“Probes are the single-stranded nucleic acid employed in the hybridization while primers are used in the amplification.”
Hybridization and amplification are two key methods or we can say, techniques used in almost all type of molecular genetic laboratories.
This clearly means one should learn hybridization as well as amplification to acquire different technique.
We all know about the primers right!
A primer is used during the DNA replication, synthesised by the primase enzyme and helps in synthesise of growing DNA strand.
Now, what is the role of the primer in that? Is it actually helps in adding nucleotides?
No!
It only provides the free 3’ OH end for the DNA polymerase to work.
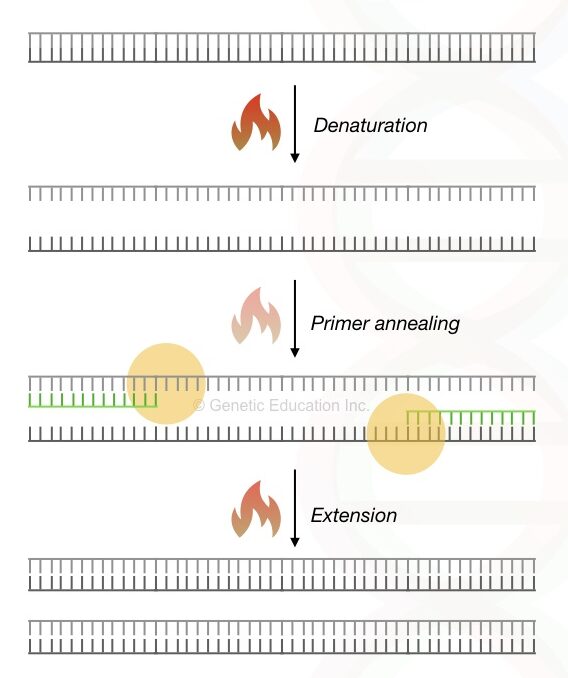
Amplification is a method for synthesis of DNA done through the polymerase chain reaction, often known as a PCR and done in a thermocycler.
Using different temperature gradient, a special type of DNA polymerase called Taq DNA polymerase utilises dNTPs, primer and reaction buffer to synthesise new DNA strand, in vitro.
On the other size, the hybridization is different.
Applying stringent condition, a complementary labelled nucleotide (known as a probe) binds to its exact complementary sequence of the target nucleic acid molecule.
Simply, binding of the complementary nucleic acid sequences is known as hybridization.
Well, as we said both methods have their own importance and drawbacks as well as similarities and differences.
In the present article, we will talk about some of the differences and similarities of the probes and primers used in the hybridization and amplification, respectively.
Key Topics:
Differences between probe vs primer:
The length:
One of the crucial factor in any of the reaction is the length of the nucleic acid molecule used as a primer or probe.
The primer must be shorter ranging from 15 to 25 nucleotides while the probes are longer; ranging from 100 to 1000 or sometimes 10,000 nucleotides in length.
Function:
The primary function of the probe whether it is DNA or RNA probe is to detect the presence or absence of target nucleic acid present in a sample.
While the function of a primer or a DNA primer is to provide a free 3’ end for Taq DNA polymerase to synthesise new DNA strand.
The probe binds at our target location whereas the primer binds from where the amplification will starts.
Probes are used for the detection through hybridization thus, there must be a colour molecule attached to it.
The probes are generally labelled with the radiolabelled or non-labelled compounds while the primers are not labelled.
Because of that, there is a little risk associated with the use of probes, on the other side, the primers are safe to use.
Read more on probe designing: DNA Probes: Labelling, Types And Uses.
