“Restriction enzymes or endonucleases are the class of enzymes that perform a catalytic reaction to cleave the DNA. EcoR1, BamH1 and HinfI are examples of some restriction enzymes used in genetic engineering.”
Genetic engineering is all about cutting, modifying and ligating DNAs. It is a kind of ‘garage’ to modify and repair DNA. The purpose to modify or repairing DNA is to make it useful for us. For instance, the techniques of genetic engineering are used to modify DNA in order to cure genetic disorders.
Another example is to produce economically important plant species which are useful for us and so many others.
Tools like nuclease, helicase, ligase or plasmid make it possible to edit the DNA of the host organism. Every tool is employed to perform a different function. Here are some of the functions:
Vector: to transfer the gene of interest to the target location
Plasmid: to express and transfer the genetic material (Plasmid).
Ligase: to seal or join the DNA fragments (Ligase).
Helicase: to release the tension on DNA (Helicase).
Restriction endonucleases: to cut the DNA at a specific location.
Polymerase: to synthesize the DNA artificially (Polymerase).
Restriction endonucleases are the class of nuclease enzymes that have the power to cut the DNA. The endonucleases are different from the exonucleases having a specialized function to cut DNA inside.
On the other side, the exonucleases cut or cleave the DNA outside, on the ends. Every endonuclease has its own recognition site where it binds and cleaves the DNA. To read more on the present topic you can read this article: endonuclease vs exonuclease.
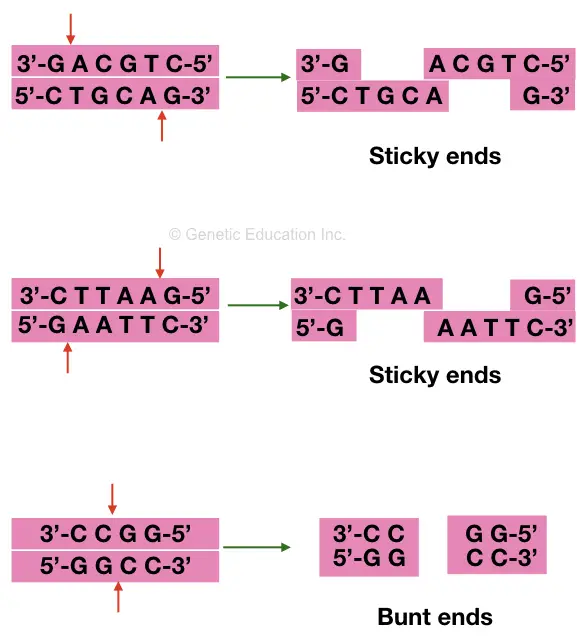
Sticky ends and blunt ends are two types of cleaved products, generated on digestion. Usually, the restriction enzyme generates both types of ends.
The nuclease, especially endonuclease, is derived from the prokaryotes and found in them only. It is present in prokaryotes to protect the bacterial DNA from phage attacks.
Here in the present article, we will concentrate on restriction enzymes, one of the important genetic engineering tools. We will try to cover all the details of it and its possible applications.
In addition, we will also give you the list of 10 important restriction enzymes, commercially available and widely used in recombinant DNA technology.
Key Topics:
Restriction enzymes: Definition, Types and mechanism and application
What are the restriction enzymes?
The restriction enzymes are a protein, obviously- a type of DNA enzymes that are widely used in site-specific cutting DNA viz- in genetic engineering and biotechnology experiments. They are used to cut DNA at a location we wish to study.
Much like other enzymes (ligase or helicase), restriction enzymes are pivotal in ‘in vitro’ experiments of DNA modification. Their function gives us the flexibility to play with the target DNA, to remove or insert some DNA sequence.
Werner Arber, Hamilton-Smith and Daniel Nathas are the three pioneers who discovered restriction enzymes during 1960 and 1970.
The restriction enzymes are the type of nuclease known as endonuclease more precisely- restriction endonucleases having the capacity to cleave or cut DNA inside. Nucleases are of two types, endonucleases that are our restriction enzymes and exonucleases that can cut DNA outside or on the ends.
Although the endonucleases are widely used in the ‘in Vitro studies in comparison with the exonucleases. It was believed that the endonuclease enzymes were evolved from the transposable elements.
Read more: What is Restriction Digestion and how to do it?
