA fragments can be amplified with a long-range PCR protocol.