Mechanism of action:
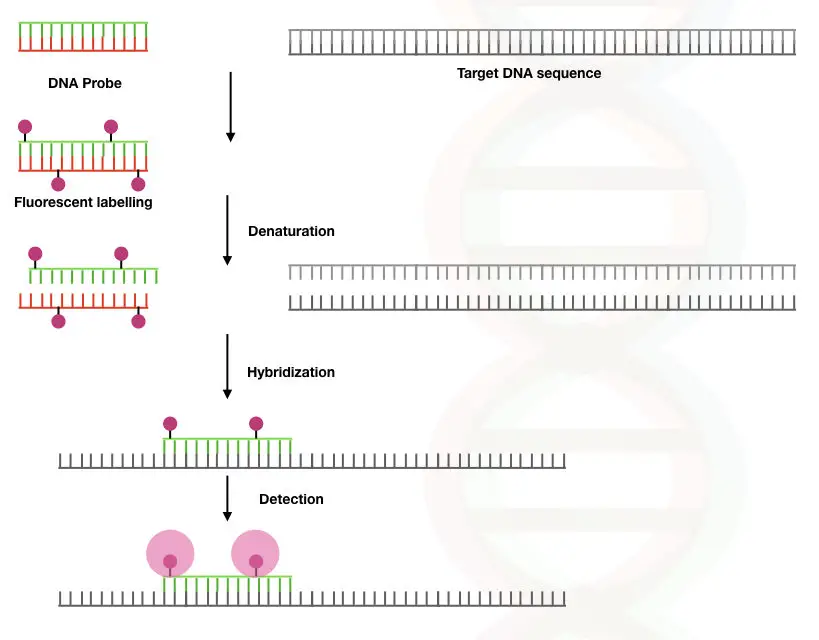
The long chain of the single-stranded probe contains the fluorochrome or a radiolabelled compound on its end.
Once it finds its complementary sequence on the template DNA it binds to it, the fluorochrome is released and emits fluorescence which is measured by the detector.
Thus the amount of the fluorescence emitted is directly proportional to the probe hybridization.
On the other side,
Under the exact annealing temperature, the primer binds to its complementary sequence. Once the annealing happens, the Taq DNA polymerase starts inserting the dNTPs next to the 3’ end of it until it reaches the end of the strand.
At the end of the reaction, a new DNA molecule is generated.
Read more: The Function Of dNTPs in PCR Reaction.
Application:
The probes are used in the hybridization experiments such as ISH- in situ hybridization, fluorescence in situ hybridization and DNA microarray.
The primer is used widely, in PCR experiments and DNA sequencing.
The nucleic acid probe, in the hybridization, binds to the target location and finds copy number variations.
It can also be employed directly on the chromosome for detecting mutations and copy number variation.
Larger copy number variations such as larger duplication, deletions, inversion or insertion can be encountered using the probe-based hybridization method.
Contrary to this,
The primer based amplification methods are used for generating DNA amplicons of interest which can be used for DNA sequencing or for constructing cDNA library or DNA library.
Minor alterations such as SNPs- single nucleotide polymorphisms in which a single base change can cause a mutation, can be detected using the primer based amplification methods.
The hybridization method is applicable in the detection of copy number variations, genome-wide association studies and disease screening.
The PCR is used for diagnosis of various inherited disease, DNA fingerprinting and genetic engineering experiments.
Limitations:
The major limitations of the probe-based hybridization method are its low efficiency. The probe can bind to non-target locations and gives false results.
Furthermore, FISH like methods are unable to detect minor variations, SNPs can not be detected accurately.
New mutations can not be identified, because sequence information is required to construct the probes.
On the other side,
The major limitation of the primer based PCR methods is its inability to amplify larger DNA sequences.
Furthermore, hard templated such as high GC rich DNA templates can not be amplified properly.
Although sequence information is required to construct a primer, still, using DNA sequencing like methods, new variations and mutations can be discovered using the primer based method.
As per my practical knowledge, the probes are available in different variations, for example, some probes like TaqMan are linear while some are structurally different- molecular beacons are hairpin shaped probe.
On the other side, the primers are a linear molecule- shape variations are not available for enhancing amplification.
Read more: Molecular Beacon: A hairpin that enhances real-time PCR specificity.
Similarities between probe vs primer:
Even though nucleic acid probes and primers are used for different applications still several similarities occur between both.
Both are the strand of nucleic acid either DNA or RNA.
For designing either probe or primer, sequence information must be required.
Both can be designed artificially or by using the polymerase chain reaction.
Both primer and probe are used in the realtime PCR for quantification of nucleic acid and identification of microorganism.
A high level of expertise required for using both the elements thus the chance of reaction failure is higher in both.
Also, the chance of false-positive result is higher in both amplification and hybridization method.
Related article: Comparison Between DNA Primer And RNA Primer.
Conclusion:
Probe and primer are important elements in molecular genetics, one should know about it. There are so many online software is available to design probes as well as primers for different sequences.
The primer 3 is one of them. we have covered an article on it. You can read it here: PCR primer design guidelines.