Definition:
A special class of DNA enzymes known as “restrictase” or restriction endonucleases that has the power to cut or cleave DNA at a very specific location and so used in genetic engineering experiments is known as restriction enzymes.
Restriction modification system:
You may wonder, are restriction enzymes present in eukaryotes?
The restriction enzymes are originally discovered from bacteria. Indeed, it is present in prokaryotes only.
Restriction enzymes are evolved in prokaryotes to protect the host bacteria from viruses and pathogenic invasion. Once it recognizes foreign DNA through its recognition site, it immediately cleaves it and destroys the pathogen, virus or phase.
Nonetheless, it can cut any kind of DNA having their recognition sites, but can’t cut their own DNA, thanks to one special mechanism- methylation, also performed through restrictases.
Methylation of DNA protects bacteria’s DNA from their own restriction enzymes.
Another enzyme known as methyltransferase inserts methyl groups at the location where the REase act, resultantly, makes them unable to cut their own DNA. The insertion of the methyl group prevents self-restriction digestion.
More than 600 different restriction enzymes are used in genetic engineering and molecular cloning experiments, though a total of 300 REase is well studied to date.
Types of restriction enzymes:
Based on their action, cutting site, the requirement of cofactors and recognition sites all the restriction enzymes are categorized into four broad categories.
Type I:
The type I restriction enzyme doesn’t have a specific cleaving activity, actually because it cleaves DNA randomly far away from their recognition site, even at anywhere.
It has digestion as well as methylation activity, which means it cleaves and inserts methyl groups too, into the host bacterial DNA. Note that to perform the methylation and for the properly catalytic activity of S-adenosyl-L- methionine, ATP as a cofactor is required.
ATP is an energy molecule, we all know that right!
Type II:
The type II restriction enzymes are a bit different than the type I and III, cleave DNA within or near the recognition site. Due to this reason, type II RE is commonly used in gene cloning and genetic engineering experiments.
They can produce a distinct banding pattern in a gel, the type II RE uses the magnesium ion as their cofactor. But here, unlike Type, I, ATP or energy molecules are not consumed.
On cleaving it can create either blunt ends or sticky ends, which depends on where it performs the catalytic activity. Restriction enzymes of whole class II have only restricted digestion activity and lack methylation activity.
HindIII, NotI and Hhal are common examples of this family.
Type III:
The type III REes is much like the type I but are slightly different, having cleaving sites near the recognition site after 20 to 30 base pairs.
Most bacterial restriction digestion systems possess these types of enzymes frequently and henceforth helps in invading viral attacks.
Though ATP and S-adenosyl- L- methionine is present, but not required during the catalytic reaction.
EcoP15, EcoPI and HinfIII are the three examples of it.
Besides these three major categories, other enzymes are classified in Type IV which typically act on the methylated DNA and Type V that uses the guided RNA for invading the viral attack.
To understand the present topic more precisely, we need to look after one of the important topics -the recognition site to be learned first.
Recognition site:
Suppose if restriction enzymes cleave DNA randomly, it may be useless! But that is not the case. This class of enzymes has a special characteristic to identify the target location either in vivo or in vitro cut, the recognition site.
The recognition site is a double-stranded DNA sequence identifiable by a specific restriction enzyme and usually is the palindromic sequence.
The palindromic nature of sequences is like a mirror effect, reading from either end, the sequences are the same, see the example.

Almost every restriction enzyme has recognition sites having palindromic in nature and are 4, 6 or 8 nucleotide long double strands.
Some enzymes cleave DNA after the recognition site, some inside the site, some far away from it, and some very near to it. Broadly the recognition site is utilized as a ‘marker’ for identifying the cleaving site by the enzyme-restriction endonuclease.
Two types of sequences are generated on restriction digestion; that are sticky ends and blunt ends. Enzymes that produce sticky ends are highly recommended in genetic engineering experiments.
Here are examples of some recognition sites.
| Restriction endonuclease | Recognition site | Blunt ends or sticky ends. |
| EcoR1 | 5’—G/AATTC—3’
3’—CTTAA/G—5’ | Sticky ends |
| EcoRV | 5’—GAT/ATC—3’
3’—CTA/TAG—5’ | Blunt ends |
| PstI | 5’—CTGCA/G—3’
3’—G/ACGTC—5’ | Sticky ends |
| BamHI | 5’—G/GATCC—3’
3’—CCTAG/G—5’ | Sticky ends |
| BccI | 5’—CCATC(N)/—3’
3’—GGTAG(N)/—5’ | Blunt ends |
| HaeII | 5’—GCGC/—3’
3’—/CGCG—5’ | Sticky ends |
| HaeIII | 5’—GG/CC—3’
3’—CC/GG—5’ | Blunt ends |
| HindIII | 5’—A/AGCTT—3’
3’—TTCGA/A—5’ | Sticky ends |
| HinfI | 5’—G/ANTC—3’
3’—CTNA/G—5’ | Sticky ends |
| NIaIII | 5’—CATG/—3’
3’—/GTAC—5’ | Sticky ends |
Mechanism of action:
Either in vivo or in vitro systems, the catalytic activity of restriction enzymes starts after they find their recognition sequence.
However, nowadays scientists have developed novel nucleases that can even recognize more than 36bp long sequences to use in gene cloning.
The larger catalytic subunit of the enzyme binds to the cleaving site on the DNA. It uses metal ions as its cofactor to boost its activity and cleave the phosphodiester bonds between the DNA.
The phosphodiester bonds are between the two adjacent nucleotides. One subunit of enzyme cleaves phosphodiester bonds on one strand and another one cleaves the other strand.
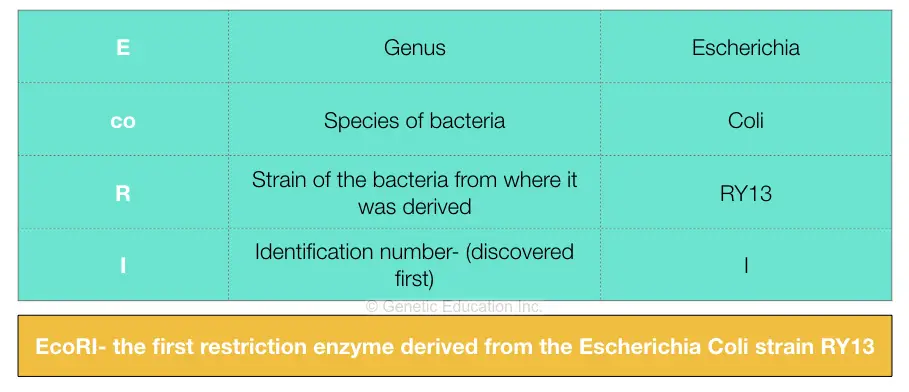
Nomenclature of restriction enzymes:
We have explained the nomenclature process in our previous article, here we have comprehensively described it in tabular form.

Applications of restriction enzymes:
Nucleases, especially endonucleases are widely used in genetic engineering, gene cloning, and PCR-based diagnostic approaches.
RFLP- restriction fragment length polymorphism is one of the well-known and widely used genetic markers that rely on the mechanism of restriction digestion.
Hereby observing the length of different fragments generated through the digestion, the results can be evaluated.
Also, the PCR-based RFLP analysis system is widely used in the identification and characterization of various genes and mutations. For example, the HinfI is used for C677T, MTHFR gene mutation studies and is quite popular.
It is also used in screening various mutations using the PCR- polymerase chain reaction-based restriction digestion.
All the results of PCR-based or direct digestion through RFLP are analyzed on the agarose gel electrophoresis. 3% of agarose gel is employed in studying the banding pattern of digestion.
A single nucleotide polymorphism can also be distinguished using the SNP-based restriction digestion and can differentiate SNP allele and non-SNP allele.
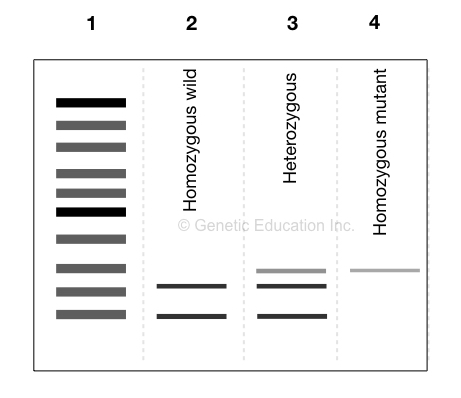
The use of restriction digestion also allows one to distinguish homozygous from heterozygous. Let us understand it with an example,
Suppose, one allele of a gene has an SNP on the recognition site of EcoRI and another allele is normal. EcoRI can cleave the normal allele, not the mutant one.
So you can imagine, on running a gel, two bands in the normal allele lane, a single band on mutant allele lane and three bands on heterozygous alleles lane are observed.
Like this!-
During gene cloning and genetic engineering, it is used to insert DNA at the target location by cleaving at a specific location on the plasmid.
ZNF- zinc finger protein is an artificial restriction endonuclease, specially designed for gene therapy experiments that can cleave DNA at a precise location.
It is widely used in generating restriction digestion maps and southern blotting.
Read more: gene mapping.
Now let’s go deeper into the topic and understand some technical things! Here are some of the commercial REs.
Top 10 Restriction endonucleases used in genetic engineering
EcoR1:
The EcoRI is one of the most widely used and known restriction endonucleases in molecular genetics. It was derived from the E.Coli strain RY13 and hence named EcoRI.
It cleaves DNA at the site of G/AATTC palindromic sequences which is present after 46 bases. It generates sticky ends.
Technically, it is a high-quality, high-speed enzyme that digests the DNA sample in 5 to 15 minutes.
EcoRV:
The EcoRV is another enzyme derived from E.Coli and types II restriction endonuclease which is also denoted as Eco32I.
It is used as a blunt end cutter and generates blunt ends. Its recognition site is 5’GAT/ATC. The EcoRV is also powerful enough to digest DNA samples within 15 minutes. It is a fast endonuclease.
PstI:
The PstI is derived from the species of gram-negative Providencia stuartii. It is categorized into type II endonuclease and has a recognition sequence of 5’ CTGCA/G3’.
It generates sticky ends during the restriction digestion.
Note that in the natural bacterial system the present endonuclease not only performs digestion but also works as a methyltransferase to protect the native DNA strand of bacteria.
It transfer’s methyl groups to protect their own DNA. PstI is widely used in gene cloning and genetic engineering experiments.
BamHI:
The BamHI is categorized into type II restriction endonuclease and is derived from the Bacillus amyloliquefaciens.
Its recognition site is 5’-GGATCC-3’ and it generates sticky ends when it digests DNA.
BccI:
The BccI is derived from the Bacteroides caccae whose recognition site is 5’CCATC-3’. It generates sticky ends when digestion.
HaeIII:
HaeII is derived from the bacteria Haemophilus aegyptius. The recognition sequence of HaeII is 5’-GG/CC-3’, a four nucleotides long common sequence that generates the blunt ends.
The heat denaturation temperature of HaeII is 80C for 20 minutes.
HaeII:
The present restriction enzyme is also derived from the Haemophilus aegyptius and it is a type II RE. Its recognition sequence is 5’-GCGC/(TC)-3,
HindIII:
HindIII is another popular molecular genetic tool used in genetic engineering experiments that are derived from the Haemophilus influenzae bacteria.
Its recognition sequence is the 6 nucleotides long palindromic sequence of 5’-A/AGCTT-3’.
It generates sticky ends. It is widely used in site-directed mutagenesis experiments.
HinfI:
HinfI is another important restriction endonuclease used in genetic labs for since long. A fast enzyme HinfI can digest DNA within 5 to 15 minutes. It generates stick ends on digestion.
NIaIII:
The NIaIII is also generating sticky ends upon digestion. It is also available in the fast digestion version with digest DNA in 15 minutes.
How restriction endonucleases are used in genetic engineering experiments?
So far we have enlisted some of the often used restriction endonucleases but you may wonder how they are used?
Well, first of all, these are enzymes and therefore should be processed at 4C temperature. At higher temperatures, its activity might decrease.
Around 10 microliter or 2 to 5 units of the enzyme is used in the reaction.
The enzyme is added to the amplicon tube and mixed with the buffer (provided by the manufacturer). Then it is placed at 37°C overnight for the catalytic reaction to happen. If it is a fast enzyme then 5 to 10 minutes are enough to digest the sample.
The enzyme cleaves the phosphodiester bond between the nucleotides, exactly where its recognition site is located.
Afterward, the products are run on 3% agarose gel. If digestion occurs two or more than two DNA fragments appear.
That is the whole process to use the restriction endonucleases in genetic engineering.
Conclusion:
Restriction enzymes are an important tool in genetics and genetic engineering experiment. ZNF like artificial restriction enzymes makes it possible to modify the recognition sites at our convenience.



This reminds me of the experiment, I did on enzymes